Impact Factor ISSN: 1449-2288
Int J Biol Sci 2018; 14(12):1610-1620. doi:10.7150/ijbs.26518 This issue Cite
Review
Extracellular Vesicles: Potential Participants in Circadian Rhythm Synchronization
1. Department of Orthopaedic Surgery, Shanghai Jiao Tong University Affiliated Sixth People's Hospital, 600 Yishan Road, Shanghai 200233, China
2. Institute of Microsurgery on Extremities, Shanghai Jiao Tong University Affiliated Sixth People's Hospital, 600 Yishan Road, Shanghai 200233, China
Received 2018-4-6; Accepted 2018-7-15; Published 2018-9-7
Abstract
The circadian rhythm (CR) is a set of autonomous endogenous oscillators. Exposure to the 24-hour day-night cycle synchronizes our CR system, maintaining homeostasis and human health. Several mechanisms for the CR system have been proposed, including those underlying the function (transcriptional-translational negative-feedback loops, or TTFLs), mechanisms regulating the TTFLs, and the mechanism by which the “server clock” is synchronized to environmental time. Several pathways downstream of the “server clock” perform well-characterized biological functions. However, the synchronization between the “server clock” (the endogenous master clock seated in the suprachiasmatic nucleus within the hypothalamus) and the “client clock” (imbedded in nearly every cell in the form of interlocking TTFLs) is difficult to explain with current theories. Extracellular vesicles (EVs), which are involved in intercellular communication and have recently been found to participate in regulation of the “client clock”, might be the answer to this question. In this review, we summarize the current knowledge of CRs, TTFLs, and EVs, examine research findings about the functions of EVs in the CR system, and discuss the issues requiring attention in future research.
Keywords: circadian rhythms, extracellular vesicles, exosomes, transcriptional-translational negative-feedback loops, post-translational modifications, noncoding RNAs
1. Introduction
Almost every life-form on Earth is exposed to environmental changes over the 24-hour day-night cycle (environmental time), resulting in the evolution of circadian rhythms (CRs) to adapt to daily changes [1]. A wide variety of physiological processes are influenced by CRs, which are essential for maintaining the health of mammals, including humans [2]. In addition to the sleep-wake cycle—the most conspicuous output of CRs—there are diurnal variations in other physiological systems, including the cardiovascular system, digestive system, endocrine system, body temperature regulation, metabolism, and immune functions [3-7].
The molecular-level core of the CR system consists of oscillating clock-related genes that constitute the transcriptional-translational negative-feedback loops (TTFLs) [8]. Although the oscillators of the CR system are endogenous, exposure to the normal light/dark cycle is essential for the CR system to synchronize with environmental time and keep the body functioning normally by adapting to the environment [9-11]. However, in modern society, sun-free environments in the daytime and the use of artificial light at night have become commonplace, as these technologies have allowed human lifestyles to become increasingly flexible—people can eat, sleep, work, and exercise whenever they want [2, 9-11]. These lifestyle changes correlate with rising rates of several disorders, including heart disease [12], ulcers [13], cancer [14], somnipathy [15], diabetes [16], depression [17], and cognitive disorders [18]. There is solid evidence that night-shift work increases the risk of malignant tumours, diabetes, and cardiomyopathy [19-25]. Thus, a thorough understanding of the operating mechanism of the CR system is essential for disease prevention, early diagnosis, and effective intervention.
The endogenous CR system is mainly composed of two parts: the endogenous master clock (or “server clock”) and subordinate clocks (or “client clocks”). The endogenous “server clock” of the CR system is seated in the suprachiasmatic nucleus (SCN) in the hypothalamus [26, 27]. Through specialized retinal ganglion cells, the “server clock” is calibrated by environmental time rather than simply obeying the endogenous rhythms generated by TTFLs [28, 29]. The “client clocks” are imbedded in nearly every cell in the form of interlocking TTFLs, which are composed of clock genes that exert their biological effects via target genes (clock-controlled genes) [30, 31]. Hence, understanding of the synchronization mechanisms between “client clock” and “server clock” is crucial for a complete understanding of the CR system.
Recently, post-translational modifications (PTMs) and noncoding RNAs (ncRNAs) have been found to regulate the physiological processes of CRs [2, 32-35]. In addition, extracellular vesicles (EVs), important intercellular couriers of proteins and ncRNAs, have attracted increasing attention [36]. EVs might carry the synchronization signal to correct the “client clocks” in accordance with the “server clock”.
In this review, we will first introduce the molecular mechanism of CRs. Then, we will highlight the current knowledge concerning EVs and the regulatory mechanisms in the CR system. Finally, we will discuss how EV-mediated gene regulation could regulate the CR system. Given their emerging potential in diagnostics and even therapeutics, a thorough understanding of the role of EVs in the CR system could help tackle CR-related diseases.
2. Circadian Rhythms
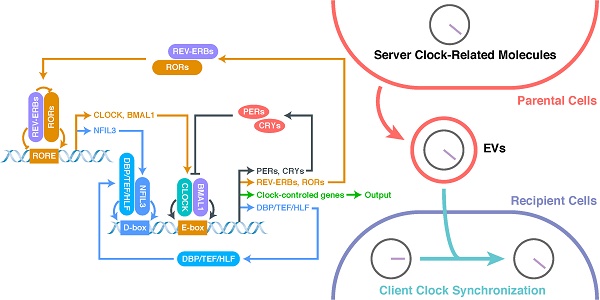
The most basic structure of the CR system is the “client clock”, a cell-autonomous circadian oscillator that exists in all cells, which is composed of clock-related proteins and is described as a network of interlocking TTFLs (Figure 1) [21, 37-40]. The core loop of TTFLs is constituted by both transcriptional activator genes [CLOCK, Brain and muscle Arnt-like protein 1 (BMAL1)] and repressor genes [Period-1 (PER1), PER2, PER3, Cryptochrome-1 (CRY1) and CRY2] [21]. The CLOCK-BMAL1 complex binds to the E-box-containing regulatory elements in repressor genes [41-43]. The working mechanism of the core loop, in brief, is that CLOCK-BMAL1 transactivation activates the transcription of repressor genes (including CRYs and PERs) together with the output genes (clock-controlled genes), and then, after translation, the accumulated CRY and PER proteins interact with each other to suppress CLOCK-BMAL1 activation (negative feedback) [38]. As the repression progresses, the protein levels of repressors, which have short half-lives and can be degraded by proteasomes, decrease, allowing a new CR cycle to begin [33, 44].
The network of interlocking TTFLs.
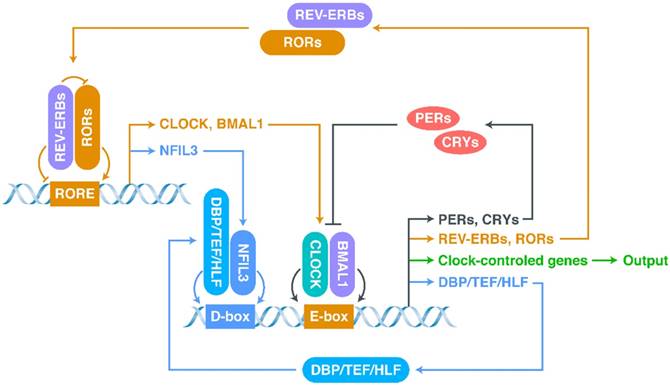
In addition to the core loop, there are two sub-loops, which are coupled to the core loop to consummate the oscillation. The first sub-loop consists of retinoic acid receptor-related orphan receptors (RORs) and nuclear receptors called REV-ERBs (including REV-ERBα and REV-ERBβ) [21, 38, 39, 45-47]. REV-ERBs compete with RORs for binding to ROR-binding elements (RORE): REV-ERBs repress, whereas RORs activate, CLOCK and BMAL1 [39, 45]. The second sub-loop consists of D-box-related proteins, including D‐box binding protein (DBP), thyrotroph embryonic factor (TEF), and hepatic leukaemia factor (HLF) [39, 48, 49]. These proteins interact with nuclear factor interleukin-3-regulated protein (NFIL3), a downstream protein of RORs/REV-ERBs, at D-boxes [39, 48, 49]. This molecular-level oscillator network is the foundation of the CR system.
The “server clock” seated in the SCN is the master of the CR system in mammals [50]—it receives the input signal (light from the external environment) via the retinohypothalamic tract to synchronize the “server clock” to environmental time [51]. The “server clock” then communicates with the wider central nervous system (CNS) and non-CNS organs [52]. Ablation of the SCN leads to dysregulation of clock genes in the “client clock” of most tissues, causing arhythmicity of behaviours and physiological functions [8]. Thus, the duty of the “server clock” is to synchronize the “client clock” to environmental time [53].
