Impact Factor ISSN: 1449-2288
Int J Biol Sci 2021; 17(4):1125-1137. doi:10.7150/ijbs.55724 This issue Cite
Review
Paradoxical HBsAg and anti-HBs coexistence among Chronic HBV Infections: Causes and Consequences
1. National Center for Clinical Laboratories, Beijing Hospital, National Center of Gerontology; Institute of Geriatric Medicine, Chinese Academy of Medical Sciences, Beijing, P.R. China.
2. Beijing Engineering Research Center of Laboratory Medicine, Beijing Hospital, P.R. China.
3. Graduate School, Peking Union Medical College, Chinese Academy of Medical Sciences, Beijing, P.R. China.
Received 2020-11-10; Accepted 2021-3-11; Published 2021-3-11
Abstract
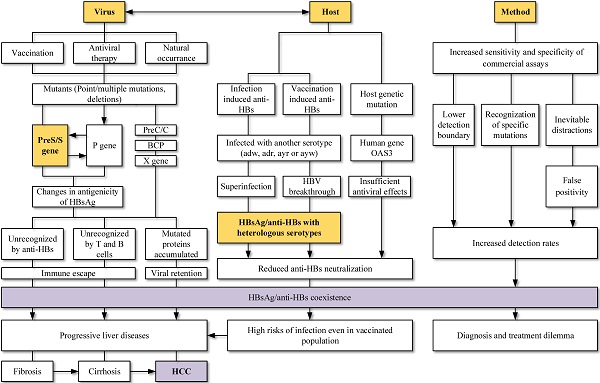
Hepatitis B surface antigen (HBsAg) and Hepatitis B surface antibody (anti-HBs) were reported simultaneously among Hepatitis B virus (HBV) infections. HBsAg is a specific indicator of acute or chronic HBV infections, while anti-HBs is a protective antibody reflecting the recovery and immunity of hosts. HBsAg and anti-HBs coexist during seroconversion and then form immune complex, which is rare detected in clinical cases. However, with the promotion of vaccination and the application of various antiviral drugs, along with the rapid development of medical technology, the coexistence of HBsAg and anti-HBs has become more prevalent. Mutations in the viral genomes, immune status and genetic factors of hosts may contribute to the coexistence. Novel HBsAg assays, with higher sensitivity and ability to detect mutations or immune complexes, can also yield HBsAg/anti-HBs coexistence. The discovery of coexistence has shattered the idea of traditional serological patterns and raised questions about the effectiveness of vaccines. Worth noting is that HBsAg/anti-HBs double positivity is strongly associated with progressive liver diseases, especially hepatocellular carcinoma. In conclusion, viral mutations, host factors, and methodology impacts can all lead to the coexistence of HBsAg and anti-HBs. This coexistence is not an indicator of improvement, as an increased risk of adverse clinical outcomes still exists.
Keywords: Hepatitis B surface antibody, Hepatitis B surface antigen, Mutation, Hepatocellular carcinoma
Introduction
Hepatitis B virus (HBV) infection is an enormous public health problem, seriously threatening a large number of undeveloped countries worldwide. In 2015, chronic hepatitis B infection (CHB) affected 257 million persons globally (3.5% of the world population) [1]. Based on an epidemiology survey [2], China was suffering from the heaviest burden of HBV with 93 million HBsAg carriers, of which approximately 20 million were CHB patients. It was also estimated that nearly one million people died of HBV-related liver diseases every year with half coming from China [3].
During the infection, antibody response plays an important role in eliminating HBV particles and infected liver cells. Among the infected population, patients may naturally eliminate HBsAg and generate hepatitis B surface antibody (anti-HBs), while others may progress from chronic hepatitis to cirrhosis and hepatocellular carcinoma (HCC) [4, 5]. Hepatitis B surface antigen (HBsAg) is the most important marker for the diagnosis of HBV infection. Anti-HBs is a specific protective antibody produced by the stimulation of exposed antigenic determinant, which can neutralize HBsAg in serum. The widely accepted goal of HBV therapy is seroconversion from HBsAg to anti-HBs. The dynamic transition may be the reason for the rare detection of HBsAg/anti-HBs coexistence. However, cases of simultaneous HBsAg and anti-HBs were frequently reported and difficult to be explained by the regular serological patterns. Arnold [6] first reported the concurrent anti-HBs and HBsAg with different subtypes both in serologic and fluorescence histologic studies. Though it has been over 40 years since they discovered this serological pattern, the underlying molecular mechanisms are still unclear and whether the coexistence is beneficial or detrimental to the recovery remained to be elucidated. In this review, we discuss the possible causes and clinical significance of the paradoxical HBsAg/anti-HBs coexistence and provide insights for unusual serological patterns.
Mechanisms of coexistence
Mutations in the viral genome
Using high-affinity monoclonal anti-HBs against separate determinants on HBsAg revealed the antigenic heterogeneity of HBV [7]. Accordingly, antiviral therapy, vaccination, and natural immune pressure [8, 9] could account for these mutations.
Mutations are generally not limited to specific open reading frames (ORFs) and take place in all viral genes and regulatory elements [10]. However, they tend to cluster together in specific regions, like the preS/S gene, reverse transcriptase region (RT) in the polymerase gene, the pre-core region, basal core promoter (BCP), and the X gene.
Mutations in the preS/S gene
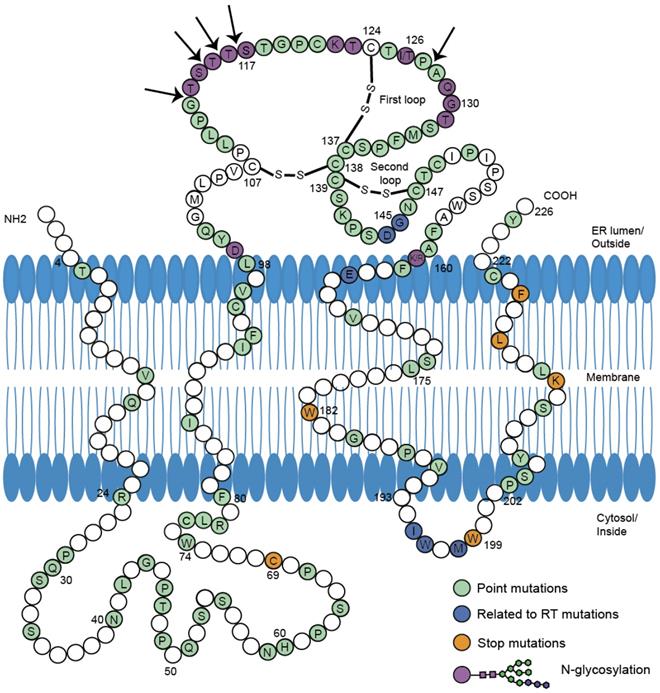
Point mutations in the preS/S gene have been studied extensively in coexisting patterns (Figure 1). The major hydrophilic region (MHR) of HBsAg comprises the main conformational epitopes of viral particles [11]. According to functional studies in vivo and in vitro, variations in the MHR can alter the antigenicity and immunogenicity of HBsAg, and diminish the reaction of anti-HBs [12, 13] (Table 1). Lada [14] found that mutation rates during coexistence group were 2.7-fold higher than during HBsAg alone. The rate of aa variability in the 'α' determinant of the MHR was also higher during coexistence (9.5% vs. 2.4%, p=0.009) than during HBsAg alone. In the first loop of 'α' determinant, mutations occurred in more positions in genotype C (Table S1). Another study found that the mutation rate at 126 aa (I126S/T/V) during coexistence was the highest and 6-fold higher [15]. The high prevalence of I126 mutations in genotype C was consistent with what other studies found [16, 17].
Mutations in S protein associated with HBsAg/anti-HBs coexistence. 226 amino acids of HBV small S protein were demonstrated in wild-type sequence with specific colors indicating different mutations. The conformation of the two major loops of the 'a' determinant is dependent on the presence of disulfide bonds (-S-S-). The proposed four transmembrane domains (TMD) and two cytosol loops were shown in the picture. The amino acid sequence of HBV S protein was retrieved from GenBank. The division of specific domains (TMD1:4-24 aa, TMD2:80-98 aa, TMD3:160-193 aa, TMD4:202-222 aa, MHR:99-160 aa) was referred to previous report[11]. According to the reference sequence ABE01542 (Genotype C, isolate S05014765), positions in white were not reported to mutate in HBsAg/anti-HBs coexistence. Positions in green were reported to occur in concurrent HBsAg/anti-HBs situation individually or jointly. Positions in dark blue were in connection with mutations in reverse transcriptase region of polymerase gene. Stop mutations were in orange and N-glycosylation in purple. Arrows were indicating insertions into amino acid sequence. All the mutations were listed in the Table S1.
