Impact Factor ISSN: 1449-2288
Int J Biol Sci 2022; 18(6):2292-2303. doi:10.7150/ijbs.69411 This issue Cite
Review
The role of ubiquitination and deubiquitination in tumor invasion and metastasis
1. Department of Radiology, The Third Xiangya Hospital of Central South University, Changsha, Hunan 410013, China
2. Cell Transplantation and Gene Therapy Institute, The Third Xiangya Hospital, Central South University, Changsha, Hunan, 410013, China
3. Department of Ultrasonography, The Third Xiangya Hospital of Central South University, Changsha, Hunan, 410013, China
Received 2021-11-24; Accepted 2022-2-19; Published 2022-3-6
Abstract
Ubiquitination is vital for multiple cellular processes via dynamic modulation of proteins related to cell growth, proliferation, and survival. Of the ubiquitination system components, E3 ubiquitin ligases and deubiquitinases have the most prominent roles in modulating tumor metastasis. This review will briefly summarize the observations and underlying mechanisms of multiple E3 ubiquitin ligases and deubiquitinases to regulate tumor metastasis. Further, we will discuss the relationship and importance between ubiquitination components and tumor progression.
Keywords: ubiquitination, E3 ubiquitin ligase, deubiquitinase, metastasis
A brief overview of the ubiquitination process
Protein ubiquitination is a dynamic, multifaceted post-translational modification involved in multiple cellular processes. Ubiquitin (Ub), a 76-amino acid protein, features seven lysine residues (K6, K11, K27, K29, K33, K48, and K63), which can each be ubiquitinated to form distinctive forms of polyubiquitin chains. Different polyubiquitin chains mediate distinct signaling pathways to determine the fate of substrate proteins [1]. For example, K48/K11-linked chains are responsible for targeting substrate proteins for proteasomal degradation, while other chains perform non-degradative roles in controlling protein interactions, cellular localization, and signaling transduction (Fig. 1).
A brief overview of the ubiquitination pathway. Ubiquitination is catalyzed by a three-enzyme cascade composed of the E1 Ub-activating enzyme, the E2 Ub-conjugating enzyme, and the E3 Ub ligase. The E3 ligase selectively recognizes substrate proteins by forming an iso-peptide bond and recruits the Ub-E2 complex to catalyze the transfer of Ub to the substrate from E2. Elongation and distinct polyubiquitin chains are involved in protein degradation, signal transduction, and transcriptional activity. Deubiquitinases remove Ub moieties from substrate proteins with high specificity and reverse Ub signals to maintain cellular dynamic ubiquitination.
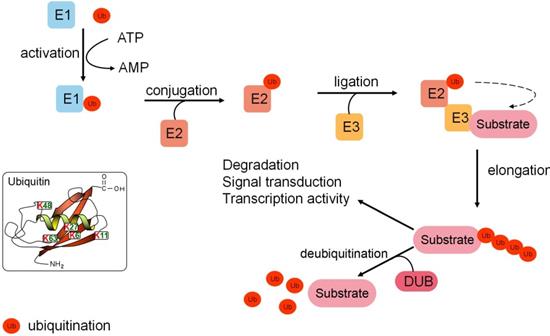
Ubiquitination is catalyzed by a three-enzyme cascade composed of the E1 Ub-activating enzyme, the E2 Ub-conjugating enzyme, and the E3 Ub ligase. E1 recruits and activates Ub by utilizing the energy of ATP. Activated Ub is transferred to E2, which can transfer Ub to the target substrate. E3 Ub ligase selectively recognizes a substrate protein by forming an iso-peptide bond between the COOH-terminal glycine of Ub and a lysine residue of the substrate. In addition, E3 Ub ligase recruits the E2-Ub complex and catalyzes the transfer of Ub to the substrate from E2 [2]. Different subtypes of E3s (the RING type E3s, the HECT type E3s, and the RBR type E3s) are the most critical component of the ubiquitination cascade for the substrate recognition capacity [2]. Meanwhile, ubiquitination is a dynamic and reversible process. Deubiquitinases (DUBs) can act as an “eraser” that reverses Ub signals. Most DUBs remove Ub moieties from proteins to prevent substrate proteins from degradation. However, some proteasome-related DUBs, including USP14, UCHL5, PSMD14, and PSMD7, are localized in 19S particles of the proteasome [3]. The roles of these DUBs are to deubiquitinate the substrates and facilitate their degradation in 20S particles of the proteasome. Alternatively, DUBs can alter signals by non-degradation ubiquitination [4] (Fig. 1).