There are three classical neuroendocrine pathways governing the peripheral effects induced by the “server clock”. First, the “server clock” induces the release of epinephrine/norepinephrine via the nerve endings of the autonomic nervous system [54]. Second, it induces the release of glucocorticoids by the adrenal gland via the hypothalamic-pituitary-adrenal axis [55]. Third, it induces melatonin release by activating the pineal gland [56]. These neuroendocrine pathways and their downstream effects can explain many of the peripheral functions of the “server clock” [56]. However, these pathways cannot explain how the “client clocks” synchronize with the “server clock”.
There is mounting evidence that disruption of the CR system is intimately implicated in the pathology of neurodegenerative diseases [57-59], metabolic disorders [60], chronic inflammatory diseases [61], cardiovascular disease [62], and malignant tumours [19]. The peripheral blood cells of Parkinson's disease patients have abnormal clock gene expression [63, 64]. Kim et al. found that deletion of the secretory vesicle proteins IA-2 and IA-2β, which have also recently been found to be closely related to EV secretion, affects the CR system [65, 66]. These findings suggest that the pathogenesis of CR-related diseases is complicated and multi-dimensional (at the level of the “server clock”, the “client clock”, or in the signal transduction between them, for example).
3. Regulatory Mechanisms of the TTFLs
Recent research has been focused on the multi-level regulatory mechanisms of TTFLs, which can be classified into groups depending on their main level of action: PTMs and ncRNAs. Both the “server clock” and the “client clock” are precisely regulated by PTMs [33-35].
Both the total level and the phosphorylated level of PER proteins dramatically influence the oscillations of the CR system [38, 67]. The main modification of PERs are shown in Figure 2A. In most cases, PER1-3 are phosphorylated by casein kinase I (CKI) and then poly-ubiquitinated by the Skp-Cullin-F-box (SCF) complex containing the F-box type E3 ligase β-TrCP1 or β-TrCP2, before degradation by the proteasome [38, 68, 69]. PER2 is phosphorylated at Ser53 by CKII and then degraded [38]. However, the different phosphorylation sites on PER2 seem to have different effects on the stability of PER2 and thus on oscillations. Phosphorylation of Ser662 (and the four downstream serines Ser665, Ser668, Ser671, and Ser674) leads to the stabilization of PER2 (Figure 2D) [38, 70, 71].
Regulatory Mechanisms of the TTFLs. The modification of (A) PERs, (B) CRYs and (C) BMAL1-CLOCK as well as some special cases (D and E).
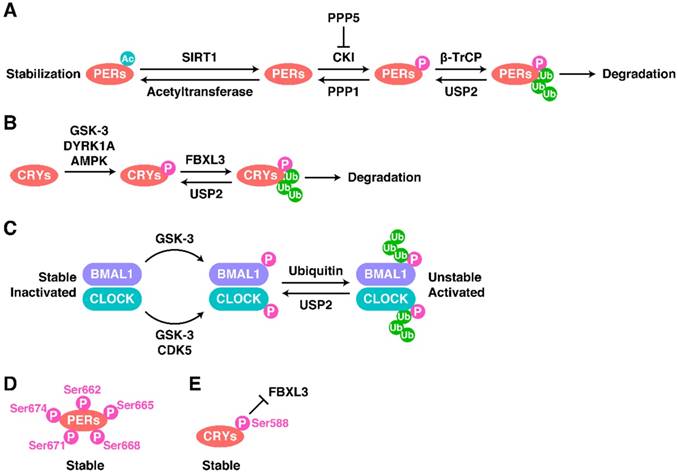
The opposite of phosphorylation is de-phosphorylation. Phosphoprotein phosphatase 1 (PPP1) can de-phosphorylate PERs and thus antagonize the CKI-mediated degradation of PERs [38, 72]. PPP5 regulates the stabilization of PERs and the CR cycle by preventing the auto-phosphorylation of CKI [72, 73]. Moreover, de-ubiquitination and acetylation can also stabilize proteins. For example, BMAL1, PER1, and CRY1-2 can be de-ubiquitinated by ubiquitin-specific protease 2 (USP2), stabilizing them [72, 74, 75]. SIRT1 binds to PER2, inducing its de-acetylation and promoting its degradation [76, 77].
The phosphorylation and ubiquitination of CRY proteins is also important, and a diagram showing the main modification of CRYs is shown in Figure 2B. AMP-activated protein kinase (AMPK) phosphorylates CRY1 at Ser71 to induce the degradation of CRY1 by recruiting FBXL3 [78]. The FBXL3-containing SCF complex induces the ubiquitination of CRY proteins and terminates their transcriptional repression activity by promoting their degradation [79, 80]. Dual-specificity tyrosine-phosphorylation-regulated kinase 1A (DYRK1A) phosphorylates Ser557 on the C-terminus of CRY2 [81], and glycogen synthase kinase 3 (GSK-3) induces the secondary phosphorylation of CRY2 at Ser553 [81, 82]. After this two-step phosphorylation, CRY2 is degraded by an unknown E3 ligase [81]. The phosphorylation of CRY1 at Ser588 blocks the interaction between FBXL3 and CRY1, stabilizing CRY1 (Figure 2E), but the responsible kinase remains unknown [83].
The mechanisms regulating CLOCK and BMAL1 act in a completely different way to those just described (Figure 2C). First, the total protein levels of CLOCK and BMAL1 are relatively stable, but their phosphorylation levels are circadian [67, 84]. Phosphorylation decreases the stability of CLOCK and BMAL1 but promotes their activity [85, 86]. Cyclin-dependent kinase 5 (CDK5) phosphorylates CLOCK at Thr451 and Thr461 [87]. GSK-3 phosphorylates CLOCK at Ser427 and BMAL1 at Ser17 [85, 86].
Noncoding RNAs are also involved in the regulation of the CR system. Researchers have developed a computational model involving PERs, CRYs, CLOCK, BMAL1, and two microRNAs (miRNAs; miR-219 and miR-132) [88-90]. Using this model, it was found that the most studied CR-related miRNAs to date, miR-219 and miR-132, can activate the translation of PER1 and regulate the mammalian CR system [88, 89, 91, 92]. MiR-219 in the SCN shows a rhythmicity correlated to the CR and has its peak expression level at midday [88, 93]. Photo-activated expression of miR-132 requires CREB and MAPK/ERK [88, 94, 95].
In addition, circulating miR-494 and miR-142-3p can modulate CRs by targeting the clock gene BMAL1 [96, 97], while miR-433 regulates the expression of PER2 and BMAL1 [98]. BMAL1 is also targeted by miR-27b-3p [113] and miR-155 [99], while PER1 is targeted by miR-34a [100], and the miR-192/194 cluster can inhibit the PER family [101]. Finally, translation of CRY1 is regulated by miR-185 [102].
The lncRNA TUG-1 is required for photoreceptor differentiation, but the underlying mechanism is still unknown [88, 103]. The level of the lncRNA HULC is positively correlated with the expression level of CLOCK and can upregulate its downstream output genes [104]. However, it is essential to screen more potential CR-related ncRNAs, and study their mechanisms and functions in order to broaden understanding of the regulatory network of CR.
4. Basic Concepts of Extracellular Vesicles
EV secretion was initially regarded as a process to eliminate unwanted compounds from cells [105, 106]. However, pioneers in the field (Raposo et al.) found, in 1996, that EVs play important roles in immune responses [107]. Since then, the functions of EVs in intercellular communication have drawn increasing attention and the number of annual citations has dramatically increased, from 28 in 1996 to 24,765 in 2016 [108]. Almost all eukaryotic cells take up and secrete EVs, and these minute EVs contain genetic instructions (nucleic acids and proteins) that regulate the function of the recipient cells, whether under normal or pathological conditions, sometimes mildly, sometimes strongly [108]. Moreover, the communication process mediated by EVs is conserved from bacteria to plants and animals [109, 110]. Most interestingly, there is much evidence of cross-species communication via EVs, even between micro-organisms and mammals [111].
