Impact Factor ISSN: 1449-2288
Int J Biol Sci 2022; 18(3):1303-1312. doi:10.7150/ijbs.66931 This issue Cite
Review
Role of Slit/Robo Signaling pathway in Bone Metabolism
1. State Key Laboratory of Oral Diseases and National Clinical Research Center for Oral Diseases, West China Hospital of Stomatology, Sichuan University, Chengdu 610041, China
2. Department of Conservative Dentistry and Endodontics, West China Hospital of Stomatology, Sichuan University, Chengdu 610041, China
Received 2021-9-8; Accepted 2021-12-26; Published 2022-1-9
Abstract
Slit/Robo signals were initially found to play an essential role in nerve development as axonal guidance molecules. In recent years, with in-depth study, the role of Slit/Robo in other life activities, such as tumor development, angiogenesis, cell migration, and bone homeostasis, has gradually been revealed. Bone is an organ with an active metabolism. Bone resorption and bone formation are closely related through precise spatiotemporal coordination. There is much evidence that slit, as a new bone coupling factor, can regulate bone formation and resorption. For example, Slit3 can promote bone formation and inhibit bone resorption through Robo receptors, which has excellent therapeutic potential in metabolic bone diseases. Although the conclusions of some studies are contradictory, they all affirm the vital role of Slit/Robo signaling in regulating bone metabolism. This paper reviews the research progress of Slit/Robo signaling in bone metabolism, briefly discusses the contradictions in the existing research, and puts forward the research direction of Slit/Robo in the field of bone metabolism in the future.
Keywords: Slit/Robo, Coupling factors, Bone Metabolism, Osteoblast, Osteoclast
Introduction
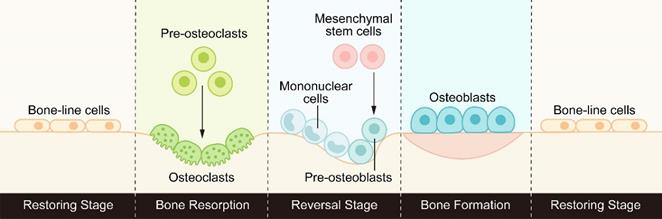
Bone is a highly dynamic tissue and organ that goes through the process of bone formation and bone resorption throughout life to maintain good mechanical properties and maintain the steady state of minerals in the tissue. This process of continuous replacement is called bone remodeling [1]. Bone remodeling usually consists of three consecutive stages: osteoclasts initiate bone resorption, a transition from osteoclast-dominated catabolism to osteoblast-dominated anabolism, and finally osteoblasts form the bone matrix and replace old bone with new bone. (Figure 1).
The process of bone remodeling: osteoclasts initiate bone resorption, then the reverse stage begins under the activation of signal molecules, transforming from bone catabolism to bone synthesis metabolism, and finally osteoblasts form new bone to replace old bone.
In healthy bone, each stage of bone remodeling is subject to precise spatiotemporal regulation to ensure that the absorbed bone is equal to the newly formed bone, maintaining a balance between osteoclasts and osteoblasts. This coordinated relationship between bone resorption and bone formation in time and space is called bone coupling [2, 3]. Since this concept was proposed, the subtle interaction between osteoclasts and osteoblasts has been a hot topic. Understanding the mechanism of bone coupling has important guiding significance for the treatment of metabolic bone diseases caused by bone resorption or steady-state destruction of bone formation, such as osteoporosis and ossification [4].
The growth, development, and repair of bones depend on correct innervation. Neuronal axons and osteoblasts form membrane-membrane contacts, which mediate direct communication between cells through molecules such as glutamate and substance P[5-7]. The physical and functional connection between the nervous system and the skeletal system forms a neuroskeletal network [8]. To date, it has been reported that BMPs [9, 10], CGRP [11, 12], CXCR4/CXCL12 [13, 14], Eph/ephrin [15, 16], and other molecules mediate the crosstalk between bone and the nervous system and participate in the formation of the neuroskeletal network.
In recent years, an increasing number of studies have found that axon guiding molecules such as Sema [17], Netrins [18, 19], and Slit [20, 21] are also members of the neuroskeletal network. They play an essential role in neural development and are expressed in the bone microenvironment to mediate the interaction between osteoblasts and osteoclasts. Sema3A can inhibit the differentiation of osteoclasts by suppressing RhoA and enhance the differentiation of osteoblasts mediated by β-catenin [22]. At the same time, Netrin-1 can promote the activation of RhoA in osteoclasts via UNC5B receptors, which promotes the fusion of osteoclasts and antagonizes the role of Sema3A in the bone microenvironment [23, 24]. The Slit family controls axonal rejection and axonal guidance by binding to Robo receptors, which are essential in nerve development [25]. With the in-depth study of this family, it was found that they are also involved in the regulation of cancer development, cell migration, cell proliferation, and angiogenesis and play an essential role in bone metabolism [26-28]. This article reviews the research progress and future research direction of the Slit/Robo signaling pathway in bone metabolism.
Slit Ligands and Robo Receptors
The Structure and Proteolysis of Slit Ligands
Slit family proteins are highly conserved secretory proteins that were first found in the central nervous system of Drosophila embryos. They are synthesized by midline glial cells and control midline crossover, axon rejection, and axon guidance [29]. Since then, Slit homologs have been found from nematodes to fish, birds, amphibians, and mammals, which are highly conserved among species. There is only one type of slit (Slit1) in invertebrates, while there are three slit genes (Slit1~3) in most vertebrates, which encode approximately 200 kDa [26]. The Slit1 gene is located on human chromosome 10q24.1, and the Slit2 gene is located on 4p15.31. The Slit3 gene is located on 5q34-q35.1 [30].