The Ub cascade and DUBs synergistically regulate protein turnover and function in numerous signaling pathways to maintain cellular invasion and metastasis.
A novel insight into tumor metastasis
Tumor metastasis remains the primary cause of cancer-associated mortality. Metastasis involves tumor cell motility, intravasation into the adjacent tissues, circulation, and extravasation to distant organs. Simultaneously, the process is caused by genome instability where cancer cells can reprogram tumor metabolism, resist cell death, avoid immune destruction, and constitute the tumor microenvironment [5, 6]. The epithelial-mesenchymal transition (EMT) is an equally crucial determinant during the metastatic cascade [7].
Ubiquitination and deubiquitination broadly participate in various processes involved in protein modification and regulation. Aberrant dysregulation induces tumorigenesis. This review is primarily focused on recent novel observations and underlying mechanisms concerning E3 ligases and DUBs in order to contribute to further elucidating the role of ubiquitination and deubiquitination in tumor invasion and metastasis.
E3 Ub ligases inhibiting metastasis
F-box and WD-repeat domain-containing 2 (FBXW2)
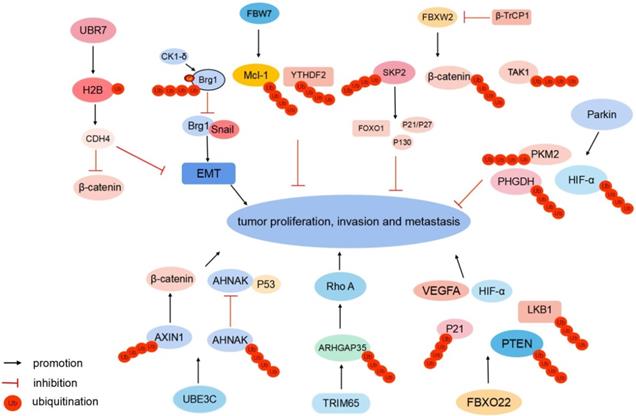
FBXW2 is a substrate recognition receptor in the SKP1-Cullin1-F-box protein (SCF) E3 Ub ligase complex. FBXW2 suppresses proliferation and invasion of lung cancer cells by targeting S phase kinase-associated protein 2 (SKP2) and β-catenin (Fig. 2). FBXW2 is downregulated and negatively correlated with β-catenin in lymph-node metastasis [8, 9]. Meanwhile, FBXW2 is a novel substrate of β-transducin repeat-containing protein 1 (β-TrCP1). Following growth factor stimulation, β-TrCP1 targets FBXW2 for ubiquitination degradation. The accumulation of SKP2 subsequently leads to the degradation of tumor suppressors and apoptosis-inducing substrates, such as p21, p27, p130, and FOXO1, to promote tumor cell proliferation, growth, and survival [8]. The β-TrCP-FBXW2-SKP2 signaling cascade forms the oncogene (β-TrCP1)-tumor suppressor gene (FBXW2)-oncogene (SKP2) axis that regulates the growth and survival of lung cancer cells via targeting each other for degradation, which is a process of crucial crosstalk among F-box proteins. In hepatocellular carcinoma (HCC), FBXW2 targets transforming growth factor-β-activated kinase 1 (TAK1) for K48-linked polyubiquitination and degradation to inhibit cancer progression [10] (Fig. 2). Clinically, FBXW2 levels could be a significant indicator of the prognosis and survival of patients with cancer.