EVs can be roughly classified into three categories: exosomes (EXOs), microvesicles (MVs), and apoptotic bodies (ABs) [106, 112-114]. EXOs and MVs are the most important EVs in intercellular communication, while ABs are rapidly eliminated by immune cells [115]. EXOs (also known as inward-budding vesicles; 30-100 nm in diameter) are generated on endosomal membranes by inward budding during the maturation process of multi-vesicular bodies (MVBs) [105, 106, 116, 117]. Before MVBs fuse with the plasma membrane and release EXOs, EXOs in MVBs are called intra-luminal vesicles [106, 116, 117]. MVs (also known as outward-budding vesicles; 50-1,000 nm in diameter) are generated and released on the plasma membrane by outward budding [106, 113] and were initially studied for their important roles in blood coagulation [106, 118, 119]. However, new research suggests MVs are important players in intercellular communication [106, 120].
Although there are differences between MVs and EXOs in terms of their biogenesis and release, most research on the biological function of EVs has not strictly accounted for the differences in their intracellular origins. Many researchers do not strictly consider small EVs, isolated by gradient centrifugation or 0.22 μm filters, as EXOs [121, 122]. The mainstream view now holds that there is no need to make a detailed distinction between EXOs and MVs, if the research is focused on their pathological and physiological functions, rather than their biogenesis. In this review, we have called them both EVs, in accord with most reviews discussing EVs [123, 124].
5. Extracellular Vesicle-Mediated Intercellular Communication
EVs contain and transport several types of molecules, including membrane proteins and cytosolic proteins, messenger RNAs, and ncRNAs (such as miRNAs, lncRNAs, and circular RNAs) (Figure 3A) [113, 115, 125]. The regulatory functions of EVs in signalling pathways of recipient cells are based on either receptor-ligand interactions or direct content delivery after internalization [115, 123]. Their phospholipid bilayers enable EVs to protect their cargo from the external environment, and they are found in almost all body fluids [124, 126, 127]. The different surface proteins expressed on EVs give them different targeting properties [128].
Potential Participants in CR Synchronization. (A) The constitution of EVs. (B) The cargoes of EVs might be the key of CR synchronization.
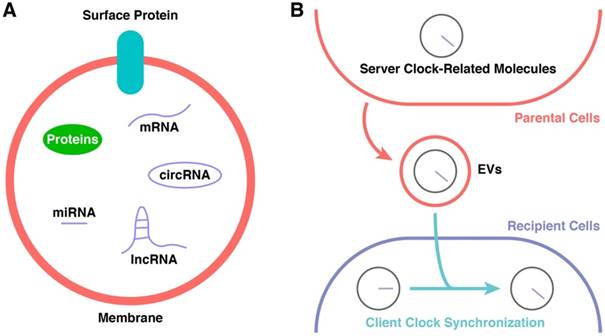
EVs can be derived from nearly every type of cell, both normal [129, 130] and malignant [131, 132]. Normal cells use EVs to coordinate, communicate, and cooperate with their “colleagues” [115]. Malignant cells use EVs to mislead normal cells and issue aberrant orders. EVs derived from malignant cells can deliver signals to establish a pre-metastatic niche, a suitable environment for metastasis [133-136]. The constitution of EVs derived from malignant cells is markedly different from that of EVs derived from normal cells [137, 138], and this makes them potential biomarkers in liquid biopsies. EVs also show potential as diagnostic markers and progression markers of infectious diseases [139-141]. Hence, it is also possible to choose EVs as potential markers to monitor the functions of the CR and to detect early warning signals related to dysfunction of the internal CR system.
EVs are also potential therapeutic tools to regulate and maintain the internal CR system and prevent/treat disease caused by dysfunction of the internal CR system. Stem cells once showed great potential in regenerative medicine because of their unparalleled proliferation/differentiation potential [142]. However, embryonic stem cells and induced pluripotent stem (iPS) cells have shown key weaknesses, including immunological rejection, tumour formation, and circulation damage [124, 143]. In recent years, EVs have become rising stars in the field of regenerative medicine for use in cell-free therapy. Increasing evidence suggests that stem/progenitor cell-derived EVs have therapeutic functions similar to or even better than their parent cells [124, 144]. In addition, EVs are stable and easy to preserve because they can retain bioactivity during lyophilisation and other extreme conditions during handling, owing to the protective effect of their lipid bilayer [124, 145]. Moreover, EVs can cross barriers such as the blood-brain barrier [136], so the route of administration is relatively easy. Above all, therapeutic use of EVs is safer than direct use of their parent cells because of their hypoallergenic nature and lack of oncogenic potential [124]. Evidence of their hypoallergenicity includes the fact that human-derived EVs work well upon first injection and repeated injections in animal models, including rats, mice, and pigs [108]. Adamiak et al. found that over half of the mice they injected with iPS cells developed teratomas, whereas mice injected with EVs derived from iPS cells (even from the same cells that had caused teratomas) did not develop teratomas [144]. Stem/progenitor cell-derived EVs (or even EVs derived from fragments of cytoplasm such as platelets) regulate the biological processes of the target cells by delivering parent cell-originated nucleic acids and proteins [124, 126, 146-148].
Natural EVs cannot always meet therapeutic needs, and developing improved/modified EVs will be an exciting area of future research [124, 126, 127, 149, 150]. For example, EVs derived from synovial mesenchymal stem cells enhance the proliferation and migration of chondrocytes, but reduce the formation of extracellular matrix [127]. This problem can be solved by enhancing the levels of miR-140-5p in these EVs [127]. Modified or optimized EVs might therefore be the future of EV-based therapeutic medicine. With advances in deep knowledge of the CR system and of the technology of EV-based therapy, preventing or reversing malfunctions, caused by modern life such as night shift work and jet lag, will become a reality.
6. Circadian Rhythms and Extracellular Vesicles
Recent research has suggested that EVs act as a bridge between the “server clock” and “client clock” (Figure 3B). Khalyfa et al. employed a mouse model of chronic nocturnal shift work and found altered intestinal flora and increased colonic cell permeability accompanied by changes in the components of plasma EVs, including clock genes [36]. SIRT1, which is involved in the regulation of PER acetylation (see previous section), is regulated by EVs [151, 152]. EVs also regulate the phosphorylation of GSK-3 [153], which plays an important role in the regulation of CRYs and BMAL1/CLOCK (see previous section). AMPK, which participates in the regulation of CRYs (see previous section), is regulated by EVs and is probably related to ncRNAs in EVs [154-156].
MiR-132, which plays an important role in regulating PERs (see previous section), is enriched in EVs derived from fibrocytes, adipose tissue-derived stem cells, cardiac progenitor cells, neurons, serum, and umbilical cord blood [157-162]. MiR-219, which participates in activating PER1 translation (see the previous section), is enriched in serum-derived EVs [163]. Therefore, although direct links have yet to be shown, these results suggest that EVs in the circulatory system are communicators between the “server clock” and the “client clock”.
An online database of EVs, EVpedia (http://student4.postech.ac.kr/evpedia2_xe/xe/), could help to discover the existence of CR-related genes in EVs [164-168]. The database confirmed the existence of PER2, PER3, CKI families, AMPKβ, AMPKγ, GSK-3α, and GSK-3β, as well as miR-219a, miR-219b, miR-132, miR-494, miR-142, miR-433, miR-27b-3p, miR-192, and miR-194 in EVs. Thus, EVs contain enough bioactive molecules to regulate the “client clock” of recipient cells, so it is essential to verify whether EVs participate in liaising between the “server clock” and the “client clock”. It is important not only to detect any additional CR-related molecules carried by EVs but also to verify the idiographic functions of these molecules.
Shende et al. found that several circulating miRNAs (miR-152, miR-494, and miR-142-3p) correlate with diurnal oscillation (expression peaks near midday and 8/12 h later) and participate in the regulation of clock genes in a mouse model [96]. However, this study did not distinguish whether these miRNAs were free in the plasma or packaged in EVs. Circulating miR-494 and miR-142-3p can regulate CRs by targeting the clock gene BMAL1 [96], but the locations of these miRNAs were not confirmed.
In blood, there are two major forms of circulating miRNAs: miRNAs as “cargos” of EVs (miRNA-EVs) and miRNAs bound to Argonaute (AGO) proteins (miRNA-AGOs) [169]. It is generally believed that miRNA-AGOs cannot be internalized by recipient cells because of their size and lack of bioactivity for penetration [169-171], but miRNA-EVs can be efficiently internalized with their miRNAs by recipient cells via penetration [170, 172]. When viewed from this perspective, the circulating miRNAs that regulate CR-related functions stand a good chance of being carried in EVs. Nonetheless, it is important to evaluate not only the total content of genetic material in plasma but also their specific locations (whether enclosed in EVs or not).