Mutations associated with HBsAg/anti-HBs coexistence
| Regions | Mutation patterns | Possible mechanisms of coexistence | Representative mutations | ||
|---|---|---|---|---|---|
| More frequently/only found in HBsAg/anti-HBs coexistence | Associated with progressive liver diseases | ||||
| S gene (encoding small HBsAg) | MHR (99-160 aa) | Point or double mutations (including glycosylation) | Alter immunogenicity of HBsAg and reduce reactivity for anti-HBs induced by the recombinant HBV vaccine | Genotype B: sG145R [17] Genotype C: sI126S/T [15] | Genotype B: sT131N+sM133T Genotype C: sT116N, sQ129N, sT131N, sG130N+sT131N, sG130N+sT131I [22, 23] |
| Inserted mutations or deletions | Change the antigenicity of HBsAg in a greater extent | Genotype B: s126-127 “SARIVNTT” insertion Genotype C: s112-113 “KNA” insertion, s114-115 “NTSTT” insertion, s115-116 “INGTST” insertions, s116-117 “STGR” insertions [22, 24] | Genotype C: s112-113 “KNA” insertion, s114-115 “TTN”/”NTSTT” insertion, s115-116 “INGTST” insertions; sT113N+ deletion in 114-116 aa [22] | ||
| Outside of MHR | Point mutation and stop mutations | Mutations in cytotoxic lymphocyte epitope affect the recognition of immune cells and affect binding to anti-HBs; Early stop mutation in the transmembrane domain influence the stability and secretion of HBsAg, affect the structure of virus-like particles and the binding of anti-HBs | sC69stop, sW182stop [17] | Genotype C: sW182stop [25] | |
| preS/S gene (encoding large and middle HBsAg) | preS1 (1-119 aa) 4 epitopes for T cells; 9 epitopes for B cells | Deletions and stop mutations | Interrupt viral surface epitopes and evade immune-surveillance; Mutations are unable to be recognized by immune cells | Genotype B: Deletion in 98-106 aa Genotype C: Deletion in 57-117 aa, 66-112 aa, 107-113 aa[21, 26] | |
| preS2 (120-174 aa) 2 epitopes for T cells; 1 epitope for B cells | Deletions and stop mutations | Mutation at the epitope for T and B cells influence the recognition of immune cells; Alter the expression and secretion of HBV envelope proteins and result in intracellular accumulation of mutant proteins | Genotype C: Deletion in 125-135 aa, 122-139 aa, 125-138 aa, 132-141 aa[21, 27] | ||
| Initiation codon mutations | Affect the secretion of the virus and cause the accumulation of viral proteins in host cells; Mutant proteins are unable to be recognized by immune cells | Genotype B: M120I/T/V[24, 28] | |||
| P gene (Polymerase) | RT region (1-344 aa) | Point or multiple mutations | Corresponding S gene mutants possess antigenically distinct HBsAg proteins which decrease HBsAg antigenicity and impair the neutralization ability of the antibodies | Genotype A: rtI16T Genotype B: rtR153H Genotype D: rtD134E, rtS135T/Y/N, rtR138N/K, rtN139Q, rtR/W153Q/K[29, 30] | Genotype B: rtR153Q, rtA181T Genotype C: rtR153Q, rtA181T, rtM204V, rtL180M+rtM204V, rtV173L+rtL180M+rtM204V [31-33] |
| BCP (1742-1849 nt) and X gene (1374-1838 nt) | Mutations in nucleotide level | Affect binding of BCP to liver-specific transcription factors | Genotype B and C: A1762T+G1764A [24, 34] | Genotype B and C: A1762T+G1764A [35] Genotype C: G1613A, C1653T, T1674C, T1753 V, G1764A [36] | |
| Mutations in amino acid level (HBxAg) | Mutant HBxAg may not stimulate CTL activity and favor the clearance of HBsAg | K130M+V131I [24, 34] | K130M+V131I [36] | ||
| preC/C (1814-2452 nt) | Point or double mutation | Mechanism remained exploration but the related mutations are classical risk factors for HBV-related HCC | Genotype C: G1896A (W28*) [34] | Genotype B: G1896A+G1899A; Genotype C: A1846T [36] | |
Abbreviations: HBsAg: hepatitis B surface antigen; anti-HBs: antibodies against HBsAg; MHR: major hydrophilic region; RT: reverse transcriptase; BCP: basal core promoter; HBxAg: hepatitis B x antigen; HCC: hepatocellular carcinoma; nt: nucleotide; aa: amino acid; CTL: cytotoxic lymphocyte.
In the second loop, mutations existed in all the positions. The G145R mutation was widely accepted as the most common immune-escape mutation [18]. Variants with G145R could dramatically decrease the affinity for anti-HBs [19] and increase the compactness and stability of HBsAg by enhancing the rigidity of the 'a' determinant [20]. However, G145R's detection varied in different genotypes. In genotype C, G145R during coexistence or G145A during HBsAg alone was rarely detected [15, 21]. Among patients with genotype B, G145R occurred in coexistence group but was not detected in HBsAg alone group [17]. G145R in genotype B and I126 mutations in genotype C can thus be the representative MHR substitutions in HBsAg/anti-HBs coexistence.
Glycosylation, insertion and early termination in S proteins influence the expression, secretion and recognition of HBsAg [29, 37]. Glycosylation reduces the affinity between anti-HBs and HBsAg by decreasing HBsAg antigenicity and impairing the neutralization ability of anti-HBs. N-glycosylation occurred more frequently during coexistence than during HBsAg alone (47/216 vs. 1/182, p<0.001) [23]. Position 129-131 aa displayed highest N-glycosylation rates. N-glycosylation insertions clustered around position 112 to 116 aa and some large insertion patterns shown in Table S1 were first reported during coexistence [22, 34, 38]. The insertions may affect biological properties and antigenicity of HBsAg to a greater extent. Stop mutations W182* and W199* [24] have been reported in the transmembrane domain (TMD) near the C-terminal of S proteins. Stop codons and the resulting early termination in HBsAg lacked the major epitopes recognized by specific antibodies, which could favor persistent viral replication and prolonged mutant HBsAg existence in the presence of anti-HBs.
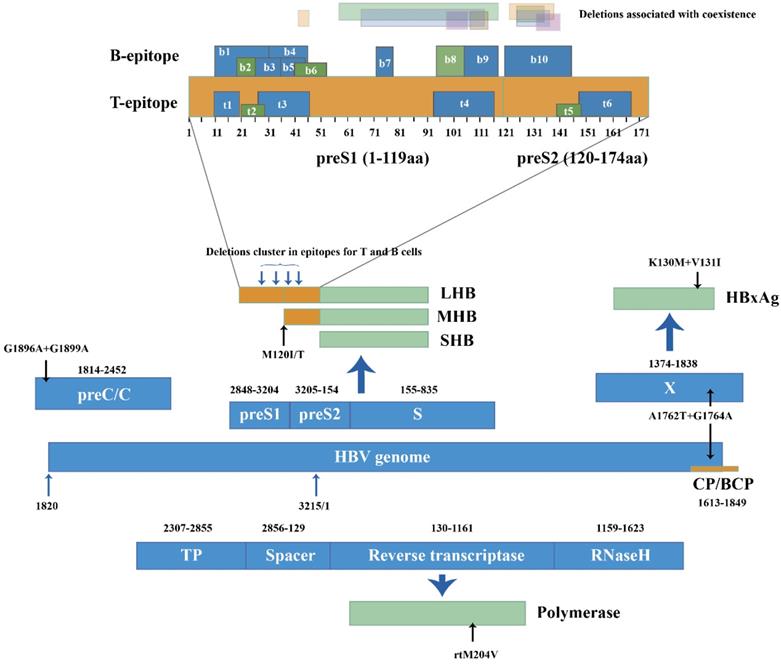
Mutations and deletions in the epitopes for T and B cells can also explain the coexistence situation. In genotype C, mutations I92T, F93C and C95W occurred in cytotoxic lymphocyte (CTL) epitope of S proteins and led to higher aa variation compared to HBsAg alone group (p=0.002) [34]. Huang [21] reported coexistence group yielded more deletions than HBsAg alone group at the T and B cell epitopes of the preS gene (25.0% vs. 3.6%). The deletions clustered within the C-terminal of the proteins encoded by preS1. Deletions specific to preS2 were located at the N-terminal of preS2 proteins, covering two epitopes for T cells and one epitope for B cells [27, 39] (Figure 2). Moreover, preS2 deletions were only found in coexistence group from genotype C-infected patients [21]. The mutations and deletions appear at C-terminal of preS1 proteins and N-terminal of preS2 proteins, impairing the recognition sites for immune cells and preventing CTL activation. Because the immune system produces few or even no active anti-HBs, immune-escape mutants survive and coexist with anti-HBs.
HBV genome (1-3215 nucleotide, nt) was represented in linear diagram. Different open reading frames and their encoded viral proteins were connected with bold blue-filled arrows. Light black arrows pointed out specific mutations in nucleotide level and amino acid level, respectively. PreS/S gene encoded three products: Large-HBsAg, Middle HBsAg and Small HBsAg (LHB, MHB and SHB). B and T epitopes in the preS products were numbered from N to C terminal. Translucent bars in different colors represented preS1 and pre S2 deletion patterns in HBsAg/anti-HBs coexistence.
Mutations in the transcriptase region
Studies on coexistence due to polymerase gene mutations are limited and are mainly focused on the RT region. Considering the overlapping structure of the S gene and RT region, mutations within S gene may account for changes in the P genes [40]. A significantly higher aa variability was observed within RT regions overlapping the MHR when compared between coexistence group and the HBsAg alone group (1.14 vs. 0.17, p=0.003), and mutations rtI16T, rtD134E, rtS135T/Y/N, rtR138N/K, rtN139Q and rtR/W153Q/K/H in genotype A, B or D were significantly more frequent in the coexistence group (p<0.05) [29, 31]. These mutations were harboring at the functional finger subdomain of the enzyme [41]. Other than rtI16T, the other mutations had corresponding HBsAg mutants located in the 'α' determinant. The dual mutations rtR153Q/sG145R, which occur in separate genes can inhibit HBV replication by affecting polymerase activity [42] and greatly decrease the neutralizing ability of anti-HBs by altering the immunogenicity of HBsAg.