Different E3 Ub ligases regulate tumor metastasis. FBXW2 targets SKP2, β-catenin, and TAK1, FBW7 targets Brg1, Mcl-1, and YTHDF2, and Parkin targets HIF-1α, PHGDH, and PKM2 for ubiquitination degradation to suppress tumor proliferation and metastasis, respectively. UBR7 monoubiquitinates histone H2B to suppress EMT and nuclear β-catenin. UBE3C targets AHNAK and AXIN1 for ubiquitination-induced degradation. TRIM65 ubiquitinates ARHGAP35, FBXO22 ubiquitinates nuclear PTEN and p21 to enhance cancer cell migration, respectively. FBXO22 mediates Lys-63-linked LKB1 ubiquitination. FBXO22 upregulates HIF-1α and VEGFA to promote tumor proliferation and metastasis.
F-box and WD repeat domain-containing 7 (FBW7)
FBW7 is a substrate recognition receptor of the SCFFBW7 E3 Ub ligase complex. It functions as a tumor suppressor and mediates ubiquitination-induced degradation of various oncogenic proteins, including c-MYC, NOTCH, c-JUN, and cyclin E [11]. In gastric cancer, transcription activator Brg1 (Brg1) binds to the promoter of Snail, which subsequently promotes EMT and metastasis. Meanwhile, Brg1 is a Ub substrate of the SCFFBW7 E3 ligase complex. Casein kinase 1 (CK1)δ-mediated phosphorylation at Ser31/Ser35 sites of Brg1 strengthens FBW7-binding capacity, thus accelerating the ubiquitination of Brg1[12] (Fig. 2). Methylation induces epigenetic silencing of the FBW7 gene. Decitabine (DAC) epigenetically activates FBW7 expression via its demethylation. DAC-activated FBW7 promotes myeloid leukemia cell differentiation protein Mcl-1 ubiquitination and degradation to suppress lung cancer growth [13]. YTH domain-containing family protein 2 (YTHDF2) is the N6-methyladenosine (m6A) reader protein and promotes the decay of the m6A-modified mRNAs. FBW7 can degrade YTHDF2 by ubiquitination and suppress the propagation of ovarian cancer [14] (Fig. 2).
Putative E3 Ub-protein ligase UBR7 (UBR7)
UBR7 belongs to the Ub-protein ligase E3 component N-recognin (UBR) family and has a unique plant homeodomain (PHD) finger [15]. PHD fingers are central “readers” of histone post-translational modifications [16]. UBR7-PHD finger monoubiquitinates histone H2B in triple-negative breast tumors at lysine 120 (H2BK120Ub). H2BK120Ub enhances cadherin-4 (CDH4) transcription activity and expression level [15]. CDH4 overexpression notably suppresses EMT and reduces cellular proliferation, migration, and invasion [15] (Fig. 2). Meanwhile, CDH4 can regulate the Wnt/β-catenin signaling pathway. It alters the nuclear localization of β-catenin to the cytoplasm, which downregulates β-catenin target genes, including AXIN2, G1/S-specific cyclin-D1, C-MYC, COX2, and MMP7, to suppress tumor metastasis [17, 18].
Parkin
Notably, as an E3 Ub ligase, Parkin can degrade substrate proteins associated with Parkinson's Disease (PD) [19]. Meanwhile, Parkin acts as a tumor suppressor, and its expression is downregulated in various tumors [20]. Parkin is an E3 Ub ligase for hypoxia-inducible factor 1α (HIF-1α) and can ubiquitinate HIF-1α at lysine 477 (K477), inhibit HIF-1α transcriptional activity, and induce its degradation to suppress breast cancer cells invasion and metastasis [21, 22]. Parkin regulates HIF-1α in a Von Hippel-Lindau-independent manner, unveiling an additional layer of regulation for HIF-1α in cells. Phosphoglycerate dehydrogenase (PHGDH) is the first rate-limiting enzyme of serine synthesis. PHGDH overexpression activates serine synthesis to promote cancer progression. Parkin expression is inversely correlated with PHGDH expression in breast and lung cancer. Parkin interacts with PHGDH and ubiquitinates PHGDH at lysine 330, leading to PHGDH degradation to suppress serine synthesis [23]. In intrahepatic cholangiocarcinoma (ICC), Parkin targets pyruvate kinase PKM2 for ubiquitination degradation to suppress migration and proliferation [24] (Fig. 2).