7. Discussion and Outlook
As outlined above, the mechanism underlying the CR appears to be a complicated, interconnected, and multi-level system. Although the fundamental structure and some regulatory mechanisms have been identified, many questions remain. In particular, current knowledge cannot explain the specific synchronization methods between “server clock” and “client clock”. Before reaching full understanding of the synchronization methods, the regulation of TTFLs at the molecular level needs to be better understood. In this review, we have summarized current understanding of the regulation of TTFLs, including by PTM and ncRNAs, in mammals. It is possible that the “server clock” transports PTM-related molecules or ncRNAs to the “client clock” through some as-yet unidentified mechanism, which may be EVs.
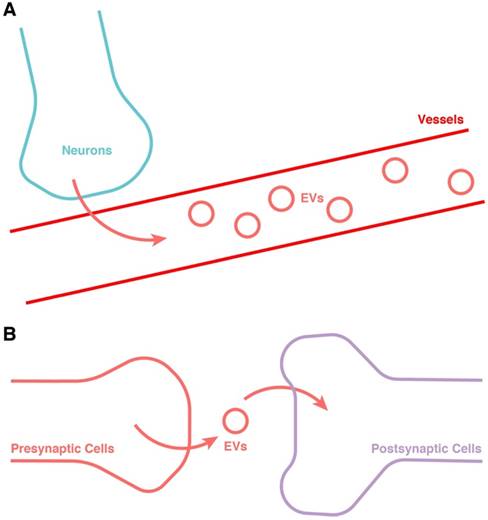
In addition to the transport of EVs in blood, the transmission of EVs between neurons is also worthy of serious attention [173] (Figure 4). The release of EVs has been found to be dependent on synaptic activity [174], and could be an important intermediary in neuron-neuron communication [175, 176]. In the larvae of Drosophila, EVs have also been found to be involved in controlling the retrograde postsynaptic signal [177]. The function of EVs related to synaptic activity could be explained by the function of EV-containing cargos including miRNAs and synaptic-associated proteins [178]. This might be not only an important addition to the communication between “server clock” and CNS besides electroneurographic signals, but also a potential jigsaw of the entirety of CR synchronization. Hence, this research area also merits much further attention.
Potential approach for the release of EVs from the “server clock”. (A) Release into the blood. (B) Transportation from neuron to neuron through the synapse.
Some evidence indicates that EVs participate in the regulation of individual cells' “client clocks”. Circulating RNAs also have important roles in the regulation of “client clocks”. Although the localization of these RNAs is not known, it is well recognized that circulating RNAs with biological effects are mainly found in EVs. It is very important to verify the localization of these circulating RNAs and their specific biological effects in the regulation of “client clocks”. An online database containing a large amount of high-throughput data such as EVpedia can greatly advance the progress of research. However, it will be necessary to establish an online database of the CR system, and carry out multi-database joint analysis to bring a breakthrough in this research area.
Acknowledgements
The National Natural Science Foundation of China [Nos. 81871834, 81802226 and 81301589] supported this work.
Author Contributions
Shi-Cong Tao planned and wrote this manuscript. Shang-Chun Guo helped with planning and writing. All authors reviewed the manuscript.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Konopka RJ, Benzer S. Clock mutants of Drosophila melanogaster. Proceedings of the National Academy of Sciences of the United States of America. 1971;68:2112-6
2. Eckel-Mahan K, Sassone-Corsi P. Metabolism and the circadian clock converge. Physiological reviews. 2013;93:107-35
3. Hardin PE, Hall JC, Rosbash M. Feedback of the Drosophila period gene product on circadian cycling of its messenger RNA levels. Nature. 1990;343:536-40
4. Partch CL, Green CB, Takahashi JS. Molecular architecture of the mammalian circadian clock. Trends in cell biology. 2014;24:90-9
5. Yin L, Wu N, Curtin JC, Qatanani M, Szwergold NR, Reid RA. et al. Rev-erbalpha, a heme sensor that coordinates metabolic and circadian pathways. Science (New York, NY). 2007;318:1786-9
6. Raghuram S, Stayrook KR, Huang P, Rogers PM, Nosie AK, McClure DB. et al. Identification of heme as the ligand for the orphan nuclear receptors REV-ERBalpha and REV-ERBbeta. Nature structural & molecular biology. 2007;14:1207-13
7. Bass J, Lazar MA. Circadian time signatures of fitness and disease. Science (New York, NY). 2016;354:994-9
8. Musiek ES, Holtzman DM. Mechanisms linking circadian clocks, sleep, and neurodegeneration. Science (New York, NY). 2016;354:1004-8
9. Bedrosian TA, Nelson RJ. Influence of the modern light environment on mood. Molecular psychiatry. 2013;18:751-7
10. Fonken LK, Nelson RJ. The effects of light at night on circadian clocks and metabolism. Endocrine reviews. 2014;35:648-70
11. Hastings MH, Reddy AB, Maywood ES. A clockwork web: circadian timing in brain and periphery, in health and disease. Nature reviews Neuroscience. 2003;4:649-61
12. Kivimaki M, Virtanen M, Elovainio M, Vaananen A, Keltikangas-Jarvinen L, Vahtera J. Prevalent cardiovascular disease, risk factors and selection out of shift work. Scandinavian journal of work, environment & health. 2006;32:204-8
13. Koda S, Yasuda N, Sugihara Y, Ohara H, Udo H, Otani T. et al. [Analyses of work-relatedness of health problems among truck drivers by questionnaire survey]. Sangyo eiseigaku zasshi = Journal of occupational health. 2000;42:6-16
14. Ball LJ, Palesh O, Kriegsfeld LJ. The Pathophysiologic Role of Disrupted Circadian and Neuroendocrine Rhythms in Breast Carcinogenesis. Endocrine reviews. 2016;37:450-66
15. Trenell MI, Marshall NS, Rogers NL. Sleep and metabolic control: waking to a problem? Clinical and experimental pharmacology & physiology. 2007;34:1-9
16. Morikawa Y, Nakagawa H, Miura K, Soyama Y, Ishizaki M, Kido T. et al. Shift work and the risk of diabetes mellitus among Japanese male factory workers. Scandinavian journal of work, environment & health. 2005;31:179-83
17. Bildt C, Michelsen H. Gender differences in the effects from working conditions on mental health: a 4-year follow-up. International archives of occupational and environmental health. 2002;75:252-8
18. Cho K, Ennaceur A, Cole JC, Suh CK. Chronic jet lag produces cognitive deficits. The Journal of neuroscience: the official journal of the Society for Neuroscience. 2000;20:Rc66
19. Stevens RG, Brainard GC, Blask DE, Lockley SW, Motta ME. Breast cancer and circadian disruption from electric lighting in the modern world. CA: a cancer journal for clinicians. 2014;64:207-18