Antiviral treatments may stimulate immune selection pressure and induce viral mutation acceleration. rtL180M and rtM204V appeared in a patient who was undergoing anti-HBV therapy when HBsAg/anti-HBs coexistence was first detected [29]. Several RT mutations corresponding to HBsAg mutants including sE164D, sI195M, sW196S, sM198I and sE164D+sI195M were also selected during lamivudine treatment and existed in coexistence group [33], raising the possibility that therapy-resistant mutants possess antigenically distinct HBsAg proteins that cannot be neutralized by anti-HBs. Another vaccine escape mutation that occurred in coexistence was rtA181T/sW172*, which had a dominant negative secretion effect and led to clearance of wildtype HBV antigen from the serum [43]. This mutation may help vaccine escape mutants coexist with the anti-HBs induced by vaccine [44, 45]. Although treatment-induced mutations in RT regions did not display high frequency in coexistence group, their concurrence raised questions about the effectiveness of antiviral drugs and vaccines.
Mutations overlapping the preC/BCP and X genes
BCP and pre-core mutations are prevalent in HBV chronic infection and associated with the severity of liver diseases [46]. During coexistence, point mutations G1742A, T1753C, A1755C, T1768A, C1773T, A1775G, T1802C and A1846T occur in the BCP region while G1896A and G1899A occur in the pre-core at the nucleotide level [24]. The BCP double mutations A1762T and G1764A are found more frequently in coexistence patients than in HBsAg alone patients (p=0.012) [34]. The dual mutations can enhance viral genome replication and may partially resist neutralization of anti-HBs [47]. A1762T+G1764A at the nucleotide level can also lead to K130M+V131I at the HBxAg coded by the X gene [36], which is CTL's target and can mediate the immune recognition of CTL. Mutations and early termination in HBxAg may not stimulate CTL activity and can slow down the clearance of HBsAg, thus generating uncommon serological patterns. Mutations in the preC/BCP and X region associated with coexistence were reported in limited samples. To investigate coexistence and advanced-disease-related mutants, more studies with larger sample sizes must be carried out.
Immune status and genetic factors of hosts
Adverse immune status of hosts can lead to superinfection of different HBV strains when no mutation exists in the HBV genome. Breakthrough or reactivation of HBV may occur and facilitate the heterologous HBsAg/anti-HBs coexistence [18, 29, 48].
HBsAg subtype determinants had diverse physical and chemical properties, and infection by one subtype generated incomplete cross-immunity, favoring the mixed infection of different subtypes. Thus, heterologous subtype-specific antibody in coexisted serological pattern is commonly produced through the infection with different HBV serotypes. Purcell [49] vaccinated several seronegative chimpanzees with prepared antigen of subtypes ayw or adr, and infected them with the ayw subtype HBV six months later. The chimpanzees inoculated with the same subtypes did not show evidence of developing HBsAg or hepatitis. However, one of the chimpanzees vaccinated with adr subtype was infected, indicating the absence of complete protection between subtypes. Japanese scientists then discovered HBsAg and anti-HBs of different serotypes in chronic HBV carriers [50]. Reinfection with a second subtype may be the reason as anti-HB against a different subtype than that of the circulating HBsAg was found [51]. In a neutralization experiment, 74.3% (26/35) anti-HBs in coexistence patients could not neutralize the HBsAg in their serum and 28.6% (10/35) of HBsAg in the same group could not be fully neutralized by the anti-HBs from vaccinated persons [17]. Moreover, antibodies from four patients could not neutralize the three major HBV serotypes (adw, adr and ayw) in China [52]. These experiments are strong evidence that HBsAg and anti-HBs develop from different serotypes and the coexistence pose an underlying threat to general populations, even to vaccinated persons.
HBV reactivation associated with immunosuppressive conditions is another possible mechanism [53]. The concurrent detection of HBsAg and anti-HBs is a possible signal of HBV reactivation, which occurs mainly in HBV resolved patients who received chemotherapy or immunosuppressive therapy without antiviral prophylaxis. Patients with resolved HBV infection often have negative HBsAg, positive hepatitis B core antibody (anti-HBc) and protective anti-HBs. Wang [48] emphasized that active viral replication or occult infection still occurs in these patients, and anti-HBs offers no protection against the virus. In patients with HBV reactivation, viral replication would accelerate again to produce a large amount of HBsAg in a short time, leading to the coexistence situation and therapy-related HBV reactivation. The reactivation may finally lead to lethal consequences such as fulminant hepatitis or hepatic failure [54].
Host genetic factors could play a role in the clearance of HBV and influence the occurrence of anti-HBs in chronic HBsAg carriers [55]. The human gene oligoadenylate synthetases 3 (OAS3) may degrade viral RNAs and restrict the HBV replication, which was related to HBsAg/anti-HBs coexistence through a burden study. The overexpression of mutated OAS led to inactivation of IFN-induced OAS/RNase L pathways; and the resulting insufficient antiviral effects may cause failed clearance of HBsAg. Although further functional study of OAS3 gene is needed to evaluate its contribution to the coexistence group, host factors' influence on antiviral responses cannot be overlooked.
Impact of methodology
The improvement of diagnosis methods rather than faulty methodology in HBsAg or anti-HBs detection could primarily account for the coexistence [34]. Although HBV genetic variability poses threats against the current assays, in-depth serological and molecular researches contribute to the development of the reagents. Over the years, the sensitivity and specificity of assays have greatly improved. Huzly [56] compared nine commercial assays for anti-HBs quantification. An accurate measurement for a series of diluted international standard was achieved by most of the systems, including both enzyme immunoassays (EIAs) and chemiluminescent immunoassays (CIAs). After evaluating and comparing the performance of available anti-HBs reagents [57], new assay formats were designed to optimize their specificity [58].
Qualitative assays for HBsAg had the ability to detect classical mutations in the MHR [59], including point mutations at positions 123, 129, 133, 144, 145 and P142L/S+G145R double mutations. Lou [60] reported an ultra-sensitive Architect assay for the detection of HBsAg, which can detect up to 97 designed mutations covering the key positions in S proteins. From qualitative detection to quantitative determination, the limit of HBsAg reached the boundary of 50 mIU/ml [61]. Pancher [62] demonstrated favorable overall performance of quantitative assays for HBsAg from coexistence patients and that the circulating anti-HBs did not influence HBsAg quantification. Moreover, in sandwich CIAs applying both polyclonal and monoclonal antibodies, no specific protein pattern can impair the quantification which was supported by all the assays [63].
Development of highly‐sensitive assays was a trend for future HBsAg quantitation. A novel chemiluminescent enzyme immunoassay (CLEIA) was designed for quantitative HBsAg, which used monoclonal antibodies targeting both common 'a' determinant and loop inside the lipid bilayer [64]. The prototype of the two-step sandwich assay was further applied on Lumipulse system (Fujirebio Inc). The sensitivity of Lumipulse HBsAg-HQ was 5 mIU/ml, approximately 10-fold higher than previously reported assays. By disrupting HBV particles, dissociating HBsAg from HBsAg/anti-HBs complexes and denaturing epitopes to a linear form, thus fully exposing the outer and inner epitopes of HBsAg. As a result, in 26 HBsAg seronegative patients, Lumipulse HBsAg-HQ assay detected HBsAg in 4 patients with a protective anti-HBs concentration (over 10 mIU/ml), 3 of whom had no HBsAg escape mutations [65]. The highly sensitive CLEIA Lumipulse HBsAg-HQ is suitable for HBV monitoring, which may raise the questions of vaccine effectiveness and disrupt the awareness of traditional serological patterns.
The false positive of serological markers caused by sample storage, specimen handling, as well as reagent usage should also be dealt with [53], as the detection can be influenced by contamination, interfering substances and the loading amount of plasma [66]. The results of different studies need cautious analysis and comparisons [67, 68]. The identification of the coexistence group in clinical experiment needs re-examination and the use of reliable reagents is essential to reduce errors caused by reagents and operating process. Detection of HBV serum markers can be combined with HBV DNA, liver function and other clinical symptoms. A dynamic and comprehensive analysis can provide a dependable basis for clinical diagnosis and treatment.
Clinical characteristics and significances in coexistence
Clinical characteristics and advanced liver diseases in coexistence
The prevalence of simultaneous anti-HBs and HBsAg positivity was found in 2.4%-5.8% of chronic HBV infections in mainland China from 2007-2019 (Table 2). However, the higher prevalence rates (21%) were obtained in Japan and Singapore in 1996, which might overestimate the occurrence of concurrent HBsAg/anti-HBs due to limited sample size and inclusion of acute HBV infection.
