Impact Factor ISSN: 1449-2288
Int J Biol Sci 2018; 14(1):87-99. doi:10.7150/ijbs.22498 This issue Cite
Research Paper
Comparison of microRNA Profiles between Bovine Mammary Glands Infected with Staphylococcus aureus and Escherichia coli
1. College of Animal Science and Technology, National Beef Cattle Improvement Centre, Northwest A&F University, Yangling Shaanxi, China;
2. Key Laboratory of Zoology in Hunan Higher Education, College of Life Science, Hunan University of Arts and Science, Changde Hunan, China.
* These authors contributed equally to this work.
Abstract
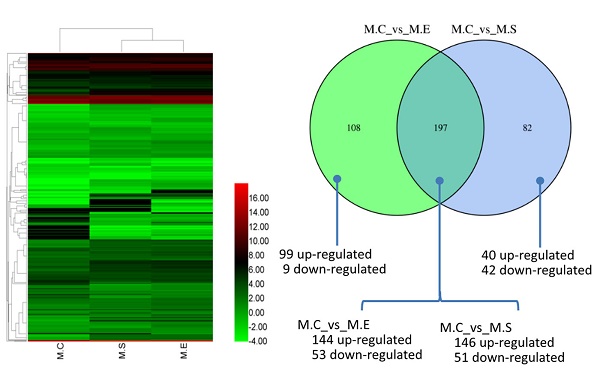
MicroRNAs (miRNAs) play crucial roles in regulating innate and adaptive immunity in humans and animals. Infection with E. coli or S. aureus can cause inflammation of the mammary glands, which results in significant economic losses in dairy cattle. However, the regulatory mechanisms of miRNAs in response to E. coli or S. aureus infection in bovine mammary glands have not been thoroughly explored. To discover the differential expression of miRNA in bovine mammary gland challenged with E. coli or S. aureus, we performed miRNA sequencing on tissue samples. A total of 1838 miRNAs were identified, including 580 known-miRNAs (included in the miRbase database) and 1258 predicted novel miRNAs. The miRNA expression patterns indicated that, compared with control samples, 279 miRNAs and 305 miRNAs were differentially expressed miRNAs (DIE-miRNA) in S. aureus and E. coli infected tissues, respectively. Moreover, the results of comparison the DIE-miRNAs between the E. coli and S. aureus infected groups showed that 197 DIE-miRNAs are identical, 108 DIE-miRNAs are specific to the E. coli group, and 82 DIE-miRNAs are specific to the S. aureus group. Many DIE-miRNAs, such as bta-miR-144, bta-miR-451 and bta-miR-7863, might be the useful biomarkers of mastitis caused by E. coli and S. aureus. In addition, target genes of the DIE-miRNAs were predicted. The Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis indicated that these DIE-miRNAs are likely involved in many immune signaling pathways, including the Toll-like receptor signaling pathways, MAPK signaling pathway, cell adhesion molecules, TGF-β signaling pathway, leukocyte trans endothelial migration, cytokine-cytokine receptor interaction, and chemokine signaling pathways. This study has provided supportive evidence that miRNAs may serve as diagnostic biomarkers of mastitis in dairy cows, and suggests potentially of effective strategies to combat mastitis.
Keywords: miRNA, E. coli, S. aureus, mastitis, dairy cattle, deep sequencing

 Global reach, higher impact
Global reach, higher impact