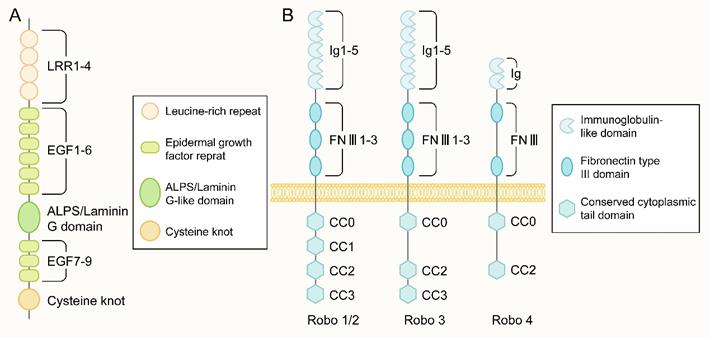
The protein encoded by the Slit gene mainly consists of five structural parts from the N-terminus to the C-terminus: four leucine-rich repeats (LRRs), six epidermal growth factor (EGF) repeat sequences, and a domain found in agrin, laminin, perlecan, and slit named the ALPS or laminin-G-like domain. In addition, there were one to three EGF repeats and one cysteine knot (Figure 2A). The main difference in Slit1-3 lies in the number of EGF repeat sequences. Slit protein can form a homodimer through the reaction between the LRR2 domain and the LRR4 domain [31]. In addition, Slit can bind to other molecules in the extracellular matrix, including Type IV Collagen [32], Netrin 1[33] and dystroglan [34].
The structure of Slit and Robo. (A) The molecular structure of the slit ligand, from the N-terminus to the C-terminus, is mainly composed of five structural parts: four leucine-rich repeats (leucine-rich repeats, LRRs), six epidermal growth factor (epidermal growth factor, EGF) repeats, an ALPS domain or laminin-G-like domain, another three EGF repeats and one cysteine junction. (B) The molecular structure of the Robo receptor protein: Robo1-3 has five Ig-like domains, three fibronectin type III repeat sequences, one transmembrane part, and a long cytoplasmic tail. The cytoplasmic tail of Robo1/2 contains four conserved proline-rich cytoplasmic motifs, called CC0-CC3. In contrast, the cytoplasmic tail of Robo3 has CC0, CC2, and CC3 domains. There are only two Ig-like domains and two FNIII domains in the extracellular part of Robo4 and only CC0 and CC2 domains in the intracellular region.
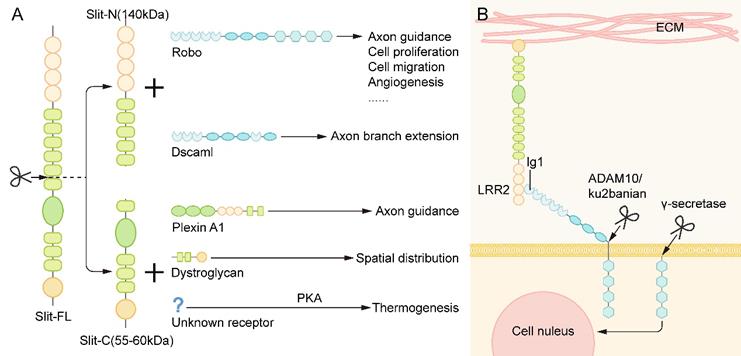
The slit protein has a conserved proteolysis site between the fifth and sixth EGF repeats and can be cleaved by an unknown protease to grow an N-terminal fragment (Slit-N) and short C-terminal fragment (Slit-C) [35]. These different forms of Slit proteins have different functions (Figure 3A).
The Proteolysis of Slit and Robo. (A) The slit protein has a conserved proteolysis site between the fifth and sixth EGF repeats and is cleaved by an unknown protease to grow the N-terminal fragment (Slit-N) and short C-terminal piece (Slit-C). They can bind to different receptors to perform various functions. (B) The binding of the Slit protein fixed to the extracellular matrix to the Robo on the cell membrane results in the exposure of the cleavage site of metalloproteinases in the area adjacent to the membrane of Robo protein. Under the action of Kuzbanian in Drosophila (ADAM10 in mammals), the Robo protein is cleaved, and its extracellular domain falls away. In human cancer cells, the lagged residues of the extracellular domain of Robo are further cleaved by γ secretase, and the resulting C-terminal fragments can then be transferred to the nucleus.
Receptors of Slit Ligands
Robo family proteins are the main receptors of Slit ligands. In the majority of tissue microenvironments, they combine with each other to transmit cellular signals. Both full-length Slit (Slit-FL) and Slit-N can bind to Robo receptors and regulate axonal guidance, cell proliferation, cell migration, angiogenesis, and other life activities [26]. However, the interaction between Slit and Robo is not unique. Several novel receptors of Slit have been found in recent studies.
Dan et al. [36] found that Slit-N can directly bind to the N-terminal Ig domains of Dscam1, which enhances the interaction between Dscam1 and receptor tyrosine phosphatase 69D (RPTP69D) and stimulates the dephosphorylation of Dscam1. It does not depend on Robo to promote the extension of specific axon collaterals.
Slit-C is mainly released into the extracellular space. PlexinA1, a novel receptor of Slit, can bind to the C-terminal fragment of Slit specifically and mediate commissural growth cone collapse independently of the Robo. Slit/PlexinA1 and Slit/Robo signaling play a complementary role in the guidance of commissural axons [37].
In addition, Kevin et al. [38] found that the C-terminus of Slit can bind to the floor plate and basement membrane scaffold protein Dystroglycan to regulate the spatial distribution of cells. In mutant mice with dystroylycan deletion, the distribution of endogenous Slit was wrong and led to developmental abnormalities. The combination of Slit and Dystroglycan is an important factor affecting the correct distribution of Slit in vivo.
A recent study [39] has shown that the C-terminal fragment of Slit2 can activate PKA-dependent signaling pathways to regulate adipose tissue thermogenesis and improve blood glucose homeostasis, which shows therapeutic potential for the treatment of obesity. However, the exact receptor of Slit2-C in this signaling pathway remains unclear.
The Structure and Proteolysis of Robo Receptors
The most important receptor of the Slit family is the Robo protein, which is a type of transmembrane receptor protein composed of 1000 to 1600 amino acids that has a highly conserved intracellular domain. There are three Robo receptors (Robo1-3) in Drosophila, chickens, and Xenopus laevis and four Robo receptors (Robo1-4) in zebrafish and mammals [40, 41]. The Robo1 and Robo2 genes are located on human chromosome 3p12.3, and the Robo3 and Robo4 genes are located on human chromosome 11q24.2 [42].