Prevalence rate, assays and characteristics in HBsAg/anti-HBs coexistence
| Prevalence rate | Assays for HBsAg | Assays for anti-HBs | Genotypes in the study | Particular characteristics (compare with HBsAg positive alone group) | Countries | Year of publication | |
|---|---|---|---|---|---|---|---|
| 2.4% | 39/1606 | Architect quantification assays (CMIA, Abbott Laboratories) 1 | Genotype B and C | None | China, Wuhan | 2015 [74] | |
| 2.6% | 145/5513 | Quantification on Architect i2000 system (CMIA, Abbott Laboratories) 1 and LIAISON-XL quantification assays (CLIA, Diasorin) 2 | Genotype B and C | Lower proportion of patients with HBsAg >250 IU/ml; Lower proportion of patients with HBV DNA > 104 IU/ml | China, Fujian | 2017 [17] | |
| 2.9% | 122/4169 | Elecsys System (ECLIA, Roche Diagnostics) | Genotype B, C and D | Higher genotype D proportion | China, Gansu | 2016 [75] | |
| 2.9% | 953/32467 | Architect quantification assays (CMIA, Abbott Laboratories) 1 | Genotype B and C | None | China, Shanghai | 2014 [23] | |
| 2.9% | 54/1862 | AxSYM assay (MEIA, Abbott Diagnostics) | Genotype B and C | Higher anti-HBe positive rate and genotype C proportion | China, Shanghai | 2012 [30] | |
| 3.0% | 36/1194 | Architect quantification assays on Architect i2000 system (CMIA, Abbott Laboratories) 1 | Genotype C | Higher anti-HBs level | China, Wuhan | 2016 [31] | |
| 3.3% | 436/13080 | Architect quantification assays on Architect i2000 system (CMIA, Abbott Laboratories) 1 | Genotype B and C | Lower proportion of patients with HBsAg >250 IU/ml | China, Jiangsu | 2016 [16] | |
| 3.4% | 34/1000 | Architect quantification assays on Architect i2000 system (CMIA, Abbott Laboratories) 1 | Genotype B and C | None | China, Shanghai | 2010 [21] | |
| 3.6% | 72/1985 | Architect quantification assays on Architect i2000 system (CMIA, Abbott Laboratories) 1 | Genotype B and C | Lower HBsAg level and HBV DNA concentration | China, Zhejiang | 2011 [34] | |
| 4.0% | 179/4455 | Architect quantification assays on Architect i2000 system (CMIA, Abbott Laboratories) 1 | Genotype B and C | None | China, Guangzhou | 2018 [38] | |
| 4.9% | 20/411 | Commercial EIAs by Abbott Laboratories, Roche Diagnostics and Dade Behring | Genotype B2 and C1 | Lower serum HBV DNA concentrations | China, Shanghai, Beijing, Guangzhou, Changchun | 2007 [18] | |
| 5.8% | 505/8687 | Cobas E601 quantitative electrochemical luminescence assays (ECLIA, Roche Diagnostics) 3 | Genotype B and C | Higher genotype C proportion | China, Tianjin | 2019 [76] | |
| 1.2% | 18/1462 | Elecsys system (ECLIA, Roche Diagnostics) | Enzyme-linked immunosorbent assay (ELISA, Diasorin Inc) | Genotype A2, B, C and E | Older median age; lower platelet counts; higher prevalence of HBeAg; lower HBsAg levels | North America | 2020 [77] |
| 2.8% | 13/459 | AxSYM assay (MEIA, Abbott Laboratories) | Genotype A, B, D, E and A/D | HBV genotypes D was preponderant; HBV DNA levels were significantly higher | France | 2007 [29] | |
| 2.9% | 353/12191 | Elecsys system (ECLIA, Roche Diagnostics), ADVIA Centaur qualitative assays (CLIA, Bayer Diagnostics), and AxSYM assay (MEIA, Abbott Laboratories) | None | Subjects > 50 years old was higher; Higher AST and ALT levels | Korea | 2013 [78] | |
| 3.3% | 3/92 | Enzyme-linked immunosorbent assay (ELISA, Diasorin) | Genotype D | None | Northeastern Brazil | 2017 [79] | |
| 5.0% | 129/2578 | Quantification on Architect i2000 system (CMIA, Abbott Laboratories) 1, Elecsys system (ECLIA, Roche Diagnostics) and LIAISON-XL quantification assays (CLIA, DiaSorin) 2 | Genotype B~E and A/F | Lower anti-HBs concentration (<50 IU/L) | France | 2015 [80] | |
| 6.4% | 48/755 | Commercial kits (EIA, Abbott Laboratories) | Genotype C | HBeAg positive rate was higher | South Korea | 2009 [72] | |
| 7.0% | 73/1042 | Commercial assay kits (Abbot Laboratories) | Genotype C | Older median age | South Korea | 2014 [71] | |
| 7.1% | 166/2341 | Quantification on Architect i2000 system (CMIA, Abbott Laboratories) 1 | Genotype C | Lower proportion of patients with HBsAg >250 IU/ml | South Korea | 2019 [53] | |
| 8.9% | 77/866 | AxSYM assay (MEIA, Abbott Laboratories) | Genotype A~E | Higher anti-HBs levels | France | 2006 [14] | |
| 20.5% | 15/73 | Radioimmunoassay AUSRIAII-I25 and AUSAB (Abbott Laboratories) | None | Higher serum titers of anti-HBc | Japan | 1996 [81] | |
| 21.0% | 234/1132 | Auszyme II (EIA, Abbott Laboratories) | AxSYM AUSAB (MEIA, Abbott Laboratories) | None | None | Singapore | 1996 [48] |
1 Architect HBsAg and anti-HBs quantification assays (CMIA, Abbott Laboratories): critical value for positive HBsAg is 0.05 IU/ml, for positive anti-HBs is 10 mIU/mL.
2 LIAISON-XL quantification assays (CLIA, Diasorin): critical value for positive HBsAg is 0.05 IU/ml, for positive anti-HBs is 3 mIU/mL.
3 Cobas E601 quantitative electrochemical luminescence assays (ECLIA, Roche Diagnostics): critical value for positive HBsAg is 0.05 IU/ml, for positive anti-HBs is 10 IU/L.
The coexistence of HBsAg and anti-HBs might be associated with important clinical concerns and this profile could be linked to advanced fibrosis, hepatocellular carcinoma (HCC) and liver failure [69]. Among 92 patients with simultaneous HBsAg/anti-HBs occurrence in Korea, 6.5% suffered from liver cirrhosis and 2.2% were diagnosed with HCC [70]. With the follow-up of 1042 non-HCC patients for a median 4.3 years, 13.7% (10/73) patients with coexistence developed HCC, which was higher compared with HBsAg alone group (6.9%, 67/969) [71]. Coexistence of HBsAg/anti-HBs independently increased the risk of HCC in CHB infection [71, 72]. There is a 3.08-fold (95%CI: 1.26-7.55) higher risk of HCC than the HBsAg alone patients [53]. Characteristics in HBsAg/anti-HBs coexistence such as age over 50 years and abnormal ALT levels were also risk factors for HCC, which was linked to a long history of HBV infection with advanced liver fibrosis and active inflammation [29, 72]. Serum PIVKA-II, better correlated with portal vein invasion, larger tumor size and recurrence of HBV-associated HCC [73], was also observed with higher levels in coexistence group [69].
Mutations and HCC
Mutations in HBV ORFs and the encoding mutated proteins connected the progressive liver diseases with the concurrent HBsAg/anti-HBs. HCC patients with HBsAg/anti-HBs coexistence had a significantly higher frequency of N-glycosylation mutations in the first loop of S proteins compared to non-HCC patients (22.4% vs. 8.0%, p<0.01) [22]. Longitudinal observation revealed these N-glycosylation mutations were detected 1-4 years prior to HCC occurrence, which was also found in other mutation patterns, the preS deletions [72] and T1762+A1764 double mutations [82].
PreS/S deletions in HCC were consistent with those from coexistence group, which displayed lower prevalence in HBsAg alone group. These deletions in HCC patients collected from their serum or liver tissues [83], clustered in the immune epitopes for T and B cells, especially in the C-terminal of preS1 and N-terminal of preS2, affecting the recognition of virus by immune cells [72]. These deletions can trigger a protein kinase C (PKC) dependent activation of signal transduction cascade and induce the AP-1 and NF-κB transcription factors to enhanced hepatocytes proliferation [84], demonstrating the mutant proteins encoded by preS/S can be the transcriptional activators facilitating the invasions of the tumors [85]. Mutations in the preS2 initiation codons affect the secretion of the virus and cause the accumulation of viral proteins in host cells. The mutation at the site 120 in genotype B, utilizing a different pathway in liver disease progression that involves high expression of NF-κB subunit p50 [28], activated NF-κB as one of the mechanisms in inducing advanced liver disease. PreS1 and preS2 deletions in ground glass hepatocytes is characterized by an abnormal retention of mutant large and middle surface proteins (LHBs and MHBs) in endoplasmic reticulum (ER) [86]. Deletions in preS from coexistence patients may generate truncated LHB and MHB. The mutants may also retain in the ER and induce ER stress signals as well as oxidative stress responses, upregulating COX-2 and cyclin A to induce cell cycle progression and generating a large amount of reactive oxygen species (ROS). Genomic instability in hepatocytes [87] and prolonged hepatocellular injuries can initiate a programmed response characterized by inflammation, regenerative hyperplasia, transcriptional deregulation and aneuploidy [72]. Such effects can lead to the development of HCC, as proven in studies with transgenic mice [83].