E3 Ub ligases promoting metastasis
Ub-protein ligase E3C (UBE3C)
UBE3C belongs to the HECT family of E3 Ub ligases. It functions as a tumor promoter and is aberrantly expressed in breast cancer [25], hepatocellular carcinoma [26], and renal cell carcinoma [27]. In non-small cell lung cancer (NSCLC) tissues, UBE3C maintains cancer stemness by ubiquitinating and promoting neuroblast differentiation-associated protein AHNAK (AHNAK) degradation [28]. AHNAK is a cofactor that assists P53 binding to stemness-related gene promoters. UBE3C-mediated degradation of AHNAK abrogates P53-AHNAK complex-mediated inhibition of gene expression, which enhances lung cancer cell stemness and NSCLC growth and metastasis [28, 29]. UBE3C targets AXIN1 for ubiquitination degradation to activate β-catenin signaling in gastric cancer [30] (Fig. 2).
Tripartite motif-containing protein 65 (TRIM65)
Tumor metastasis involves the reorganization of the cytoskeleton, whose activities are controlled by GTPases[31]. When bound to guanosine diphosphate (GDP), GTPases are inactivated, which is regulated by GTPase-activating protein (GAP)[32]. Rho A belongs to the Rho family of GTPases and regulates the cytoskeleton. Rho GTPase-activating protein 35 (ARHGAP35), a Rho GAP, regulates polarized cell migration and inhibits Rho GTPase. In colorectal cancer (CRC), E3 Ub ligase TRIM65 ubiquitinates and degrades ARHGAP35, which leads to subsequent elevated Rho GTPase activity and cytoskeleton remodeling [33] (Fig. 2). The TRIM65-ARHGAP35-Rho A axis enhances cancer cell migration by modulating the actin cytoskeleton.
F-box only protein 22 (FBXO22)
FBXO22, one of the F-box-only proteins, is the substrate-recognizing subunit of the SCF E3 Ub ligase complex [34]. FBXO22 ubiquitinates nuclear PTEN at lysine 221 (K221). Nuclear PTEN exerts potent tumor inhibition capacity. In CRC tissues, FBXO22 overexpression contributes to the downregulation of nuclear PTEN to promote tumorigenesis, which is reversed by the mutation of K221 [35]. FBXO22 is upregulated and negatively correlated with p21 in HCC. FBXO22 functions as an oncogene by mediating the ubiquitination and degradation of p21 to promote HCC pathogenesis and progression [36]. FBXO22 mediates Lys-63-linked liver kinase B1 (LKB1) polyubiquitination and inhibits LKB1-AMPK-mTOR signaling in lung adenocarcinoma [37]. FBXO22 promotes melanoma angiogenesis and migration of tumor cells via upregulating HIF-1α and vascular endothelial growth factor A (VEGFA) [38] (Fig. 2).
DUBs
DUBs can reverse ubiquitination by cleaving the isopeptide bond between the Ub and the substrate. Currently, over 100 DUBs have been identified and can be divided into six subclasses: i) Ub-specific proteases (USPs); ii) ovarian tumor proteases (OTUs); iii) Ub C-terminal hydrolases (UCHs); iv) Machado-Joseph disease proteases; v) JAB1/MPN/Mov34 metalloenzymes; and vi) monocyte chemotactic protein-induced protein[39]. DUBs act as tumor suppressors or oncogenes and play essential roles in regulating various types of tumors (Table 1). DUBs have emerged as promising therapeutic targets in cancer.