20. Breast cancer on the night shift. Lancet (London, England). 2009; 373: 1054.
21. Bass J, Takahashi JS. Circadian integration of metabolism and energetics. Science (New York, NY). 2010;330:1349-54
22. Pan A, Schernhammer ES, Sun Q, Hu FB. Rotating night shift work and risk of type 2 diabetes: two prospective cohort studies in women. PLoS medicine. 2011;8:e1001141
23. van den Berg MP, Viersma JW, Beaufort-Krol GC, Bink-Boelkens MT, Bezzina CR, Veldkamp MW. et al. A large family characterised by nocturnal sudden death. Netherlands heart journal: monthly journal of the Netherlands Society of Cardiology and the Netherlands Heart Foundation. 2002;10:304-12
24. Vetter C, Devore EE, Wegrzyn LR, Massa J, Speizer FE, Kawachi I. et al. Association Between Rotating Night Shift Work and Risk of Coronary Heart Disease Among Women. Jama. 2016;315:1726-34
25. Steptoe A. Night shift work and the cardiovascular health of medical staff. European heart journal. 2009;30:2560-1
26. Stephan FK, Zucker I. Circadian rhythms in drinking behavior and locomotor activity of rats are eliminated by hypothalamic lesions. Proceedings of the National Academy of Sciences of the United States of America. 1972;69:1583-6
27. Moore RY, Eichler VB. Loss of a circadian adrenal corticosterone rhythm following suprachiasmatic lesions in the rat. Brain research. 1972;42:201-6
28. Berson DM, Dunn FA, Takao M. Phototransduction by retinal ganglion cells that set the circadian clock. Science (New York, NY). 2002;295:1070-3
29. Panda S, Provencio I, Tu DC, Pires SS, Rollag MD, Castrucci AM. et al. Melanopsin is required for non-image-forming photic responses in blind mice. Science (New York, NY). 2003;301:525-7
30. Zhang EE, Kay SA. Clocks not winding down: unravelling circadian networks. Nature reviews Molecular cell biology. 2010;11:764-76
31. Yamamoto T, Nakahata Y, Soma H, Akashi M, Mamine T, Takumi T. Transcriptional oscillation of canonical clock genes in mouse peripheral tissues. BMC molecular biology. 2004;5:18
32. Hardin PE. Molecular genetic analysis of circadian timekeeping in Drosophila. Advances in genetics. 2011;74:141-73
33. Gallego M, Virshup DM. Post-translational modifications regulate the ticking of the circadian clock. Nature reviews Molecular cell biology. 2007;8:139-48
34. Reischl S, Kramer A. Kinases and phosphatases in the mammalian circadian clock. FEBS letters. 2011;585:1393-9
35. Stojkovic K, Wing SS, Cermakian N. A central role for ubiquitination within a circadian clock protein modification code. Frontiers in molecular neuroscience. 2014;7:69
36. Khalyfa A, Poroyko VA, Qiao Z, Gileles-Hillel A, Khalyfa AA, Akbarpour M. et al. Exosomes and Metabolic Function in Mice Exposed to Alternating Dark-Light Cycles Mimicking Night Shift Work Schedules. Frontiers in physiology. 2017;8:882
37. Dunlap JC. Molecular bases for circadian clocks. Cell. 1999;96:271-90
38. Hirano A, Fu YH, Ptacek LJ. The intricate dance of post-translational modifications in the rhythm of life. Nature structural & molecular biology. 2016;23:1053-60
39. Takahashi JS. Transcriptional architecture of the mammalian circadian clock. Nature reviews Genetics. 2017;18:164-79
40. Lowrey PL, Takahashi JS. Mammalian circadian biology: elucidating genome-wide levels of temporal organization. Annual review of genomics and human genetics. 2004;5:407-41
41. Gekakis N, Staknis D, Nguyen HB, Davis FC, Wilsbacher LD, King DP. et al. Role of the CLOCK protein in the mammalian circadian mechanism. Science (New York, NY). 1998;280:1564-9
42. Kume K, Zylka MJ, Sriram S, Shearman LP, Weaver DR, Jin X. et al. mCRY1 and mCRY2 are essential components of the negative limb of the circadian clock feedback loop. Cell. 1999;98:193-205
43. Shearman LP, Sriram S, Weaver DR, Maywood ES, Chaves I, Zheng B. et al. Interacting molecular loops in the mammalian circadian clock. Science (New York, NY). 2000;288:1013-9
44. Preussner M, Heyd F. Post-transcriptional control of the mammalian circadian clock: implications for health and disease. Pflugers Archiv: European journal of physiology. 2016;468:983-91
45. Preitner N, Damiola F, Lopez-Molina L, Zakany J, Duboule D, Albrecht U. et al. The orphan nuclear receptor REV-ERBalpha controls circadian transcription within the positive limb of the mammalian circadian oscillator. Cell. 2002;110:251-60
46. Sato TK, Panda S, Miraglia LJ, Reyes TM, Rudic RD, McNamara P. et al. A functional genomics strategy reveals Rora as a component of the mammalian circadian clock. Neuron. 2004;43:527-37
47. Zhang Y, Fang B, Emmett MJ, Damle M, Sun Z, Feng D. et al. GENE REGULATION. Discrete functions of nuclear receptor Rev-erbalpha couple metabolism to the clock. Science (New York, NY). 2015;348:1488-92
48. Mitsui S, Yamaguchi S, Matsuo T, Ishida Y, Okamura H. Antagonistic role of E4BP4 and PAR proteins in the circadian oscillatory mechanism. Genes & development. 2001;15:995-1006
49. Gachon F, Fonjallaz P, Damiola F, Gos P, Kodama T, Zakany J. et al. The loss of circadian PAR bZip transcription factors results in epilepsy. Genes & development. 2004;18:1397-412
50. Hood S, Amir S. The aging clock: circadian rhythms and later life. The Journal of clinical investigation. 2017;127:437-46
51. Welsh DK, Takahashi JS, Kay SA. Suprachiasmatic nucleus: cell autonomy and network properties. Annual review of physiology. 2010;72:551-77
52. Dibner C, Schibler U. Circadian timing of metabolism in animal models and humans. Journal of internal medicine. 2015;277:513-27
53. Mohawk JA, Green CB, Takahashi JS. Central and peripheral circadian clocks in mammals. Annual review of neuroscience. 2012;35:445-62
54. Geiger SS, Fagundes CT, Siegel RM. Chrono-immunology: progress and challenges in understanding links between the circadian and immune systems. Immunology. 2015;146:349-58
55. Spies CM, Hoff P, Mazuch J, Gaber T, Maier B, Strehl C. et al. Circadian rhythms of cellular immunity in rheumatoid arthritis: a hypothesis-generating study. Clinical and experimental rheumatology. 2015;33:34-43
56. Paganelli R, Petrarca C, Di Gioacchino M. Biological clocks: their relevance to immune-allergic diseases. Clinical and molecular allergy: CMA. 2018;16:1
57. Kondratova AA, Kondratov RV. The circadian clock and pathology of the ageing brain. Nature reviews Neuroscience. 2012;13:325-35
58. Mattis J, Sehgal A. Circadian Rhythms, Sleep, and Disorders of Aging. Trends in endocrinology and metabolism: TEM. 2016;27:192-203
59. Abbott SM, Videnovic A. Chronic sleep disturbance and neural injury: links to neurodegenerative disease. Nature and science of sleep. 2016;8:55-61
60. McFadden E, Jones ME, Schoemaker MJ, Ashworth A, Swerdlow AJ. The relationship between obesity and exposure to light at night: cross-sectional analyses of over 100,000 women in the Breakthrough Generations Study. American journal of epidemiology. 2014;180:245-50
61. Lucassen EA, Coomans CP, van Putten M, de Kreij SR, van Genugten JH, Sutorius RP. et al. Environmental 24-hr Cycles Are Essential for Health. Current biology: CB. 2016;26:1843-53
62. Morris CJ, Purvis TE, Hu K, Scheer FA. Circadian misalignment increases cardiovascular disease risk factors in humans. Proceedings of the National Academy of Sciences of the United States of America. 2016;113:E1402-11
63. Breen DP, Vuono R, Nawarathna U, Fisher K, Shneerson JM, Reddy AB. et al. Sleep and circadian rhythm regulation in early Parkinson disease. JAMA neurology. 2014;71:589-95
64. Cai Y, Liu S, Sothern RB, Xu S, Chan P. Expression of clock genes Per1 and Bmal1 in total leukocytes in health and Parkinson's disease. European journal of neurology. 2010;17:550-4