The Robo family is a family of immunoglobulin (immunoglobulin, Ig) superfamilies. Robo1-3 has five Ig-like domains, three fibronectin type III repeats, a transmembrane part, and a long cytoplasmic tail (Figure 2B). The cytoplasmic tail of Robo1 and Robo2 contains four conserved proline-rich cytoplasmic motifs, while the cytoplasmic tail called CC0-CC3; Robo3 contains CC0, CC2, and CC3 domains. Robo4 was initially considered a vascular endothelial cell-specific receptor of the Robo family[41], with only two Ig-like domains and two FNⅢ domains in the extracellular part and only CC0 and CC2 domains in the intracellular part(Figure 2B). A recent study [43] showed that Robo4 also exists in the newborn cerebral cortex to regulate the directional migration of newborn neurons. No apparent catalytic region was found in the Robo family proteins, mainly through the CC0-CC3 domain in the cytoplasm to bind to various intracellular signal molecules to mediate signal transmission.
Robo proteins can also be cleaved, which seems to be essential for the signal transduction of the Slit/Robo pathway into the cell. The binding of the Slit protein, which is fixed to the extracellular matrix, to the Robo on the cell membrane results in the exposure of the cleavage site of metalloproteinases in the area adjacent to the membrane of Robo protein. Under the action of Kuzbanian in Drosophila (ADAM10 in mammals), the Robo protein is cleaved, and its extracellular domain falls away [44, 45] (Figure 3B). Some studies have found that the cleavage of Robo receptors is necessary to recruit downstream signaling molecules. In human cancer cells, the residue of the extracellular domain of Robo is further cleaved by γ secretase, and the resulting C-terminal fragment can then be transferred to the nucleus. However, the specific function of this fragment has not been determined [45] (Figure 3B). Nevertheless, it remains unclear whether the Robo of all vertebrates will exfoliate extracellular domains and play a role in the development.
Ligands of Robo Receptors
Slit protein is considered the only ligand of Robo receptors. The second leucine-rich repeat (LRRD2) of the Slit protein is its active site, which binds to the active site in the first Ig domain of Robo1 and Robo2, thus transducing the extracellular signal into the cell. Robo3 and Robo4 are two unique receptors in the Robo family. Due to some amino acid changes in its first Ig domain, Robo3 almost lost its ability to bind Slit protein [46]. However, Robo3 can acquire new signaling properties simultaneously, such as NELL ligands.
NELL family proteins are newly discovered ligand proteins of Robo2/3 and cannot bind to Robo1/4. However, because the extracellular hairpin-like domain of Robo2 may mask the binding site of NELL1/2, NELL1/2 can bind to Robo2 only after the conformational change of the extracellular domain of Robo2 under mild acidic conditions [47, 48]. Both NELL1 and 2 can bind to Robo3. NELL2 can antagonize the rejection induced by Slit2-Robo1/2 and allow the commissural axon to cross the midline by binding to the Robo3 receptor [49]. Although NELL1 does not play an important role in this, it may become an effective ligand of Robo3 in other life activities [50].
Robo4 is believed to be specifically expressed in endothelial cells [28], and thus far, there have been no reports of direct interaction between Robo4 and Slit2/3. Some researchers [51-53] have proposed that Robo4 transmits signals by forming coreceptors with other molecules, such as Robo1 and heparan sulfate proteoglycans (HSPGs). The current research on Robo3 and Robo4 is imperfect. Whether they bind to Slit ligands and regulate cell activity remains to be determined.
The relationship between Slit/Robo signaling and other relevant ligands or receptors forms a complex regulatory network, which not only increases its richness and diversity but also provides a sufficient theoretical basis for its role in many life activities.
Slit-Robo Signaling in Bone Metabolism
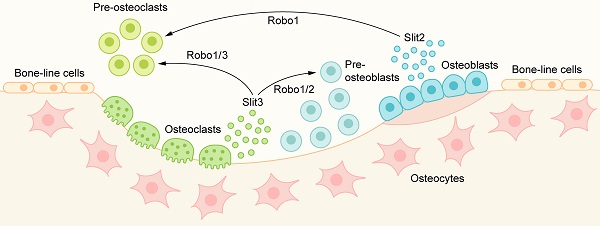
Bone remodeling is performed by a structure called basic multicellular units (BMUs), which are mainly composed of two types of cells: osteoblasts and osteoclasts [54, 55]. The whole stage of bone remodeling is composed of catabolism of osteoclasts and synthetic metabolism of osteoblasts: it begins with the activation of bone resorption of osteoclasts. After the completion of the resorption stage, preosteoblasts are recruited to the bone surface, differentiate and mature, begin the synthesis of bone matrix, and form new bone to replace old bone. The signaling molecules that coordinate the bone resorption of osteoclasts and the bone formation of osteoblasts in BMUs are called bone coupling factors. The slit protein family, the classical axon guide molecule, is a recently discovered bone coupling factor that plays a vital role in regulating bone metabolism [2]. The expression of Slit1 is the lowest in bone tissue, while Slit2 and Slit3 are expressed in both osteoblasts and osteoclasts, although the expression level may vary according to the state of cells [56, 57]. Robo1 and Robo3 are expressed in osteoclasts, Robo1 and Robo2 are expressed in osteoblasts, and Robo4 is hardly expressed in osteoblasts [58]. In recent years, there have been an increasing number of studies on the role of Slit2 and Slit3 in bone metabolism, and the regulatory role of Slit2 and Slit3 in osteoblasts and osteoclasts is gradually revealed.
The Effect of Slit2 on osteoblasts and osteoclasts
The occurrence of osteoclasts is mainly carried out through the proliferation, migration, and fusion of preosteoclasts and the subsequent differentiation of osteoclasts. Slit2 plays a role in the early stage of osteoclast development, which is directly related to the Robo1 receptor on preosteoclasts and inhibits osteoclast differentiation [59] (Figure 4). In the presence of Slit2, the expression of osteoclast surface differentiation markers, such as tartrate-resistant acid phosphatase (TRAP) and calcitonin receptor (CTR), decreased significantly. Slit2 also inhibited the activity of the small GTPase Cdc42 but had no significant effect on the activity of Rac and RhoA, resulting in a significant decrease in the migration and fusion of preosteoclast cells and resulting in the inhibition of osteoclast formation [59].
