Impact Factor ISSN: 1449-2288
Int J Biol Sci 2017; 13(8):1019-1028. doi:10.7150/ijbs.19532 This issue Cite
Research Paper
Functional Analysis of a Novel FOXL2 Indel Mutation in Chinese Families with Blepharophimosis-Ptosis-Epicanthus Inversus Syndrome Type I
Department of Ophthalmology, Ninth People's Hospital, Shanghai Jiao Tong University School of Medicine, Shanghai, P.R. China
*These authors contributed equally to this report.
Abstract
Background: Blepharophimosis-ptosis-epicanthus inversus syndrome (BPES) is an autosomal dominant disease with a low incidence rate. Indel mutations in the forkhead box L2 (FOXL2) gene cause two types of BPES that are distinguished by the presence (type I) or absence (type II) of premature ovarian failure (POF). The purpose of this study was to identify a possible deletion in FOXL2 in Chinese families with BPES and to clarify its relationship with POF.
Methods: An autosomal dominant Chinese BPES family with four generations was enrolled in this study. Peripheral venous blood was collected from all affected patients, and genomic DNA was extracted from leukocytes. The whole coding sequence and nearby 5' untranslated region (UTR) and 3'UTR of the FOXL2 gene were amplified using polymerase chain reaction (PCR) with three sets of overlapping primers, followed by sequencing analyses. The sequencing results were analysed using SeqMan software. Based on the patients' clinical manifestations and analysis of the identified indel mutation, we found that the mutation disturbed interactions between FOXL2 and the StAR gene. Furthermore, through subcellular localisation and functional studies, we observed significant mislocalisation of the mutant protein; the mutant protein was found in the cytoplasm, while the wild-type protein was found in the nucleus. Loss of function was confirmed by transcriptional activity assays, quantitative real-time PCR, and electrophoretic mobility shift assays.
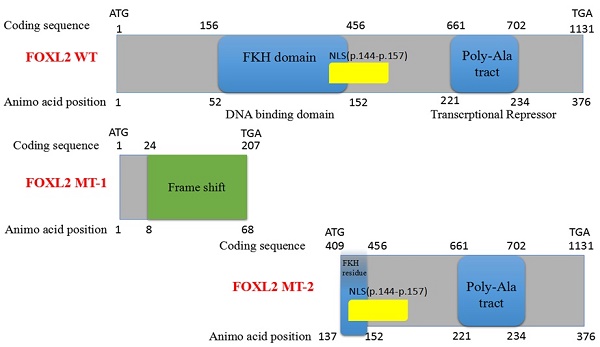
Results: All affected patients presented with clinical features of BPES type I, including small palpebral fissures, ptosis, telecanthus, and epicanthus inversus with POF. A novel FOXL2 heterozygous indel mutation, c.19_95del, a 77-bp deletion that disrupts FOXL2 protein structure, was identified in all affected members of the family. In addition, this indel mutation significantly increased StAR mRNA expression by disrupting the ability of the FOXL2 protein to bind to the StAR promoter and act as a repressor of this gene.
Conclusions: A novel FOXL2 indel mutation was identified in Chinese families with BPES. Our results expand the spectrum of known FOXL2 mutations and provide additional insight into the structure-function relationships of the FOXL2 protein. Furthermore, this novel mutation resulted in the dysfunction of FOXL2 as a transcription factor, blocking its ability to bind to the promoter region of the StAR gene, resulting in POF in the affected patient.
Keywords: BPES, FOXL2, StAR, mutation

 Global reach, higher impact
Global reach, higher impact