65. Kim SM, Power A, Brown TM, Constance CM, Coon SL, Nishimura T. et al. Deletion of the secretory vesicle proteins IA-2 and IA-2beta disrupts circadian rhythms of cardiovascular and physical activity. FASEB journal: official publication of the Federation of American Societies for Experimental Biology. 2009;23:3226-32
66. Cianciaruso C, Phelps EA, Pasquier M, Hamelin R, Demurtas D, Alibashe Ahmed M. et al. Primary Human and Rat beta-Cells Release the Intracellular Autoantigens GAD65, IA-2, and Proinsulin in Exosomes Together With Cytokine-Induced Enhancers of Immunity. Diabetes. 2017;66:460-73
67. Lee C, Etchegaray JP, Cagampang FR, Loudon AS, Reppert SM. Posttranslational mechanisms regulate the mammalian circadian clock. Cell. 2001;107:855-67
68. Eide EJ, Woolf MF, Kang H, Woolf P, Hurst W, Camacho F. et al. Control of mammalian circadian rhythm by CKIepsilon-regulated proteasome-mediated PER2 degradation. Molecular and cellular biology. 2005;25:2795-807
69. Reischl S, Vanselow K, Westermark PO, Thierfelder N, Maier B, Herzel H. et al. Beta-TrCP1-mediated degradation of PERIOD2 is essential for circadian dynamics. Journal of biological rhythms. 2007;22:375-86
70. Vanselow K, Vanselow JT, Westermark PO, Reischl S, Maier B, Korte T. et al. Differential effects of PER2 phosphorylation: molecular basis for the human familial advanced sleep phase syndrome (FASPS). Genes & development. 2006;20:2660-72
71. Shanware NP, Hutchinson JA, Kim SH, Zhan L, Bowler MJ, Tibbetts RS. Casein kinase 1-dependent phosphorylation of familial advanced sleep phase syndrome-associated residues controls PERIOD 2 stability. The Journal of biological chemistry. 2011;286:12766-74
72. Gallego M, Kang H, Virshup DM. Protein phosphatase 1 regulates the stability of the circadian protein PER2. The Biochemical journal. 2006;399:169-75
73. Partch CL, Sancar A. Cryptochromes and circadian photoreception in animals. Methods in enzymology. 2005;393:726-45
74. Scoma HD, Humby M, Yadav G, Zhang Q, Fogerty J, Besharse JC. The de-ubiquitinylating enzyme, USP2, is associated with the circadian clockwork and regulates its sensitivity to light. PloS one. 2011;6:e25382
75. Yang Y, Duguay D, Bedard N, Rachalski A, Baquiran G, Na CH. et al. Regulation of behavioral circadian rhythms and clock protein PER1 by the deubiquitinating enzyme USP2. Biology open. 2012;1:789-801
76. Asher G, Gatfield D, Stratmann M, Reinke H, Dibner C, Kreppel F. et al. SIRT1 regulates circadian clock gene expression through PER2 deacetylation. Cell. 2008;134:317-28
77. Nakahata Y, Kaluzova M, Grimaldi B, Sahar S, Hirayama J, Chen D. et al. The NAD+-dependent deacetylase SIRT1 modulates CLOCK-mediated chromatin remodeling and circadian control. Cell. 2008;134:329-40
78. Lamia KA, Sachdeva UM, DiTacchio L, Williams EC, Alvarez JG, Egan DF. et al. AMPK regulates the circadian clock by cryptochrome phosphorylation and degradation. Science (New York, NY). 2009;326:437-40
79. Godinho SI, Maywood ES, Shaw L, Tucci V, Barnard AR, Busino L. et al. The after-hours mutant reveals a role for Fbxl3 in determining mammalian circadian period. Science (New York, NY). 2007;316:897-900
80. Busino L, Bassermann F, Maiolica A, Lee C, Nolan PM, Godinho SI. et al. SCFFbxl3 controls the oscillation of the circadian clock by directing the degradation of cryptochrome proteins. Science (New York, NY). 2007;316:900-4
81. Kurabayashi N, Hirota T, Sakai M, Sanada K, Fukada Y. DYRK1A and glycogen synthase kinase 3beta, a dual-kinase mechanism directing proteasomal degradation of CRY2 for circadian timekeeping. Molecular and cellular biology. 2010;30:1757-68
82. Harada Y, Sakai M, Kurabayashi N, Hirota T, Fukada Y. Ser-557-phosphorylated mCRY2 is degraded upon synergistic phosphorylation by glycogen synthase kinase-3 beta. The Journal of biological chemistry. 2005;280:31714-21
83. Gao P, Yoo SH, Lee KJ, Rosensweig C, Takahashi JS, Chen BP. et al. Phosphorylation of the cryptochrome 1 C-terminal tail regulates circadian period length. The Journal of biological chemistry. 2013;288:35277-86
84. Yoshitane H, Takao T, Satomi Y, Du NH, Okano T, Fukada Y. Roles of CLOCK phosphorylation in suppression of E-box-dependent transcription. Molecular and cellular biology. 2009;29:3675-86
85. Spengler ML, Kuropatwinski KK, Schumer M, Antoch MP. A serine cluster mediates BMAL1-dependent CLOCK phosphorylation and degradation. Cell cycle (Georgetown, Tex). 2009;8:4138-46
86. Sahar S, Zocchi L, Kinoshita C, Borrelli E, Sassone-Corsi P. Regulation of BMAL1 protein stability and circadian function by GSK3beta-mediated phosphorylation. PloS one. 2010;5:e8561
87. Kwak Y, Jeong J, Lee S, Park YU, Lee SA, Han DH. et al. Cyclin-dependent kinase 5 (Cdk5) regulates the function of CLOCK protein by direct phosphorylation. The Journal of biological chemistry. 2013;288:36878-89
88. Bhadra U, Patra P, Pal-Bhadra M. Cardinal Epigenetic Role of non-coding Regulatory RNAs in Circadian Rhythm. Molecular neurobiology. 2017
89. Liu K, Wang R. MicroRNA-mediated regulation in the mammalian circadian rhythm. Journal of theoretical biology. 2012;304:103-10
90. Kojima S, Green CB. Circadian genomics reveal a role for post-transcriptional regulation in mammals. Biochemistry. 2015;54:124-33
91. Cheng HY, Obrietan K. Revealing a role of microRNAs in the regulation of the biological clock. Cell cycle (Georgetown, Tex). 2007;6:3034-5
92. Cheng HY, Papp JW, Varlamova O, Dziema H, Russell B, Curfman JP. et al. microRNA modulation of circadian-clock period and entrainment. Neuron. 2007;54:813-29
93. Sun E, Shi Y. MicroRNAs: Small molecules with big roles in neurodevelopment and diseases. Experimental neurology. 2015;268:46-53
94. Jagannath A, Butler R, Godinho SI, Couch Y, Brown LA, Vasudevan SR. et al. The CRTC1-SIK1 pathway regulates entrainment of the circadian clock. Cell. 2013;154:1100-11
95. Hansen KF, Sakamoto K, Obrietan K. MicroRNAs: a potential interface between the circadian clock and human health. Genome medicine. 2011;3:10
96. Shende VR, Goldrick MM, Ramani S, Earnest DJ. Expression and rhythmic modulation of circulating microRNAs targeting the clock gene Bmal1 in mice. PloS one. 2011;6:e22586
97. Shende VR, Kim SM, Neuendorff N, Earnest DJ. MicroRNAs function as cis- and trans-acting modulators of peripheral circadian clocks. FEBS letters. 2014;588:3015-22
98. Smith SS, Dole NS, Franceschetti T, Hrdlicka HC, Delany AM. MicroRNA-433 Dampens Glucocorticoid Receptor Signaling, Impacting Circadian Rhythm and Osteoblastic Gene Expression. The Journal of biological chemistry. 2016;291:21717-28
99. Curtis AM, Fagundes CT, Yang G, Palsson-McDermott EM, Wochal P, McGettrick AF. et al. Circadian control of innate immunity in macrophages by miR-155 targeting Bmal1. Proceedings of the National Academy of Sciences of the United States of America. 2015;112:7231-6
100. Han Y, Meng F, Venter J, Wu N, Wan Y, Standeford H. et al. miR-34a-dependent overexpression of Per1 decreases cholangiocarcinoma growth. Journal of hepatology. 2016;64:1295-304
101. Nagel R, Clijsters L, Agami R. The miRNA-192/194 cluster regulates the Period gene family and the circadian clock. The FEBS journal. 2009;276:5447-55
102. Lee KH, Kim SH, Lee HR, Kim W, Kim DY, Shin JC. et al. MicroRNA-185 oscillation controls circadian amplitude of mouse Cryptochrome 1 via translational regulation. Molecular biology of the cell. 2013;24:2248-55
103. Young TL, Matsuda T, Cepko CL. The noncoding RNA taurine upregulated gene 1 is required for differentiation of the murine retina. Current biology: CB. 2005;15:501-12
104. Cui M, Zheng M, Sun B, Wang Y, Ye L, Zhang X. A long noncoding RNA perturbs the circadian rhythm of hepatoma cells to facilitate hepatocarcinogenesis. Neoplasia (New York, NY). 2015;17:79-88
105. Johnstone RM, Adam M, Hammond JR, Orr L, Turbide C. Vesicle formation during reticulocyte maturation. Association of plasma membrane activities with released vesicles (exosomes). The Journal of biological chemistry. 1987;262:9412-20
