10
Impact Factor
ISSN: 1449-2288
Int J Biol Sci 2018; 14(8):811-818. doi:10.7150/ijbs.24624 This issue Cite
Research Paper
Integrated multifactor analysis explores core dysfunctional modules in autism spectrum disorder
1. Department of Child and Adolescent Health, School of Public Health, Harbin Medical University, Harbin, China;
2. College of Bioinformatics Science and Technology, Harbin Medical University, Harbin, China.
*These authors contributed equally to this work.
Abstract
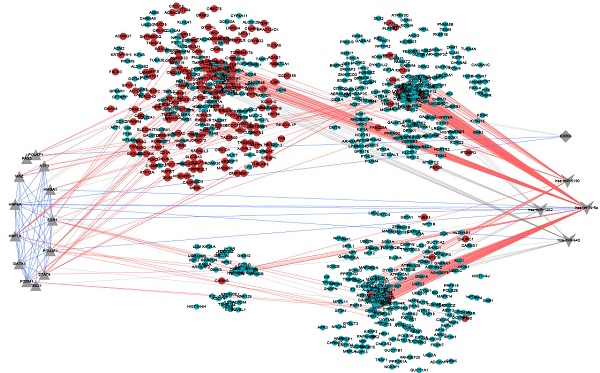
Autism spectrum disorder (ASD) is a complex neurodevelopmental disease in early childhood, and growing up to be a major cause of disability in children. However, the underlying molecular mechanism of ASD remains elusive. Hence, we represented integrated multifactor analysis exploring dysfunctional modules based on RNA-Seq data from corpus callosum in 6 patients with ASD and 6 normal individuals. According to protein-protein interactions (PPIs) and WGCNA, we performed co-expression modules analysis for ASD-associated genes, and identified 25 modules with differentially expressed genes (DEGs), observing that genes in these modules were significantly involved in various biological processes in nervous system, sensory system, phylogenetic system and variety of signaling pathways. Then, based on transcriptional and post-transcriptional regulations, integrating transcription factor (TF)-target and RNA-associated interactions, significant regulators of co-expression modules were identified as pivot regulators, including 67 pivot TFs, 13 pivot miRNAs and 6 pivot lncRNAs. GO and KEGG pathway enrichment analysis demonstrated that the pivot miRNAs significantly enriched in neural or mental-associated biological progresses. The pivot TFs were mainly involved in various regulation of transcription, immune system and organs development. Finally, our work deciphered a multifactor dysfunctional co-expression subnetwork involved in ASD, helps uncover core dysfunctional modules for this disease and improves our understanding of its underlying molecular mechanism.
Keywords: Autism spectrum disorder, Multifactor analysis, Co-expression, Core dysfunctional module

 Global reach, higher impact
Global reach, higher impact