BCP double mutations A1762T+G1764A with a significantly higher frequency in the coexistence group functioned as HCC predictors in HBV genotype B or C carriers [35, 88]. This double mutation shared the common pattern including the codon 'AGG' with the sense strand sequence of p53 [35]. Aflatoxin can bind and induce mutations in this codons of p53 during the pathogenesis of HCC [89], suggesting A1762T+G1764A might be a mutational hot spot targeted by aflatoxin or some other chemical agent. Moreover, A1762T+G1764A harbored in X gene led to L130M+V131I substitutions in HBxAg (Figure 2). Involved in many intracellular signal pathways, HBxAg was closely associated with cell proliferation and cell apoptosis [90] and the mutant proteins displayed stronger tumorigenic effects than the wildtype. In Huh-7 cell lines [91], the double mutation accelerated fibrosis and cirrhosis via inducing ROS production and mitochondrial depolarization. Their contribution to HCC may be direct involvement in host cell proliferation and hepatocarcinogenesis via altering expression of cell cycle regulatory genes. Variant with L130M+V131I mutations may also exert a synergistic effect in accelerating the progression to HCC [92], which was recently reported to promote HCC progression by activating AKT/FOXO1 pathway and inducing more severe inflammation in liver via arachidonic acid metabolism [93].
Dilemma for treatment
In the clinical cases, Galati [94] reported the dilemma of treatment when HBsAg and anti-HBs were coexisting. Interferon therapy as well as several nucleotide analogues (NAs) specific for HBV failed to achieve virologic responses. Only with the combination of second-generation NAs did the patient finally experience HBsAg loss. In another case [95], after antiviral treatment with entecavir for eight years, the female patient showed the serological profile characterized by high titers of anti-HBs and the consistent levels of HBsAg compared to three year ago. Although HBV DNA was undetectable, the patient had the risk for HBV reactivation and further treatment went into a dilemma. The irregular HBV DNA were also reported in different studies with the coexistence situation [18, 96]. Antiviral therapies may decrease serum HBV-DNA levels, concealing the influence of serum HBV DNA levels on HCC development [71]. Reversely, HBsAg/anti-HBs coexistence patients with relatively high viral loads suggested active virus replication and raised the problem of HBV transmission [14, 48]. For HBsAg/anti-HBs coexistence patients, the irregular viral load may mislead the treatment and risk evaluation.
Under some special conditions, the HBsAg/anti-HBs coexistence may increase complexity for treatment and predict poor prognosis. Approximately 10-15% of hepatitis C virus (HCV)-infected people and about 10% human immunodeficiency virus (HIV)-infected people were diagnosed with HBV [97, 98]. During coinfection, interactions between the two viruses may change the natural history of both mono-infections. Reactivation of HBV can occur in HIV-driven immune suppression and HCV-related direct-acting antivirals therapy, and then facilitate coexistence of HBsAg and anti-HBs. Moreover, the coexistence may also occur in the presence of hyperimmune function. As HBV is a common comorbidity among rheumatic patients, HBV infection may reactivate in patients with autoimmune inflammatory rheumatic diseases when they take immunosuppressive drugs or undergo biological therapies [99, 100]. The HBV infection needs surveillance, and markers like HBsAg and anti-HBs need to be detected before and after the treatment of co-infected diseases.
Clinical significance and monitoring suggestions
A certain proportion of population may be confronted with coexistence of HBsAg and anti-HBs. The emergence of anti-HBs does not demonstrate that HBsAg can be completely or effectively eliminated [96], the virus can replicate continuously with the presence of anti-HBs. While some individuals are prone to the coexistence in clinical practice. Firstly, patients who have risks of HBV breakthrough or reactivation are in need of follow-up and monitoring. They may undergo immunosuppression condition because of immunosuppressive therapies and/or coinfection with immunodeficiency diseases. Secondly, although HBV vaccines were delivered to newborns in many countries, the vaccine anti-HBs had lower binding capacity to mutant HBsAg [17] and anti-HBs levels declined with age [101], some young individuals still had risks of HBV infection, especially those born to HBsAg positive mothers. Thirdly, in occult HBV infection (OBI), HBsAg was undetectable with the existing detection methods. However, some specific mutations in preS/S gene were found in anti-HBs positive OBI samples [102] and might be adaptive substitutions selected under immune pressure. These are risk factors for virus reactivation under anti-HBs selection and changes of serological markers in OBI patients is worthy of clinical attention.
To further validate and monitor HBsAg/anti-HBs coexistence, diagnosis procedures and surveillance measures are required. Although no standard diagnosis procedure is available, what we can do is to apply reliable and sensitive detection methods for both HBsAg and anti-HBs in the first place, which can avoid false positive or false negative results. The results should also be combined with other markers such as HBV DNA and liver function indicators. Quantification of HBV DNA, along with genotyping and mutation analysis can be helpful to identify the virus, whether it's a wild-type or mutant. After ensuring the accuracy of our test results, monitoring measures are specific for different groups of people. For pregnant women and infants, we suggest to screen all pregnant women for HBsAg and provide immunoprophylaxis to infants of HBsAg-positive mothers. Moreover, routine immunization of HBV vaccine is needed for all infants. For adolescents and adults, especially those at a high risk of HBV infection (injection drug users and individuals who participate in high-risk sexual behaviors), we suggest to screen them for both HBsAg and anti-HBs periodically. If the anti-HBs titer is below the protective level, a booster dose vaccine is needed to strengthen immunity. For chronic HBV-infected patients and OBI patients, enhanced surveillance of liver function and serological indicators is urgently needed. They should go through longer follow-up of their serological patterns, and the physicians should pay close attention to the changes in the HBsAg and anti-HBs levels.
Conclusions
HBV is related to large-scale morbidity and mortality, bringing heavy burden to the developing countries. Knowledge of paradoxical serological patterns can be helpful for the control and eradication purpose of HBV. Clinical laboratories encounter HBsAg and anti-HBs double positivity more frequently at present. Its possible causes and clinical significance (Figure 3) are worth to be considered and further explored.
Viral mutation host factors, the interaction between viruses and hosts, as well as screening method were connected to the HBsAg/anti-HBs coexistence. Mutations in viral genome, especially mutations in MHR and deletions in preS gene, were widely studied and reported to change the antigenicity of HBsAg. Superinfection and breakthrough infection can lead to simultaneous detection of serotype-specific HBsAg and anti-HBs. Changed epitopes cannot be recognized by immune cells and suppressed immune system allowed the complex infections. Coexistence is not the indicator of improvement and an increased risk of adverse clinical outcomes still exists.
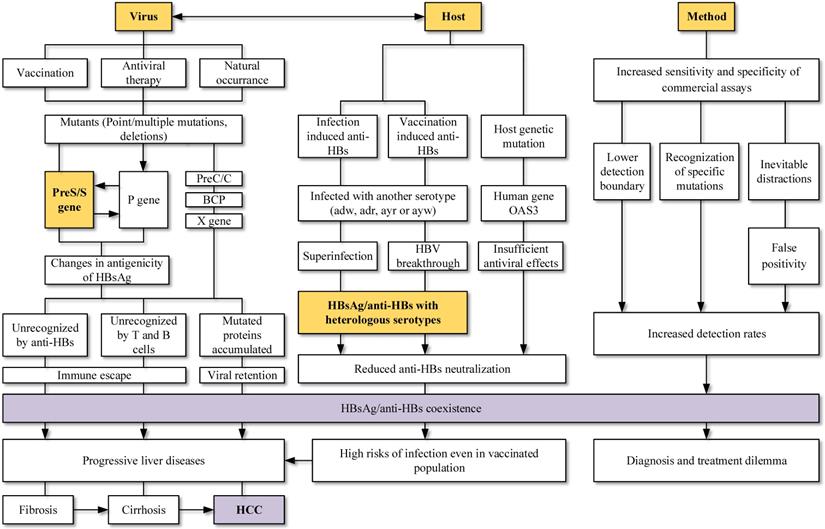
Mutations in different ORFs of HBV is a possible and convincing interpretation for the HBsAg/anti-HBs coexistence. Point mutations in the preS/S gene, RT region of polymerase gene, as well as preS deletions can both change the stability and immunogenicity of HBsAg, favoring the selection of immune escape variants and decreasing the affinity with anti-HBs. Vaccination, antiviral therapy and even HBV chronic infection without any treatment can lead to accumulation of those HBV variants. Relationship between virus and host should also be considered. Superinfection and breakthrough infection are associated with the presence of heterologous subtype-specific anti-HBs rather than mutations in the gene region, particularly in the clinical settings of immunosuppression. The improvements in screening methods may lead to the detection of lower HBsAg levels, mutants and immune complexes, increasing the detection of coexistence. Keep updating the explanations for different serological patterns is necessary.
Lastly, the coexistence is not the indicator of recovery and an increased risk of adverse clinical outcomes still exists. Patients with HBsAg/anti-HBs coexistence may progress into liver cirrhosis, HCC and liver failure. Vaccinated persons had risks of infection and the coexistence of HBsAg/anti-HBs may lead to a dilemma for diagnosis and treatment. More studies about the coexistence of HBsAg/anti-HBs should be carried out and corresponding countermeasures shall be taken by both researchers and physicians.