The role of Slit/Robo signaling in metabolic bone coupling: Slit2/3 acts on osteoclasts through the Robo1/3 receptor to inhibit the migration and fusion of osteoclasts, resulting in the inhibition of osteoclast formation. Slit3 can also bind to the Robo1/2 receptor on osteoblasts and promote the migration and proliferation of osteoblast lines, thus promoting bone formation.
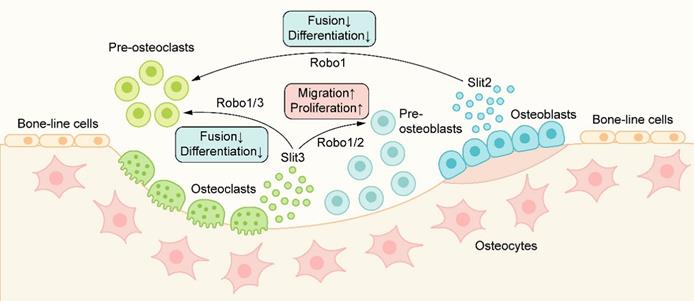
A previous study [56] pointed out that Slit2 inhibited the differentiation of osteoblast lines mainly through Robo receptors and inhibited the alkaline phosphatase activity of rat osteoblasts and MC3T3 cell lines in a concentration-dependent manner. Furthermore, Slit2 changed the cell morphology from flattened to elongated but had no significant effect on the proliferation of osteoblasts. However, a recent study [59] showed that the treatment of osteoblasts with recombinant Slit2 could not change the activity, migration ability, osteoblast differentiation markers, or alkaline phosphatase activity of osteoblast lines. The reason for this contradictory result may be that the conditioned medium for the treatment of osteoblasts in the previous study contains not only inexactly identified Slit2 but also other molecules that affect the biology of osteoblasts. In addition, cell type, cell microenvironment, treatment time, treatment conditions, and other factors may affect the effectiveness of the Slit/Robo pathway, which may also be one of the explanations for the contradictory phenomena in different studies.
The Effect of Slit3 on osteoblasts and osteoclasts
In recent years, Slit3 has been shown to be a new bone coupling factor. Kim et al. [58] found that Slit3 derived from osteoclasts acts on osteoblasts in the form of paracrine signaling to promote bone formation and acts on osteoclasts in the form of autocrine signaling to inhibit bone resorption(Figure 4). Slit3 interacts with Robo1 and Robo3 receptors on osteoclasts/preosteoclasts, which inhibits the small GTPase Rac1. In addition, Slit3 also reduces the expression of DC-STAMP in preosteoclasts, and DC-STAMP plays an important role in the fusion of preosteoclasts. Slit3 can inhibit the formation of osteoclasts and the bone resorption of osteoclasts by mediating the disturbance of migration and fusion of preosteoclasts and reducing the expression of osteoclast differentiation markers such as TRAP and CTR, but Slit3 does not affect the proliferation of preosteoclasts [58]. In osteoblasts, Slit3 binds to Robo1 and Robo2 receptors on the cell membrane, activates β-catenin, stimulates the migration and proliferation of osteoblast lines in a concentration-dependent manner, and promotes bone formation [27, 58]. In the mice with osteoclast-specific deletion of Slit3, the bone formation parameters decreased while the bone resorption parameters increased, which showed a decrease in bone mass. In contrast, bone mass was normal in mice with osteoblast-specific deletion of Slit3 or neuron-specific deletion of normal [58]. These results suggest that osteoclast-derived Slit3 may be a stop signal of bone resorption during bone remodeling and initiate the reversal stage of osteoblast bone formation.
However, Xu et al.[60] found that Slit3 was strongly expressed in osteoblasts but hardly expressed in osteoclasts; similarly, osteoblast-specific SLIT3-deficient mice showed a decrease in bone mass, while osteoclast-specific Slit3-deficient mice showed a normal bone mass, and they believed that osteoblasts were the primary source of Slit3 in bone. The results of Li et al. [61] also supported this finding that there was no significant effect on the number, differentiation, bone resorption/bone formation parameters, or total bone mass of osteoclasts in the absence of Slit3 in osteoclasts.
We suspect that the differences in the above experimental results may be partly due to the differences in tissue-specific clones used to produce conditional gene knockout mice or the existence of other unknown compensatory mechanisms in osteoclasts. Although the primary source of Slit3 in bone and its role in osteocytes are still under debate, it is certain that Slit3 plays an essential role as a bone coupling factor in bone metabolism, and further research is needed to better understand and clarify the role of Slit3-Robo signaling in bone metabolism.
Downstream of Slit-Robo Signaling in Bone Metabolism
Small GTPases of the Rho family, including RhoA, Cdc42, and Rac1, play an essential role in the downstream reaction of Slit/Robo signaling. Osteoclasts are responsible for reorganizing the cytoskeleton, which is essential for cell movement, and promoting the occurrence and development of specialized structures needed for bone resorption [62, 63]. After the activation of the Slit/Robo signaling pathway, Slit-Robo GTPase activating proteins (srGAPs) are recruited to the intracellular domain of the Robo receptor, resulting in the inactivation of Rho GTPase, thus inhibiting cell actin polymerization and stress fiber formation and affecting cell polarity and cell movement [42, 64] (Figure 5A). However, different Slit family members have different inhibition of the Rho GTPase. Slit2 mainly inhibits the Cdc42 GTPase [59], while Slit3 mainly inhibits the Rac1 GTPase [58]. There is also a feedback loop in osteoclasts. Activated Rac1 can stimulate an increase in Slit3 expression. Slit3 acts on osteoclasts in an autocrine way, forms a complex with Robo1 and recruits srGAP2, and inhibits the activation of Rac1 [64] (Figure 5B).
