10
Impact Factor
ISSN: 1449-2288
Int J Biol Sci 2021; 17(14):3717-3727. doi:10.7150/ijbs.58220 This issue Cite
Research Paper
Gene Presence/Absence Variation analysis of coronavirus family displays its pan-genomic diversity
1. Department of Bioinformatics and Biostatistics, School of Life Sciences and Biotechnology, Shanghai Jiao Tong University, 800 Dongchuan Road, Shanghai 200240, China.
2. Department of General Surgery of Ruijin Hospital, Shanghai Institute of Digestive Surgery, and Shanghai Key Laboratory for Gastric Neoplasms, Shanghai Jiao Tong University School of Medicine, 200025, Shanghai, China.
3. SJTU-Yale Joint Center for Biostatistics and Data Science, Shanghai Jiao Tong University, 800 Dongchuan Road, Shanghai 200240, China.
* To whom correspondence should be addressed.
Abstract
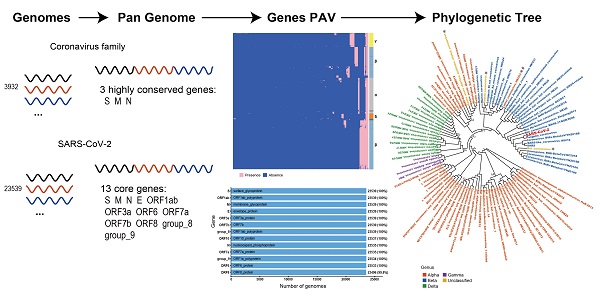
SARS-CoV-2 belongs to the coronavirus family. Comparing genomic features of viral genomes of coronavirus family can improve our understanding about SARS-CoV-2. Here we present the first pan-genome analysis of 3,932 whole genomes of 101 species out of 4 genera from the coronavirus family. We found that a total of 181 genes in the pan-genome of coronavirus family, among which only 3 genes, the S gene, M gene and N gene, are highly conserved. We also constructed a pan-genome from 23,539 whole genomes of SARS-CoV-2. There are 13 genes in total in the SARS-CoV-2 pan-genome. All of the 13 genes are core genes for SARS-CoV-2. The pan-genome of coronaviruses shows a lower level of diversity than the pan-genomes of other RNA viruses, which contain no core gene. The three highly conserved genes in coronavirus family, which are also core genes in SARS-CoV-2 pan-genome, could be potential targets in developing nucleic acid diagnostic reagents with a decreased possibility of cross-reaction with other coronavirus species.
Keywords: COVID-19, SARS-CoV-2, Genome, Diversity, Pangenomics

 Global reach, higher impact
Global reach, higher impact