Abbreviations
HBsAg: hepatitis B surface antigen; Anti-HBs: hepatitis B surface antibody; HBV: hepatitis B virus; CHB: chronic hepatitis B infection; HCC: hepatocellular carcinoma; aa: amino acids; CTL: cytotoxic lymphocyte; ORFs: open reading frames; MHR: major hydrophilic region; RT: reverse transcriptase; BCP: basal core promoter; OAS3: oligoadenylate synthetases 3; EIAs: enzyme immunoassays; CIAs: chemiluminescent immunoassays; CLEIA: chemiluminescent enzyme immunoassay; PIVKA-II: proteins induced by vitamin k absence; ER: endoplasmic reticulum; LHB: large hepatitis B surface antigen; MHB: middle hepatitis B surface antigen; SHB: small hepatitis B surface antigen; PKC: protein kinase C; ROS: reactive oxygen species; NA: nucleotide analogues; HBxAg: hepatitis B x antigen; HIV: human immunodeficiency virus; HCV: hepatitis C virus; OBI: occult hepatitis infection.
Supplementary Material
Supplementary figures and tables.
Acknowledgements
Author contributions
Xinyi Jiang contributed to the writing of the manuscript; Le Chang, Ying Yan and Lunan Wang contributed to critical revision and approval of the final manuscript. Xinyi Jiang, Le Chang, Ying Yan and Lunan Wang all approved the final draft submitted. All authors have read and agreed to the published version of the manuscript.
Funding
This work received no specific grant from any funding agency.
Competing Interests
The authors have declared that no competing interest exists.
References
1. WHO. WHO (2017) Global hepatitis report. 2017.
2. Liang X, Bi S, Yang W, Wang L, Cui G, Cui F. et al. Epidemiological serosurvey of hepatitis B in China-declining HBV prevalence due to hepatitis B vaccination. Vaccine. 2009;27:6550-7
3. Ott JJ, Stevens GA, Groeger J, Wiersma ST. Global epidemiology of hepatitis B virus infection: new estimates of age-specific HBsAg seroprevalence and endemicity. Vaccine. 2012;30:2212-9
4. Tong MJ, Pan CQ, Han SB, Lu DS, Raman S, Hu KQ. et al. An expert consensus for the management of chronic hepatitis B in Asian Americans. Aliment Pharmacol Ther. 2018
5. Hatazawa Y, Yano Y, Okada R, Tanahashi T, Hayashi H, Hirano H. et al. Quasispecies variant of pre-S/S gene in HBV-related hepatocellular carcinoma with HBs antigen positive and occult infection. Infect Agent Cancer. 2018;13:7
6. Arnold W, Hess G, Meyer zum Buschenfelde KH, Kosters W, Biswas R, Strauch M. et al. [Simultaneous occurrence in the serum of hepatitis B surface antigen (HBsAg) and antibodies against HBsAg (Anti-HBs) of different subtypes. (Serologic and fluorescence histologic studies)]. Verh Dtsch Ges Inn Med. 1976;82(Pt 1):405-7
7. Ben-Porath E, Wands JR, Marciniak RA, Wong MA, Hornstein L, Ryder R. et al. Structural analysis of hepatitis B surface antigen by monoclonal antibodies. J Clin Invest. 1985;76:1338-47
8. Alavian SM, Carman WF, Jazayeri SM. HBsAg variants: diagnostic-escape and diagnostic dilemma. Journal of clinical virology: the official publication of the Pan American Society for Clinical Virology. 2013;57:201-8
9. Wilson JN, Nokes DJ, Carman WF. Current status of HBV vaccine escape variants-a mathematical model of their epidemiology. Journal of viral hepatitis. 1998;5(Suppl 2):25-30
10. Gunther S, Fischer L, Pult I, Sterneck M, Will H. Naturally occurring variants of hepatitis B virus. Adv Virus Res. 1999;52:25-137
11. Larralde O, Dow B, Jarvis L, Davidson F, Petrik J. Hepatitis B escape mutants in Scottish blood donors. Medical microbiology and immunology. 2013;202:207-14
12. Wu C, Zhang X, Tian Y, Song J, Yang D, Roggendorf M. et al. Biological significance of amino acid substitutions in hepatitis B surface antigen (HBsAg) for glycosylation, secretion, antigenicity and immunogenicity of HBsAg and hepatitis B virus replication. Journal of General Virology. 2010;91:483-92
13. Zheng X, Weinberger KM, Gehrke R, Isogawa M, Hilken G, Kemper T. et al. Mutant hepatitis B virus surface antigens (HBsAg) are immunogenic but may have a changed specificity. Virology. 2004;329:454-64
14. Lada O, Benhamou Y, Poynard T, Thibault V. Coexistence of hepatitis B surface antigen (HBs Ag) and anti-HBs antibodies in chronic hepatitis B virus carriers: influence of "a" determinant variants. Journal of virology. 2006;80:2968-75
15. Wang L, Liu H, Ning X, Gao F. Sequence analysis of the S gene region in HBV DNA from patients positive for both HBsAg and HBsAb tests. Hepatol Res. 2010;40:1212-8
16. Liu Y, Zhang L, Zhou JY, Pan J, Hu W, Zhou YH. Clinical and Virological Characteristics of Chronic Hepatitis B Patients with Coexistence of HBsAg and Anti-HBs. PLoS One. 2016;11:e0146980
17. Fu X, Chen J, Chen H, Lin J, Xun Z, Li S. et al. Mutation in the S gene of hepatitis B virus and anti-HBs subtype-nonspecificity contributed to the co-existence of HBsAg and anti-HBs in patients with chronic hepatitis B virus infection. J Med Virol. 2017;89:1419-26
18. Zhang JM, Xu Y, Wang XY, Yin YK, Wu XH, Weng XH. et al. Coexistence of hepatitis B surface antigen (HBsAg) and heterologous subtype-specific antibodies to HBsAg among patients with chronic hepatitis B virus infection. Clinical infectious diseases: an official publication of the Infectious Diseases Society of America. 2007;44:1161-9
19. Torresi J, Earnest-Silveira L, Deliyannis G, Edgtton K, Zhuang H, Locarnini SA. et al. Reduced antigenicity of the hepatitis B virus HBsAg protein arising as a consequence of sequence changes in the overlapping polymerase gene that are selected by lamivudine therapy. Virology. 2002;293:305-13
20. Rezaee R, Poorebrahim M, Najafi S, Sadeghi S, Pourdast A, Alavian SM. et al. Impacts of the G145R Mutation on the Structure and Immunogenic Activity of the Hepatitis B Surface Antigen: A Computational Analysis. Hepatitis monthly. 2016;16:e39097-e
21. Huang X, Qin Y, Zhang P, Tang G, Shi Q, Xu J. et al. PreS deletion mutations of hepatitis B virus in chronically infected patients with simultaneous seropositivity for hepatitis-B surface antigen and anti-HBS antibodies. Journal of medical virology. 2010;82:23-31
22. Qiao Y, Lu S, Xu Z, Li X, Zhang K, Liu Y. et al. Additional N-glycosylation mutation in the major hydrophilic region of hepatitis B virus S gene is a risk indicator for hepatocellular carcinoma occurrence in patients with coexistence of HBsAg/anti-HBs. Oncotarget. 2017;8:61719-30
23. Yu D-M, Li X-H, Mom V, Lu Z-H, Liao X-W, Han Y. et al. N-glycosylation mutations within hepatitis B virus surface major hydrophilic region contribute mostly to immune escape. Journal of Hepatology. 2014;60:515-22
24. Zhou TC, Li X, Li L, Li XF, Zhang L, Wei J. Evolution of full-length genomes of HBV quasispecies in sera of patients with a coexistence of HBsAg and anti-HBs antibodies. Sci Rep. 2017;7:661
25. Lee SA, Kim K, Kim H, Kim BJ. Nucleotide change of codon 182 in the surface gene of hepatitis B virus genotype C leading to truncated surface protein is associated with progression of liver diseases. J Hepatol. 2012;56:63-9
26. Levrero M, Zucman-Rossi J. Mechanisms of HBV-induced hepatocellular carcinoma. J Hepatol. 2016;64:S84-s101
27. Zhang D, Dong P, Zhang K, Deng L, Bach C, Chen W. et al. Whole genome HBV deletion profiles and the accumulation of preS deletion mutant during antiviral treatment. BMC Microbiol. 2012;12:307
28. Siburian MD, Suriapranata IM, Wanandi SI. Pre-S2 Start Codon Mutation of Hepatitis B Virus Subgenotype B3 Effects on NF-kappaB Expression and Activation in Huh7 Cell Lines. Viral immunology. 2018;31:362-70
29. Colson P, Borentain P, Motte A, Henry M, Moal V, Botta-Fridlund D. et al. Clinical and virological significance of the co-existence of HBsAg and anti-HBs antibodies in hepatitis B chronic carriers. Virology. 2007;367:30-40
30. Liu W, Hu T, Wang X, Chen Y, Huang M, Yuan C. et al. Coexistence of hepatitis B surface antigen and anti-HBs in Chinese chronic hepatitis B virus patients relating to genotype C and mutations in the S and P gene reverse transcriptase region. Arch Virol. 2012;157:627-34