Downstream of Slit-Robo Signaling in Bone Metabolism. (A) When Slit binds to Robo, Slit-Robo GTPase activating protein (srGAP) is recruited to the intracellular domain of the Robo receptor, resulting in inactivation of Rho GTPase, which inhibits actin polymerization and stress fiber formation and affects cell polarity and cell movement. Slit2 mainly inhibits Cdc42 GTPase, while Slit3 mainly inhibits Rac1 GTPase. (B) In osteoclasts, activated Rac1 can stimulate the expression of Slit3. Slit3 acts on osteoclasts in an autocrine way, binds with Robo1 and recruits srGAP2 to form a complex, and inhibits the activation of Rac1 to form a negative feedback loop. (C) After binding to the Slit ligand, the Robo protein forms a multimolecular complex with Abl kinase, adaptor protein cables, N-cadherin and β-catenin. Then, Abl kinase phosphorylates β-catenin on tyrosine 489, resulting in the release of N-cadherin-related β-catenin, leading to the loss of N-cadherin-mediated cell adhesion. The released β-catenin can also enter the nucleus to activate target genes and regulate the life activities of cells.
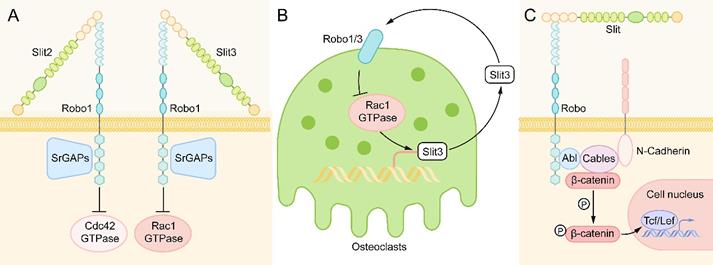
In addition to the Rho GTPase family, another major signaling molecule downstream of Robo receptors is cytoplasmic kinases, in which Abelson (Abl) tyrosine kinase is the main participant [26]. After binding to the Slit ligand, the Robo protein forms a multimolecular complex with Abl kinase, adaptor protein cables (Cdk5 and Abl enzyme substrate), N-cadherin, and β-catenin. Then, Abl kinase phosphorylates β-catenin on tyrosine 489, resulting in the release of N-cadherin-related β-catenin, which leads to the loss of N-cadherin-mediated cell adhesion [65, 66]. The released β-catenin can also enter the nucleus to activate target genes and regulate the life activities of cells [27] (Figure 5C).
Wnt/β-catenin plays an important role in the control of bone development. Slit3 has been shown to promote chondrocyte differentiation and regulate endochondral ossification by binding to the Robo2 receptor to inhibit the activity of β-catenin in chondrocytes [67], which is contrary to its effect on osteoblasts. Slit3 binds to the Robo1/2 receptor to activate β-catenin in osteoblasts to stimulate osteoblast proliferation and migration [58]. The different effects of Slit3 on the activity of β-catenin are related to the cell type. Moreover, we speculate that it is one of the reasons that different regions of the Abl tyrosine kinase have different effects on Robo receptors. Some studies [26, 66] have found that Abl kinase can inhibit Robo signaling by phosphorylating the CC1 domain of Robo, and it can also promote Robo signaling by binding to Capulet protein or Cables protein. However, in different cells, the specific mechanism of regulating the effect of Abl kinase on Robo signaling has not been reported clearly.
The Interaction of Slit-Robo Signaling in Angiogenesis and Bone Metabolism
In the skeletal system of mammals, angiogenesis plays an essential role in maintaining the dynamic balance of bone remodeling. Blood vessels can not only provide necessary nutrients, oxygen, growth factors, and hormones for bone tissue but also secrete signaling molecules to regulate bone formation [68, 69]. This close relationship between angiogenesis and bone formation in time and space is called "angiogenesis-osteogenesis coupling. " [70]
In recent studies [60], a capillary subtype, called H-type blood vessels, was significantly related to osteogenesis, which is characterized by the high expression of CD31 and Emcn on endothelial cells. H-type blood vessels are mainly located in the metaphysis and surrounded by dense Osterix+ bone progenitor cells. H-type blood vessels can guide bone formation by producing factors that stimulate the proliferation and differentiation of bone progenitor cells [71] (Figure 6).
Slit/Robo in angiogenesis-osteogenesis coupling: H-type blood vessels are surrounded by dense Osterix+ bone progenitor cells. H-type blood vessels can guide bone formation by stimulating the proliferation and differentiation of bone progenitor cells. Osteoblast-derived Slit3, as an angiogenic factor, can produce an increase in CD31hiEmcnhi endothelium. Hence, Slit3 indirectly promotes bone formation and couples the process of bone metabolism with angiogenesis.
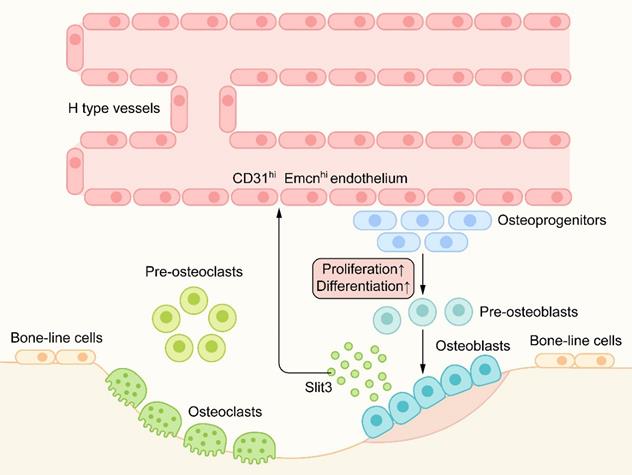
Robo1, Robo2, and Robo4 are expressed in all types of vascular endothelial cells. Slit2/3 can regulate angiogenesis by combining with the Robo1 or Robo1/Robo4 heterodimer [42, 51]. Osteoblast-derived Slit3 has been shown to increase the number of H-type vascular endothelial cells, couple the process of bone formation with angiogenesis, and indirectly promote bone formation as a pro-angiogenic factor [72] (Figure 6). Mice lacking Slit3 or Robo1 showed a significant decrease in H-type endothelial cells and bone mass. In a mouse fracture model, recombinant Slit3 significantly increased the number of H-type blood vessels and promoted fracture healing [60]. These results suggest that Slit3/Robo1 is a strong regulatory signal for inducing H-type angiogenesis and promoting bone formation and plays an important role in the angiogenesis-osteogenesis coupling.