106. van Niel G, D'Angelo G, Raposo G. Shedding light on the cell biology of extracellular vesicles. Nature reviews Molecular cell biology. 2018
107. Raposo G, Nijman HW, Stoorvogel W, Liejendekker R, Harding CV, Melief CJ. et al. B lymphocytes secrete antigen-presenting vesicles. The Journal of experimental medicine. 1996;183:1161-72
108. Marban E. The Secret Life of Exosomes: What Bees Can Teach Us About Next-Generation Therapeutics. Journal of the American College of Cardiology. 2018;71:193-200
109. Schorey JS, Cheng Y, Singh PP, Smith VL. Exosomes and other extracellular vesicles in host-pathogen interactions. EMBO reports. 2015;16:24-43
110. Robinson DG, Ding Y, Jiang L. Unconventional protein secretion in plants: a critical assessment. Protoplasma. 2016;253:31-43
111. Di Liegro CM, Schiera G, Di Liegro I. Extracellular Vesicle-Associated RNA as a Carrier of Epigenetic Information. Genes. 2017:8
112. Harding C, Heuser J, Stahl P. Receptor-mediated endocytosis of transferrin and recycling of the transferrin receptor in rat reticulocytes. The Journal of cell biology. 1983;97:329-39
113. Raposo G, Stoorvogel W. Extracellular vesicles: exosomes, microvesicles, and friends. The Journal of cell biology. 2013;200:373-83
114. Thery C, Amigorena S, Raposo G, Clayton A. Isolation and characterization of exosomes from cell culture supernatants and biological fluids. Current protocols in cell biology. 2006 Chapter 3: Unit 3.22
115. Tao SC, Guo SC, Zhang CQ. Platelet-derived Extracellular Vesicles: An Emerging Therapeutic Approach. International journal of biological sciences. 2017;13:828-34
116. Harding C, Heuser J, Stahl P. Endocytosis and intracellular processing of transferrin and colloidal gold-transferrin in rat reticulocytes: demonstration of a pathway for receptor shedding. European journal of cell biology. 1984;35:256-63
117. Pan BT, Teng K, Wu C, Adam M, Johnstone RM. Electron microscopic evidence for externalization of the transferrin receptor in vesicular form in sheep reticulocytes. The Journal of cell biology. 1985;101:942-8
118. Satta N, Toti F, Feugeas O, Bohbot A, Dachary-Prigent J, Eschwege V. et al. Monocyte vesiculation is a possible mechanism for dissemination of membrane-associated procoagulant activities and adhesion molecules after stimulation by lipopolysaccharide. Journal of immunology (Baltimore, Md: 1950). 1994;153:3245-55
119. Sims PJ, Faioni EM, Wiedmer T, Shattil SJ. Complement proteins C5b-9 cause release of membrane vesicles from the platelet surface that are enriched in the membrane receptor for coagulation factor Va and express prothrombinase activity. The Journal of biological chemistry. 1988;263:18205-12
120. Antonyak MA, Cerione RA. Microvesicles as mediators of intercellular communication in cancer. Methods in molecular biology (Clifton, NJ). 2014;1165:147-73
121. Gould SJ, Raposo G. As we wait: coping with an imperfect nomenclature for extracellular vesicles. Journal of extracellular vesicles. 2013:2
122. Kowal J, Arras G, Colombo M, Jouve M, Morath JP, Primdal-Bengtson B. et al. Proteomic comparison defines novel markers to characterize heterogeneous populations of extracellular vesicle subtypes. Proceedings of the National Academy of Sciences of the United States of America. 2016;113:E968-77
123. Tkach M, Thery C. Communication by Extracellular Vesicles: Where We Are and Where We Need to Go. Cell. 2016;164:1226-32
124. Tao SC, Guo SC, Zhang CQ. Modularized Extracellular Vesicles: The Dawn of Prospective Personalized and Precision Medicine. Advanced Science. 2018
125. Hu Y, Rao SS, Wang ZX, Cao J, Tan YJ, Luo J. et al. Exosomes from human umbilical cord blood accelerate cutaneous wound healing through miR-21-3p-mediated promotion of angiogenesis and fibroblast function. Theranostics. 2018;8:169-84
126. Tao SC, Rui BY, Wang QY, Zhou D, Zhang Y, Guo SC. Extracellular vesicle-mimetic nanovesicles transport LncRNA-H19 as competing endogenous RNA for the treatment of diabetic wounds. Drug delivery. 2018;25:241-55
127. Tao SC, Yuan T, Zhang YL, Yin WJ, Guo SC, Zhang CQ. Exosomes derived from miR-140-5p-overexpressing human synovial mesenchymal stem cells enhance cartilage tissue regeneration and prevent osteoarthritis of the knee in a rat model. Theranostics. 2017;7:180-95
128. Alvarez-Erviti L, Seow Y, Yin H, Betts C, Lakhal S, Wood MJ. Delivery of siRNA to the mouse brain by systemic injection of targeted exosomes. Nature biotechnology. 2011;29:341-5
129. Ratajczak J, Miekus K, Kucia M, Zhang J, Reca R, Dvorak P. et al. Embryonic stem cell-derived microvesicles reprogram hematopoietic progenitors: evidence for horizontal transfer of mRNA and protein delivery. Leukemia. 2006;20:847-56
130. Chen CY, Rao SS, Ren L, Hu XK, Tan YJ, Hu Y. et al. Exosomal DMBT1 from human urine-derived stem cells facilitates diabetic wound repair by promoting angiogenesis. Theranostics. 2018;8:1607-23
131. Al-Nedawi K, Meehan B, Kerbel RS, Allison AC, Rak J. Endothelial expression of autocrine VEGF upon the uptake of tumor-derived microvesicles containing oncogenic EGFR. Proceedings of the National Academy of Sciences of the United States of America. 2009;106:3794-9
132. Al-Nedawi K, Meehan B, Micallef J, Lhotak V, May L, Guha A. et al. Intercellular transfer of the oncogenic receptor EGFRvIII by microvesicles derived from tumour cells. Nature cell biology. 2008;10:619-24
133. Ciardiello C, Cavallini L, Spinelli C, Yang J, Reis-Sobreiro M, de Candia P. et al. Focus on Extracellular Vesicles: New Frontiers of Cell-to-Cell Communication in Cancer. International journal of molecular sciences. 2016;17:175
134. Peinado H, Lavotshkin S, Lyden D. The secreted factors responsible for pre-metastatic niche formation: old sayings and new thoughts. Seminars in cancer biology. 2011;21:139-46
135. Ratajczak J, Wysoczynski M, Hayek F, Janowska-Wieczorek A, Ratajczak MZ. Membrane-derived microvesicles: important and underappreciated mediators of cell-to-cell communication. Leukemia. 2006;20:1487-95
136. van der Pol E, Boing AN, Harrison P, Sturk A, Nieuwland R. Classification, functions, and clinical relevance of extracellular vesicles. Pharmacol Rev. 2012;64:676-705
137. Mahmoudi K, Ezrin A, Hadjipanayis C. Small extracellular vesicles as tumor biomarkers for glioblastoma. Molecular aspects of medicine. 2015;45:97-102
138. Xiang D, Shigdar S, Qiao G, Wang T, Kouzani AZ, Zhou SF. et al. Nucleic acid aptamer-guided cancer therapeutics and diagnostics: the next generation of cancer medicine. Theranostics. 2015;5:23-42
139. Vlassov AV, Magdaleno S, Setterquist R, Conrad R. Exosomes: current knowledge of their composition, biological functions, and diagnostic and therapeutic potentials. Biochimica et biophysica acta. 2012;1820:940-8
140. Barile L, Vassalli G. Exosomes: Therapy delivery tools and biomarkers of diseases. Pharmacology & therapeutics. 2017;174:63-78
141. Schorey JS, Harding CV. Extracellular vesicles and infectious diseases: new complexity to an old story. The Journal of clinical investigation. 2016;126:1181-9
142. Chong JJ, Yang X, Don CW, Minami E, Liu YW, Weyers JJ. et al. Human embryonic-stem-cell-derived cardiomyocytes regenerate non-human primate hearts. Nature. 2014;510:273-7
143. Hartman ME, Dai DF, Laflamme MA. Human pluripotent stem cells: Prospects and challenges as a source of cardiomyocytes for in vitro modeling and cell-based cardiac repair. Advanced drug delivery reviews. 2016;96:3-17
144. Adamiak M, Cheng G, Bobis-Wozowicz S, Zhao L, Kedracka-Krok S, Samanta A. et al. Induced Pluripotent Stem Cell (iPSC)-Derived Extracellular Vesicles Are Safer and More Effective for Cardiac Repair Than iPSCs. Circulation research. 2018;122:296-309