31. Ding F, Miao XL, Li YX, Dai JF, Yu HG. Mutations in the S gene and in the overlapping reverse transcriptase region in chronic hepatitis B Chinese patients with coexistence of HBsAg and anti-HBs. Braz J Infect Dis. 2016;20:1-7
32. Choi YM, Lee SY, Kim BJ. Naturally occurring hepatitis B virus reverse transcriptase mutations related to potential antiviral drug resistance and liver disease progression. World J Gastroenterol. 2018;24:1708-24
33. Torresi J, Earnest-Silveira L, Deliyannis G, Edgtton K, Zhuang H, Locarnini SA. et al. Reduced antigenicity of the hepatitis B virus HBsAg protein arising as a consequence of sequence changes in the overlapping polymerase gene that are selected by lamivudine therapy. Virology. 2002;293:305-13
34. Chen Y, Qian F, Yuan Q, Li X, Wu W, Guo X. et al. Mutations in hepatitis B virus DNA from patients with coexisting HBsAg and anti-HBs. Journal of clinical virology: the official publication of the Pan American Society for Clinical Virology. 2011;52:198-203
35. Kuang SY, Jackson PE, Wang JB, Lu PX, Munoz A, Qian GS. et al. Specific mutations of hepatitis B virus in plasma predict liver cancer development. Proc Natl Acad Sci U S A. 2004;101:3575-80
36. Wu IC, Liu W-C, Chang T-T. Applications of next-generation sequencing analysis for the detection of hepatocellular carcinoma-associated hepatitis B virus mutations. J Biomed Sci. 2018;25:51
37. Ponde RA. The underlying mechanisms for the "simultaneous HBsAg and anti-HBs serological profile". Eur J Clin Microbiol Infect Dis. 2011;30:1325-40
38. Liu K, Xie M, Lu X, Yu H, Wang H, Xu Y. et al. Mutations within the major hydrophilic region (MHR) of Hepatitis B virus from individuals with simultaneous HBsAg and anti-HBs in Guangzhou, Southern China. J Med Virol. 2018;90:1337-42
39. Chen B-F, Liu C-J, Jow G-M, Chen P-J, Kao J-H, Chen D-S. High prevalence and mapping of pre-S deletion in hepatitis B virus carriers with progressive liver diseases. Gastroenterology. 2006;130:1153-68
40. Ashton-Rickardt PG, Murray K. Mutants of the hepatitis B virus surface antigen that define some antigenically essential residues in the immunodominant a region. Journal of medical virology. 1989;29:196-203
41. Lin X, Yuan ZH, Wu L, Ding JP, Wen YM. A single amino acid in the reverse transcriptase domain of hepatitis B virus affects virus replication efficiency. Journal of virology. 2001;75:11827-33
42. Xiang KH, Michailidis E, Ding H, Peng YQ, Su MZ, Li Y. et al. Effects of amino acid substitutions in hepatitis B virus surface protein on virion secretion, antigenicity, HBsAg and viral DNA. J Hepatol. 2017;66:288-96
43. Yeh C-T. Development of HBV S gene mutants in chronic hepatitis B patients receiving nucleotide/nucleoside analogue therapy. Antivir Ther (Lond). 2010;15:471-5
44. Wang J, Zhu B, Lu M, Yang D. Hepatitis B virus preS/S gene mutations and their clinical implications. Annals of Blood. 2017 2
45. Warner N, Locarnini S. The antiviral drug selected hepatitis B virus rtA181T/sW172* mutant has a dominant negative secretion defect and alters the typical profile of viral rebound. Hepatology (Baltimore, Md). 2008;48:88-98
46. Hunt CM, McGill JM, Allen MI, Condreay LD. Clinical relevance of hepatitis B viral mutations. Hepatology (Baltimore, Md). 2000;31:1037-44
47. Chen L, Zheng C-X, Lin M-H, Huang Z-X, Chen R-H, Li Q-G. et al. Distinct quasispecies characteristics and positive selection within precore/core gene in hepatitis B virus HBV associated acute-on-chronic liver failure. Journal of gastroenterology and hepatology. 2012 28
48. Wang Y, Ng W, Kang J, Yap I, Seet B, Teo J. et al. Serological profiles of hepatitis B carrier patients in Singapore with special reference to the frequency and significance of concurrent presence of HBsAg and anti-HBs. Singapore medical journal. 1996;37:150-2
49. Purcell RH, Gerin JL. Hepatitis B subunit vaccine: a preliminary report of safety and efficacy tests in chimpanzees. Am J Med Sci. 1975;270:395-9
50. Hayashi J, Noguchi A, Nakashima K, Morofuji M, Kashiwagi S. Frequency of concurrence of hepatitis B surface antigen and antibody in a large number of carriers in Okinawa, Japan. Gastroenterologia Japonica. 1990;25:593-7
51. Tabor E, Gerety RJ, Smallwood LA, Barker LF. Coincident hepatitis B surface antigen and antibodies of different subtypes in human serum. J Immunol. 1977;118:369-70
52. Xia G, Nainan OV, Jia Z. [Characterization and distribution of hepatitis B virus genotypes and subtypes in 4 provinces of China]. Zhonghua Liu Xing Bing Xue Za Zhi. 2001;22:348-51
53. Kwak MS, Chung GE, Yang JI, Yim JY. Long-term outcomes of HBsAg/anti-HBs double-positive versus HBsAg single-positive patients with chronic hepatitis B. Sci Rep. 2019;9:19417
54. Su YC, Lin PC, Yu HC, Wu CC. Hepatitis B virus reactivation in patients with resolved hepatitis B virus infection receiving chemotherapy or immunosuppressive therapy. European journal of gastroenterology & hepatology. 2018
55. Wang S, Wang J, Fan MJ, Li TY, Pan H, Wang X. et al. Identified OAS3 gene variants associated with coexistence of HBsAg and anti-HBs in chronic HBV infection. Journal of viral hepatitis. 2018
56. Huzly D, Schenk T, Jilg W, Neumann-Haefelin D. Comparison of nine commercially available assays for quantification of antibody response to hepatitis B virus surface antigen. J Clin Microbiol. 2008;46:1298-306
57. Kinn S, Akhavan S, Agut H, Thibault V. Performance of the DiaSorin LIAISON((R)) anti-HBs II for the detection of hepatitis B surface antibodies: comparison with the Abbott Architect anti-HBs assay. Journal of clinical virology: the official publication of the Pan American Society for Clinical Virology. 2011;50:297-302
58. Shin YW, Cho DH, Song GW, Kim SH. A New ELISA to Overcome the Pitfalls in Quantification of Recombinant Human Monoclonal Anti-HBs, GC1102, by Commercial Immunoassays. Biological procedures online. 2018;20:18
59. Lou SC, Pearce SK, Lukaszewska TX, Taylor RE, Williams GT, Leary TP. An improved Abbott ARCHITECT assay for the detection of hepatitis B virus surface antigen (HBsAg). Journal of clinical virology: the official publication of the Pan American Society for Clinical Virology. 2011;51:59-63
60. Lou S, Taylor R, Pearce S, Kuhns M, Leary T. An ultra-sensitive Abbott ARCHITECT((R)) assay for the detection of hepatitis B virus surface antigen (HBsAg). Journal of clinical virology: the official publication of the Pan American Society for Clinical Virology. 2018;105:18-25
61. Deguchi M, Yamashita N, Kagita M, Asari S, Iwatani Y, Tsuchida T. et al. Quantitation of hepatitis B surface antigen by an automated chemiluminescent microparticle immunoassay. J Virol Methods. 2004;115:217-22
62. Pancher M, Désiré N, Ngo Y, Akhavan S, Pallier C, Poynard T. et al. Coexistence of circulating HBsAg and anti-HBs antibodies in chronic hepatitis B carriers is not a simple analytical artifact and does not influence HBsAg quantification. Journal of clinical virology: the official publication of the Pan American Society for Clinical Virology. 2015;62:32-7
63. Thibault V, Servant-Delmas A, Ly TD, Roque-Afonso AM, Laperche S. Performance of HBsAg quantification assays for detection of Hepatitis B virus genotypes and diagnostic escape-variants in clinical samples. Journal of clinical virology: the official publication of the Pan American Society for Clinical Virology. 2017;89:14-21
64. Matsubara N, Kusano O, Sugamata Y, Itoh T, Mizuii M, Tanaka J. et al. A novel hepatitis B virus surface antigen immunoassay as sensitive as hepatitis B virus nucleic acid testing in detecting early infection. Transfusion. 2009;49:585-95
65. Shinkai N, Matsuura K, Sugauchi F, Watanabe T, Murakami S, Iio E. et al. Application of a newly developed high-sensitivity HBsAg chemiluminescent enzyme immunoassay for hepatitis B patients with HBsAg seroclearance. Journal of clinical microbiology. 2013;51:3484-91
66. Mair DC, Brecher ME, Hom E, Owen HG, Shea TC. False-positive hepatitis B surface antigen screening test results in patients receiving granulocyte-colony-stimulating factor. Transfusion. 1996;36:948-51
67. Olde C, Garcia M. Hepatitis B vaccine as a cause of false positive hepatitis B surface antigen. J CANNT. 1998;8:20-1