Interestingly, some studies [42] have shown that Slit/Robo signaling can not only promote angiogenesis through some receptors but also inhibit angiogenesis through other receptors. For example, Slit2/Robo1 can promote tumor angiogenesis [73], and Slit2/Robo4 can inhibit tumor angiogenesis [74, 75]. To date, the research on the role of Slit/Robo signaling in angiogenesis is more focused on the direction of tumor development. However, few studies are related to bone development or metabolic bone balance. Whether Slit2 also plays a regulatory role in angiogenesis-osteogenesis coupling, such as Slit3 is still an interesting question worth exploring.
Conclusion and Perspectives
Slit and its main receptor Robo were first found in the study of nerve development and are considered classical axon guide molecules. With the in-depth understanding of Slit/Robo signaling, its role in other fields, such as tumor development, angiogenesis, bone development, and inflammatory regulation, has been gradually discovered. In recent years, an increasing number of studies have shown that Slit is a novel bone coupling factor that can regulate bone resorption and bone formation at the same time. In addition, because of the angiogenic effect of Slit, H-type blood vessels are associated with bone metabolism via Slit. The application of Slit in the treatment of metabolic bone diseases is also being explored; for instance, the injection of truncated recombinant Slit3 in mice can significantly save bone loss.
However, numerous unanswered questions remain regarding the role of the Slit/Robo signal in bone metabolism. For example, is the main source of Slit3 in the skeletal microenvironment from osteoblasts or osteoclasts? Does Slit2 have any effect on osteoblast biology? The Slit/Robo signaling pathway has various downstream signals in different spatiotemporal environments, resulting in the complexity of Slit/Robo signals. This makes it more difficult to clarify the specific mechanism of Slit/Robo signaling in bone metabolism.
According to the current research results, Slit shows good application prospects and therapeutic potential in the field of bone metabolism. Therefore, it is necessary to further explore and clarify the molecular mechanism of Slit/Robo in regulating bone homeostasis, which is of great significance to improve the molecular regulatory network of bone metabolism and explain the contradictory results reported in various existing studies. Simultaneously, fully exploiting the therapeutic potential of Slit family proteins in metabolic bone diseases and improving their clinical applicability are promising future research directions.
Abbreviations
BMP: Bone morphogenetic protein; CGRP: Calcitonin gene related peptide; CXCR4/CXCL12: C-X-C motif chemokine receptor 4/C-X-C motif chemokine ligand 12; RhoA: Ras homolog family member A; Rac: Ras-related C3 botulinum toxin substrate; UNC5B: Unc-5 Netrin Receptor B; LRR: Leucine-rich repeats; EGF: Epidermal growth factor; Ig: Immunoglobulin; FNⅢ: Fibronectin type Ⅲ; CC: Cytoplasmic tail; Dscam1: Down syndrome cell adhesion molecule 1; PKA: Protein kinase A; ADAM10: A Disintegrin and metalloproteinase domain-containing protein 10; HSPGs: Heparan sulfate proteoglycans; BMUs: Basic multicellular units; TRAP: Tartrate-resistant acid phosphatase; CTR: Calcitonin receptor; Cdc42: Cell division control protein 42 homolog; srGAP: Slit-Robo GTPase activating protein; Abl: Abelson.
Acknowledgements
Author Contributions
Lingyu Jiang took part in the conception of this review, drew schematic figures and drafted the article.
Jianxun Sun and Dingming Huang guided this project, critically revised the article for important intellectual content and performed the final approval of the version to be submitted.
Funding
This work was supported by the National Natural Science Foundation of China (No.81300847 [J.X.S] and No.81771063 [D.M.H]).
Competing Interests
All authors declare that there are no competing interests associated with the manuscript.
References
1. Zaidi M. Skeletal remodeling in health and disease. Nat Med. 2007;13(7):791-801
2. Kim B, Koh J. Coupling factors involved in preserving bone balance. Cell Mol Life Sci. 2019;76(7):1243-1253
3. Sims NA, Martin TJ. Osteoclasts Provide Coupling Signals to Osteoblast Lineage Cells Through Multiple Mechanisms. Annu Rev Physiol. 2020;82(1):507-529
4. Feng X, McDonald JM. Disorders of Bone Remodeling. Annual Review of Pathology: Mechanisms of Disease. 2011;6(1):121-145
5. Grässel S, Muschter D. Peripheral Nerve Fibers and Their Neurotransmitters in Osteoarthritis Pathology. Int J Mol Sci. 2017;18(5):931
6. Kodama D, Hirai T, Kondo H. et al. Bidirectional communication between sensory neurons and osteoblasts in anin vitro coculture system. Febs Lett. 2017;591(3):527-539
7. Nencini S, Ivanusic JJ. The Physiology of Bone Pain. How Much Do We Really Know? Front Physiol. 2016;7:157
8. Abeynayake N, Arthur A, Gronthos S. Crosstalk between skeletal and neural tissues is critical for skeletal health. Bone. 2021;142:115645
9. He J, Zheng X, Jiang S. et al. Sympathetic neuron can promote osteoblast differentiation through BMP signaling pathway. Cell Signal. 2013;25(6):1372-1378