145. Akers JC, Ramakrishnan V, Yang I, Hua W, Mao Y, Carter BS. et al. Optimizing preservation of extracellular vesicular miRNAs derived from clinical cerebrospinal fluid. Cancer biomarkers: section A of Disease markers. 2016;17:125-32
146. Tao SC, Yuan T, Rui BY, Zhu ZZ, Guo SC, Zhang CQ. Exosomes derived from human platelet-rich plasma prevent apoptosis induced by glucocorticoid-associated endoplasmic reticulum stress in rat osteonecrosis of the femoral head via the Akt/Bad/Bcl-2 signal pathway. Theranostics. 2017;7:733-50
147. Guo SC, Tao SC, Yin WJ, Qi X, Yuan T, Zhang CQ. Exosomes derived from platelet-rich plasma promote the re-epithelization of chronic cutaneous wounds via activation of YAP in a diabetic rat model. Theranostics. 2017;7:81-96
148. Guo SC, Tao SC, Yin WJ, Qi X, Sheng JG, Zhang CQ. Exosomes from Human Synovial-Derived Mesenchymal Stem Cells Prevent Glucocorticoid-Induced Osteonecrosis of the Femoral Head in the Rat. International journal of biological sciences. 2016;12:1262-72
149. Tao SC, Guo SC, Li M, Ke QF, Guo YP, Zhang CQ. Chitosan Wound Dressings Incorporating Exosomes Derived from MicroRNA-126-Overexpressing Synovium Mesenchymal Stem Cells Provide Sustained Release of Exosomes and Heal Full-Thickness Skin Defects in a Diabetic Rat Model. Stem cells translational medicine. 2017;6:736-47
150. Zhang ZC, Tang C, Dong Y, Zhang J, Yuan T, Tao SC. et al. Targeting the long noncoding RNA MALAT1 blocks the pro-angiogenic effects of osteosarcoma and suppresses tumour growth. International journal of biological sciences. 2017;13:1398-408
151. Forterre A, Jalabert A, Chikh K, Pesenti S, Euthine V, Granjon A. et al. Myotube-derived exosomal miRNAs downregulate Sirtuin1 in myoblasts during muscle cell differentiation. Cell cycle (Georgetown, Tex). 2014;13:78-89
152. Shi XF, Wang H, Kong FX, Xu QQ, Xiao FJ, Yang YF. et al. Exosomal miR-486 regulates hypoxia-induced erythroid differentiation of erythroleukemia cells through targeting Sirt1. Experimental cell research. 2017;351:74-81
153. Arslan F, Lai RC, Smeets MB, Akeroyd L, Choo A, Aguor EN. et al. Mesenchymal stem cell-derived exosomes increase ATP levels, decrease oxidative stress and activate PI3K/Akt pathway to enhance myocardial viability and prevent adverse remodeling after myocardial ischemia/reperfusion injury. Stem cell research. 2013;10:301-12
154. Liu L, Jin X, Hu CF, Li R, Zhou Z, Shen CX. Exosomes Derived from Mesenchymal Stem Cells Rescue Myocardial Ischaemia/Reperfusion Injury by Inducing Cardiomyocyte Autophagy Via AMPK and Akt Pathways. Cellular physiology and biochemistry: international journal of experimental cellular physiology, biochemistry, and pharmacology. 2017;43:52-68
155. Zhang C, Xiao X, Chen M, Aldharee H, Chen Y, Long W. Liver kinase B1 restoration promotes exosome secretion and motility of lung cancer cells. Oncology reports. 2018;39:376-82
156. Liu F, Bu Z, Zhao F, Xiao D. Increased T-helper 17 cell differentiation mediated by exosome-mediated microRNA-451 redistribution in gastric cancer infiltrated T cells. Cancer science. 2018;109:65-73
157. Zhu LL, Huang X, Yu W, Chen H, Chen Y, Dai YT. Transplantation of adipose tissue-derived stem cell-derived exosomes ameliorates erectile function in diabetic rats. Andrologia. 2017
158. Barile L, Lionetti V, Cervio E, Matteucci M, Gherghiceanu M, Popescu LM. et al. Extracellular vesicles from human cardiac progenitor cells inhibit cardiomyocyte apoptosis and improve cardiac function after myocardial infarction. Cardiovascular research. 2014;103:530-41
159. Cleys ER, Halleran JL, McWhorter E, Hergenreder J, Enriquez VA, da Silveira JC. et al. Identification of microRNAs in exosomes isolated from serum and umbilical cord blood, as well as placentomes of gestational day 90 pregnant sheep. Molecular reproduction and development. 2014;81:983-93
160. Cervio E, Barile L, Moccetti T, Vassalli G. Exosomes for Intramyocardial Intercellular Communication. Stem cells international. 2015;2015:482171
161. Geiger A, Walker A, Nissen E. Human fibrocyte-derived exosomes accelerate wound healing in genetically diabetic mice. Biochemical and biophysical research communications. 2015;467:303-9
162. Xu B, Zhang Y, Du XF, Li J, Zi HX, Bu JW. et al. Neurons secrete miR-132-containing exosomes to regulate brain vascular integrity. Cell research. 2017;27:882-97
163. Pusic AD, Kraig RP. Youth and environmental enrichment generate serum exosomes containing miR-219 that promote CNS myelination. Glia. 2014;62:284-99
164. Kim DK, Lee J, Kim SR, Choi DS, Yoon YJ, Kim JH. et al. EVpedia: a community web portal for extracellular vesicles research. Bioinformatics (Oxford, England). 2015;31:933-9
165. Choi DS, Kim DK, Kim YK, Gho YS. Proteomics, transcriptomics and lipidomics of exosomes and ectosomes. Proteomics. 2013;13:1554-71
166. Kim DK, Kang B, Kim OY, Choi DS, Lee J, Kim SR. et al. EVpedia: an integrated database of high-throughput data for systemic analyses of extracellular vesicles. Journal of extracellular vesicles. 2013:2
167. Choi DS, Kim DK, Kim YK, Gho YS. Proteomics of extracellular vesicles: Exosomes and ectosomes. Mass spectrometry reviews. 2015;34:474-90
168. Kim DK, Lee J, Simpson RJ, Lotvall J, Gho YS. EVpedia: A community web resource for prokaryotic and eukaryotic extracellular vesicles research. Seminars in cell & developmental biology. 2015;40:4-7
169. Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome research. 2003;13:2498-504
170. Eguchi A, Lazaro RG, Wang J, Kim J, Povero D, Willliams B. et al. Extracellular vesicles released by hepatocytes from gastric infusion model of alcoholic liver disease contain a MicroRNA barcode that can be detected in blood. Hepatology (Baltimore, Md). 2017;65:475-90
171. Warde-Farley D, Donaldson SL, Comes O, Zuberi K, Badrawi R, Chao P. et al. The GeneMANIA prediction server: biological network integration for gene prioritization and predicting gene function. Nucleic acids research. 2010;38:W214-20
172. Negash AA, Gale M Jr. Hepatitis regulation by the inflammasome signaling pathway. Immunological reviews. 2015;265:143-55
173. Zappulli V, Friis KP, Fitzpatrick Z, Maguire CA, Breakefield XO. Extracellular vesicles and intercellular communication within the nervous system. The Journal of clinical investigation. 2016;126:1198-207
174. Ghidoni R, Paterlini A, Albertini V, Glionna M, Monti E, Schiaffonati L. et al. Cystatin C is released in association with exosomes: a new tool of neuronal communication which is unbalanced in Alzheimer's disease. Neurobiology of aging. 2011;32:1435-42
175. Chivet M, Javalet C, Hemming F, Pernet-Gallay K, Laulagnier K, Fraboulet S. et al. Exosomes as a novel way of interneuronal communication. Biochemical Society transactions. 2013;41:241-4
176. Edelstein L, Smythies J. The role of epigenetic-related codes in neurocomputation: dynamic hardware in the brain. Philosophical transactions of the Royal Society of London Series B, Biological sciences. 2014:369
177. Korkut C, Li Y, Koles K, Brewer C, Ashley J, Yoshihara M. et al. Regulation of postsynaptic retrograde signaling by presynaptic exosome release. Neuron. 2013;77:1039-46
178. Goldie BJ, Dun MD, Lin M, Smith ND, Verrills NM, Dayas CV. et al. Activity-associated miRNA are packaged in Map1b-enriched exosomes released from depolarized neurons. Nucleic acids research. 2014;42:9195-208
Author contact
Corresponding authors: Shang-Chun Guo, Institute of Microsurgery on Extremities, Shanghai Jiao Tong University Affiliated Sixth People's Hospital, Shanghai 200233, China. E-mail: achunicom and Shi-Cong Tao, Department of Orthopaedic Surgery, Shanghai Jiao Tong University Affiliated Sixth People's Hospital, 600 Yishan Road, Shanghai 200233, China. E-mail: jerrytao1990com

 Global reach, higher impact
Global reach, higher impact