68. Parry JV, Mortimer PP, Friderich P, Connell JA. Faulty washers and soiled micropipettors may generate false positive serological results. Clin Diagn Virol. 1997;7:173-81
69. Jin ZZ, Jin FF, Liu X, Liu N, Wen F, Lou JL. Coexistence of low levels of HBsAg and high levels of anti-HBs may increase risk of hepatocellular carcinoma in chronic hepatitis B patients with high HBV load. Braz J Infect Dis. 2019;23:343-51
70. Chan Jeong P. et al. Coexistence of Hepatitis B Surface Antigen and Hepatitis B Surface Antibody in the Same Serum. Ann Lab Med. 1987;7:321-34
71. Seo SI, Choi HS, Choi BY, Kim HS, Kim HY, Jang MK. Coexistence of hepatitis B surface antigen and antibody to hepatitis B surface may increase the risk of hepatocellular carcinoma in chronic hepatitis B virus infection: a retrospective cohort study. J Med Virol. 2014;86:124-30
72. Jang JS, Kim HS, Kim HJ, Shin WG, Kim KH, Lee JH. et al. Association of concurrent hepatitis B surface antigen and antibody to hepatitis B surface antigen with hepatocellular carcinoma in chronic hepatitis B virus infection. J Med Virol. 2009;81:1531-8
73. Jang MK, Kim HS, Jang JS, Geon W, Kim KH, Lee JH. et al. Prognostic value of serum PIVKA-II in HBV-associated hepatocellular carcinoma: Comparison with serum alpha-feto protein. Journal of Hepatology - J HEPATOL. 2008 48
74. Ding F, Yu HG, Li YX, Cui N, Dai JF, Yu JP. Sequence analysis of the HBV S protein in Chinese patients with coexisting HBsAg and anti-HBs antibodies. J Med Virol. 2015;87:2067-73
75. Pu Z, Li D, Wang A, Su H, Shao Z, Zhang J. et al. Epidemiological characteristics of the carriers with coexistence of HBsAg and anti-HBs based on a community cohort study. Journal of viral hepatitis. 2016;23:286-93
76. Hou W, Huo Z, Du Y, Wang C, Syn WK. Characteristics of amino acid substitutions within the "a" determinant region of hepatitis B virus in chronically infected patients with coexisting HBsAg and anti-HBs. Clinics and research in hepatology and gastroenterology. 2019
77. Lee WM, King WC, Schwarz KB, Rule J, Lok ASF. Prevalence and clinical features of patients with concurrent HBsAg and anti-HBs: Evaluation of the hepatitis B research network cohort. Journal of viral hepatitis. 2020;27:922-31
78. Lee BS, Cho YK, Jeong SH, Lee JH, Lee D, Park NH. et al. Nationwide seroepidemiology of hepatitis B virus infection in South Korea in 2009 emphasizes the coexistence of HBsAg and anti-HBs. J Med Virol. 2013;85:1327-33
79. de Campos Albuquerque I, Sousa MT, Santos MDC, Nunes JDC, Moraes MJD, Gomes-Gouvêa MS. et al. Mutation in the S gene a determinant of the hepatitis B virus associated with concomitant HBsAg and anti-HBs in a population in Northeastern Brazil. Journal of medical virology. 2017;89:458-62
80. Pancher M, Desire N, Ngo Y, Akhavan S, Pallier C, Poynard T. et al. Coexistence of circulating HBsAg and anti-HBs antibodies in chronic hepatitis B carriers is not a simple analytical artifact and does not influence HBsAg quantification. Journal of clinical virology: the official publication of the Pan American Society for Clinical Virology. 2015;62:32-7
81. Kohno H, Inoue T, Tsuda F, Okamoto H, Akahane Y. Mutations in the envelope gene of hepatitis B virus variants co-occurring with antibody to surface antigen in sera from patients with chronic hepatitis B. The Journal of general virology. 1996;77( Pt 8):1825-31
82. Qu L, Kuai X, Liu T, Chen T, Ni Z, Shen X. Pre-S deletion and complex mutations of hepatitis B virus related to young age hepatocellular carcinoma in Qidong, China. PloS one. 2013;8:e59583
83. Wang HC, Huang W, Lai MD, Su IJ. Hepatitis B virus pre-S mutants, endoplasmic reticulum stress and hepatocarcinogenesis. Cancer science. 2006;97:683-8
84. Hildt E, Munz B, Saher G, Reifenberg K, Hofschneider PH. The PreS2 activator MHBs(t) of hepatitis B virus activates c-raf-1/Erk2 signaling in transgenic mice. EMBO J. 2002;21:525-35
85. Hildt E, Saher G, Bruss V, Hofschneider PH. The hepatitis B virus large surface protein (LHBs) is a transcriptional activator. Virology. 1996;225:235-9
86. Gerber MA, Hadziyannis S, Vissoulis C, Schaffner F, Paronetto F, Popper H. Electron microscopy and immunoelectronmicroscopy of cytoplasmic hepatitis B antigen in hepatocytes. Am J Pathol. 1974;75:489-502
87. Hagen TM, Huang S, Curnutte J, Fowler P, Martinez V, Wehr CM. et al. Extensive oxidative DNA damage in hepatocytes of transgenic mice with chronic active hepatitis destined to develop hepatocellular carcinoma. Proc Natl Acad Sci U S A. 1994;91:12808-12
88. Baptista M, Kramvis A, Kew MC. High prevalence of 1762(T) 1764(A) mutations in the basic core promoter of hepatitis B virus isolated from black Africans with hepatocellular carcinoma compared with asymptomatic carriers. Hepatology (Baltimore, Md). 1999;29:946-53
89. Smela ME, Currier SS, Bailey EA, Essigmann JM. The chemistry and biology of aflatoxin B(1): from mutational spectrometry to carcinogenesis. Carcinogenesis. 2001;22:535-45
90. Arbuthnot P, Kew M. Hepatitis B virus and hepatocellular carcinoma. Int J Exp Pathol. 2001;82:77-100
91. Siddiqui ZI, Farooqui SR, Azam SA, Afroz M, Wajid S, Parveen S. et al. A comparative study of hepatitis B virus X protein mutants K130M, V131I and KV130/131MI to investigate their roles in fibrosis, cirrhosis and hepatocellular carcinoma. Journal of viral hepatitis. 2017;24:1121-31
92. Al-Qahtani AA, Al-Anazi MR, Nazir N, Ghai R, Abdo AA, Sanai FM. et al. Hepatitis B virus (HBV) X gene mutations and their association with liver disease progression in HBV-infected patients. Oncotarget. 2017;8:105115-25
93. Chiu AP, Tschida BR, Sham T-T, Lo LH, Moriarity BS, Li X-X. et al. HBx-K130M/V131I Promotes Liver Cancer in Transgenic Mice via AKT/FOXO1 Signaling Pathway and Arachidonic Acid Metabolism. Mol Cancer Res. 2019;17:1582-93
94. Galati G, De Vincentis A, Vespasiani-Gentilucci U, Gallo P, Vincenti D, Solmone MC. et al. Coexistence of HBsAg and HBsAb in a difficult-to-treat chronic hepatitis B: loss of HBsAg with entecavir plus tenofovir combination. BMC Gastroenterol. 2014;14:94
95. Suceveanu A, Mazilu L, Voinea C, Suceveanu A. Concomitant Serum Presence of Hepatitis B Surface Antigen (HBsAg) and High Titers of Hepatitis B Surface Antibodies (Anti - HBsAb) in a Patient with Chronic Hepatitis B (HBV) Genotype D from Black Sea Coast Region; Case Report. Hepatitis Monthly. 2018 In Press
96. Zhu D, Chen W, Xu C, Yu X, Xi Y. Virology and Serological Characteristics of Chronic Hepatitis B Patients with the Co-existence of HBsAg and Anti-HBs Antibodies. Clin Lab. 2020 66
97. Abdelaal R, Yanny B, El Kabany M. HBV/HCV Coinfection in the Era of HCV-DAAs. Clinics in liver disease. 2019;23:463-72
98. Sarmati L, Malagnino V. HBV Infection in HIV-Driven Immune Suppression. Viruses. 2019 11
99. Ditto MC, Parisi S, Varisco V, Talotta R, Batticciotto A, Antivalle M. et al. Prevalence of hepatitis B virus infection and risk of reactivation in rheumatic population undergoing biological therapy. Clinical and experimental rheumatology. 2020
100. Canzoni M, Marignani M, Sorgi ML, Begini P, Biondo MI, Caporuscio S. et al. Prevalence of Hepatitis B Virus Markers in Patients with Autoimmune Inflammatory Rheumatic Diseases in Italy. Microorganisms. 2020 8
101. Miao N, Zheng H, Sun X, Zhang G, Wang F. Protective effect of vaccinating infants with a 5 µg recombinant yeast-derived hepatitis B vaccine and the need for a booster dose in China. Sci Rep. 2020;10:18155
102. Wang J, Zhang P, Zeng J, Du P, Zheng X, Ye X. et al. Occurrence of occult hepatitis B virus infection associated with envelope protein mutations according to anti-HBs carriage in blood donors. International journal of infectious diseases: IJID: official publication of the International Society for Infectious Diseases. 2020;92:38-45
Author contact
Corresponding author: Prof. Lunan Wang, E-mail: lunan99com; Tax numbers: +861085133609; No.1 Dahua Road, Dong Dan, Beijing 100730, P. R. China.

 Global reach, higher impact
Global reach, higher impact