10. Pederson L, Ruan M, Westendorf JJ. et al. Regulation of Bone Formation by Osteoclasts Involves Wnt/BMP Signaling and the Chemokine Sphingosine-1-Phosphate. Proceedings of the National Academy of Sciences - PNAS. 2008;105(52):20764-20769
11. Sang X, Wang Z, Shi P. et al. CGRP accelerates the pathogenesis of neurological heterotopic ossification following spinal cord injury. Artif Cells Nanomed Biotechnol. 2019;47(1):2569-2574
12. Iyengar S, Ossipov MH, Johnson KW. The role of calcitonin gene-related peptide in peripheral and central pain mechanisms including migraine. Pain. 2017;158(4):543-559
13. Zhu W, Liang G, Huang Z. et al. Conditional Inactivation of the CXCR4 Receptor in Osteoprecursors Reduces Postnatal Bone Formation Due to Impaired Osteoblast Development. J Biol Chem. 2011;286(30):26794-26805
14. Arthur A, Shi S, Zannettino ACW. et al. Implanted Adult Human Dental Pulp Stem Cells Induce Endogenous Axon Guidance. Stem Cells. 2009;27(9):2229-2237
15. Croteau LP, Kao TJ, Kania A. Ephrin-A5 potentiates netrin-1 axon guidance by enhancing Neogenin availability. Sci Rep-Uk. 2019;9(1):12009
16. Nguyen TM, Arthur A, Paton S. et al. Loss of ephrinB1 in osteogenic progenitor cells impedes endochondral ossification and compromises bone strength integrity during skeletal development. Bone. 2016;93:12-21
17. Deb Roy A, Yin T, Choudhary S. et al. Optogenetic activation of Plexin-B1 reveals contact repulsion between osteoclasts and osteoblasts. Nat Commun. 2017;8(1):15831
18. Zhu S, Zhu J, Zhen G. et al. Subchondral bone osteoclasts induce sensory innervation and osteoarthritis pain. J Clin Invest. 2019;129(3):1076-1093
19. SATO T, KOKABU S, ENOKI Y. et al. Functional Roles of Netrin-1 in Osteoblast Differentiation. In vivo. 2017;31(3):321-328
20. J I, T Y, SM K. et al. Opening windows for bone remodeling through a SLIT. The Journal of clinical investigation. 2018;128(4):1255-1257
21. B C, L C, XP D. Dynamic expression of Slit1-3 and Robo1-2 in the mouse peripheral nervous system after injury. Neural Regen Res. 2020;15(5):948-958
22. Hayashi M, Nakashima T, Taniguchi M. et al. Osteoprotection by semaphorin 3A. Nature. 2012;485(7396):69-74
23. Maruyama K, Kawasaki T, Hamaguchi M. et al. Bone-protective Functions of Netrin 1 Protein. J Biol Chem. 2016;291(46):23854-23868
24. Mediero A, Ramkhelawon B, Perez-Aso M. et al. Netrin-1 Is a Critical Autocrine/Paracrine Factor for Osteoclast Differentiation. J Bone Miner Res. 2015;30(5):837-854
25. Dun X, Parkinson D. Classic axon guidance molecules control correct nerve bridge tissue formation and precise axon regeneration. Neural Regen Res. 2020;15(1):6-9
26. Blockus H, Chédotal A. Slit-Robo signaling. Development. 2016;143(17):3037-3044
27. JM K. Osteoclast-derived SLIT3 is a coupling factor linking bone resorption to bone formation. Bmb Rep. 2018;51(6):263-264
28. Dai C, Gong Q, Cheng Y. et al. Regulatory mechanisms of Robo4 and their effects on angiogenesis. Bioscience Rep. 2019 39(7)
29. Simpson JH, Kidd T, Bland KS. et al. Short-Range and Long-Range Guidance by Slit and Its Robo Receptors: Robo and Robo2 Play Distinct Roles in Midline Guidance. Neuron. 2000;28(3):753-766
30. Katoh Y, Katoh M. Comparative genomics on SLIT1, SLIT2, and SLIT3 orthologs. Oncol Rep. 2005;14(5):1351-1355
31. Seiradake E, von Philipsborn AC, Henry M. et al. Structure and functional relevance of the Slit2 homodimerization domain. Embo Rep. 2009;10(7):736-741
32. T X, W S, E R. et al. Assembly of lamina-specific neuronal connections by slit bound to type IV collagen. Cell. 2011;146(1):164-176
33. Brose K, Bland KS, Wang KH. et al. Slit proteins bind Robo receptors and have an evolutionarily conserved role in repulsive axon guidance. Cell. 1999;96(6):795-806
34. KM W, KA L, H L. et al. Dystroglycan organizes axon guidance cue localization and axonal pathfinding. Neuron. 2012;76(5):931-944
35. Kellermeyer R, Heydman LM, Gillis T. et al. Proteolytic cleavage of Slit by the Tolkin protease converts an axon repulsion cue to an axon growth cuein vivo. Development. 2020 147
36. Dascenco D, Erfurth M, Izadifar A. et al. Slit and Receptor Tyrosine Phosphatase 69D Confer Spatial Specificity to Axon Branching via Dscam1. Cell. 2015;162(5):1140-1154
37. Delloye-Bourgeois C, Jacquier A, Charoy C. et al. PlexinA1 is a new Slit receptor and mediates axon guidance function of Slit C-terminal fragments. Nat Neurosci. 2015;18(1):36-45
38. Wright KM, Lyon KA, Leung H. et al. Dystroglycan organizes axon guidance cue localization and axonal pathfinding. Neuron. 2012;76(5):931-944
39. Svensson KJ, Long JZ, Jedrychowski MP. et al. A Secreted Slit2 Fragment Regulates Adipose Tissue Thermogenesis and Metabolic Function. Cell Metab. 2016;23(3):454-466
40. Connor RM, Key B. Expression and role of Roundabout-1 in embryonicXenopus forebrain. Dev Dynam. 2002;225(1):22-34
41. Park KW, Morrison CM, Sorensen LK. et al. Robo4 is a vascular-specific receptor that inhibits endothelial migration. Dev Biol. 2003;261(1):251-267
42. Tong M, Jun T, Nie Y. et al. The Role of the Slit/Robo Signaling Pathway. J Cancer. 2019;10(12):2694-2705
43. Zheng W, Geng A, Li P. et al. Robo4 Regulates the Radial Migration of Newborn Neurons in Developing Neocortex. Cereb Cortex. 2012;22(11):2587-2601
44. Coleman HA, Labrador J, Chance RK. et al. The Adam family metalloprotease Kuzbanian regulates the cleavage of the roundabout receptor to control axon repulsion at the midline. Development. 2010;137(14):2417-2426
45. Seki M, Watanabe A, Enomoto S. et al. Human ROBO1 is cleaved by metalloproteinases and γ-secretase and migrates to the nucleus in cancer cells. Febs Lett. 2010;584(13):2909-2915
46. Zelina P, Blockus H, Zagar Y. et al. Signaling Switch of the Axon Guidance Receptor Robo3 during Vertebrate Evolution. Neuron. 2014;84(6):1258-1272
47. Yamamoto N, Kashiwagi M, Ishihara M. et al. Robo2 contains a cryptic binding site for neural EGFL-like (NELL) protein 1/2. J Biol Chem. 2019;294(12):4693-4703
48. Barak R, Yom-Tov G, Guez-Haddad J. et al. Structural Principles in Robo Activation and Auto-inhibition. Cell. 2019;177(2):272-285
49. Jaworski A, Tom I, Tong RK. et al. Operational redundancy in axon guidance through the multifunctional receptor Robo3 and its ligand NELL2. Science (American Association for the Advancement of Science). 2015;350(6263):961-965
50. Pak JS, DeLoughery ZJ, Wang J. et al. NELL2-Robo3 complex structure reveals mechanisms of receptor activation for axon guidance. Nat Commun. 2020;11(1):1489
51. Rama N, Dubrac A, Mathivet T. et al. Slit2 signaling through Robo1 and Robo2 is required for retinal neovascularization. Nat Med. 2015;21(5):483-491
52. Li S, Huang L, Sun Y. et al. Slit2 Promotes Angiogenic Activity Via the Robo1-VEGFR2-ERK1/2 Pathway in Both In vivo and In vitro Studies. Invest Ophth Vis Sci. 2015;56(9):5210-5217
53. Fukuhara N, Howitt JA, Hussain S. et al. Structural and Functional Analysis of Slit and Heparin Binding to Immunoglobulin-like Domains 1 and 2 of Drosophila Robo. J Biol Chem. 2008;283(23):16226-16234
54. Sims NA, Martin TJ. Osteoclasts Provide Coupling Signals to Osteoblast Lineage Cells Through Multiple Mechanisms. Annu Rev Physiol. 2020;82:507-529
55. Kenkre JS, Bassett J. The bone remodelling cycle. Annals of Clinical Biochemistry: International Journal of Laboratory Medicine. 2018;55(3):308-327
56. Sun H, Dai K, Tang T. et al. Regulation of Osteoblast Differentiation by Slit2 in Osteoblastic Cells. Cells Tissues Organs. 2009;190(2):69-80
57. Zhang B, Dietrich UM, Geng JG. et al. Repulsive axon guidance molecule Slit3 is a novel angiogenic factor. Blood. 2009;114(19):4300-4309
58. Kim BJ, Lee YS, Lee SY. et al. Osteoclast-secreted SLIT3 coordinates bone resorption and formation. J Clin Invest. 2018;128(4):1429-1441
59. Park SJ, Lee JY, Lee SH. et al. SLIT2 inhibits osteoclastogenesis and bone resorption by suppression of Cdc42 activity. Biochem Biophys Res Commun. 2019;514(3):868-874
60. Xu R, Yallowitz A, Qin A. et al. Targeting skeletal endothelium to ameliorate bone loss. Nat Med. 2018;24(6):823
61. Li N, Inoue K, Sun J. et al. Osteoclasts are not a source of SLIT3. Bone Res. 2020;8(1):11
62. Croke M, Ross FP, Korhonen M. et al. Rac deletion in osteoclasts causes severe osteopetrosis. J Cell Sci. 2011;124(Pt 22):3811-3821
63. Ito Y, Teitelbaum SL, Zou W. et al. Cdc42 regulates bone modeling and remodeling in mice by modulating RANKL/M-CSF signaling and osteoclast polarization. J Clin Invest. 2010;120(6):1981-1993
64. Shin B, Kupferman J, Schmidt E. et al. Rac1 Inhibition Via Srgap2 Restrains Inflammatory Osteoclastogenesis and Limits the Clastokine, SLIT3. J Bone Miner Res. 2020;35:789-800
65. Koohini Z, Koohini Z, Teimourian S. Slit/Robo Signaling Pathway in Cancer; a New Stand Point for Cancer Treatment. Pathol Oncol Res. 2019;25(4):1285-1293
66. Rhee J, Buchan T, Zukerberg L. et al. Cables links Robo-bound Abl kinase to N-cadherin-bound β-catenin to mediate Slit-induced modulation of adhesion and transcription. Nat Cell Biol. 2007;9(8):883-892
67. Kim H, Choi YJ, Lee YS. et al. SLIT3 regulates endochondral ossification by beta-catenin suppression in chondrocytes. Biochem Biophys Res Commun. 2018;506(4):847-853
68. Gallardo-Calero I, Barrera-Ochoa S, Manzanares MC. et al. Vascularized Periosteal Flaps Accelerate Osteointegration and Revascularization of Allografts in Rats. Clinical Orthopaedics & Related Research. 2019;477(4):741-755
69. Zheng Z, Chen Y, Wu D. et al. Development of an Accurate and Proactive Immunomodulatory Strategy to Improve Bone Substitute Material-Mediated Osteogenesis and Angiogenesis. Theranostics. 2018;8(19):5482-5500
70. Grosso A, Burger MG, Lunger A. et al. It Takes Two to Tango: Coupling of Angiogenesis and Osteogenesis for Bone Regeneration. Frontiers in Bioengineering and Biotechnology. 2017;5:68
71. Peng Y, Wu S, Li Y. et al. Type H blood vessels in bone modeling and remodeling. Theranostics. 2020;10(1):426-436
72. Zhang J, Pan J, Jing W. Motivating role of type H vessels in bone regeneration. Cell Proliferat. 2020;53(9):e12874
73. Zhang X, Dong J, He Y. et al. miR-218 inhibited tumor angiogenesis by targeting ROBO1 in gastric cancer. Gene. 2017;615:42-49
74. Romano E, Manetti M, Rosa I. et al. Slit2/Robo4 axis may contribute to endothelial cell dysfunction and angiogenesis disturbance in systemic sclerosis. Ann Rheum Dis. 2018;77(11):1665-1674
75. Zhao H, Ahirwar DK, Oghumu S. et al. Endothelial Robo4 suppresses breast cancer growth and metastasis through regulation of tumor angiogenesis. Mol Oncol. 2016;10(2):272-281
Author contact
Corresponding authors: Jianxun Sun; E-mail: jxsunedu.cn; Tel: 86-28-85503469; Dingming Huang; E-mail: dingminghuangcom; Tel: 86-28-85501439

 Global reach, higher impact
Global reach, higher impact