A brief overview of different DUBs in various cancers
| DUBs | Biological effect | Brief biological mechanism | Involvement in cancer | Refs |
|---|---|---|---|---|
| USP1 | Oncogene | Phosphorylated USP1 (via ATM/ATR) deubiquitinates and stabilizes Snail. | USP1 induces platinum resistance, cancer cell stemness, and metastatic dissemination in ovarian cancer. | [68] |
| USP1 deubiquitinates KPNA2 and enhances pro-metastatic genes expression. | The intervention of USP1 via pimozide or ML323 suppresses metastasis. | [69] | ||
| USP1 deubiquitinates and increases TAZ protein stability. | Loss of USP1 reduces TAZ to inhibit cell proliferation and migration, and USP1 is a potential therapeutic target in triple-negative breast cancer (TNBC). | [70] | ||
| USP1 deubiquitinates and stabilizes ribosomal protein S16 (RPS16). | USP1-mediated RPS16 stabilization promotes cell proliferation and metastasis in hepatocellular carcinoma (HCC). | [71] | ||
| USP2a | Oncogene | On TGF-β stimulation, USP2a deubiquitinates TGFBR1 (K33-linked ubiquitin chain), recruiting SMAD2. TGFBR2 subsequently phosphorylates USP2a, facilitating SMAD2 into the cytoplasm. | It is associated with trans-activating EMT genes to promote metastasis in lung adenocarcinomas. | [72] |
| USP2a deubiquitinates and stabilizes RAB1A. | USP2a is highly upregulated and promotes hepatocellular carcinoma (HCC) cell progression. | [73] | ||
| USP3 | Oncogene | USP3 deubiquitinates and upregulates SUZ12 protein expression. | USP3 promotes TGF-β1-induced EMT and cell migration in gastric cancer. | [74] |
| USP4 | Oncogene | USP4 deubiquitinates and stabilizes Twist1 protein. | USP4 enhances cancer cell stemness which mediates tumor development and metastasis in lung cancer and breast cancer. | [75] |
| USP4 constitutes the PAK5-DNPEP-USP4 axis. Aberrant PAK5 phosphorylates DNPEP, which preferentially facilitates DNPEP ubiquitination degradation, in turn inhibiting DNPEP-mediated USP4 downregulation. | USP4 elicits cancer cell proliferation, invasion, and metastasis in breast cancer. | [76] | ||
| USP5 | Oncogene | USP5 deubiquitinates and stabilizes SLUG. | USP5 promotes EMT, tumor growth, and metastasis, rescuing by Formononetin targeting USP5 in hepatocellular carcinoma (HCC). | [77] |
| USP5 prohibits β-catenin ubiquitination degradation and upregulates β-catenin, which activates the Wnt/β-catenin signaling. | USP5 is overexpressed in non-small cell lung cancer to promote EMT, invasion, and metastasis. | [78] | ||
| USP6 | Oncogene | USP6 promotes invasion and metastasis, and acts as an efficient prognostic biomarker. | USP6 is highly overexpressed in colon cancer. | [79] |
| USP7 (HAUSP) | Oncogene | USP7 promotes the circulation of tumor cells (CTCs) to reside in the bone marrow. | Inhibition of USP7 can arrest bone marrow-resident tumor cells (BMRTC) in BM and decrease metastasis. USP7 could be a therapeutic target in melanoma. | [80] |
| K63-polyubiquitinated HAUSP deubiquitinates and stabilizes HIF-1α, and causes CBP-mediated H3K56 acetylation to regulate HIF-1α target gene promoters. | Under hypoxia, E3 ligase HectH9 is required for K63-polyubiquitinated HAUSP to promote EMT and metastasis in lung cancer. | [81] | ||
| USP7 promotes proliferation and invasion. | Overexpressed USP7 represents a worse overall survival and acts as an independent prognostic indicator in epithelial ovarian cancer (EOC) and oral squamous cell carcinoma (OSCC). | [82, 83] | ||
| USP7 overexpression activates the PI3K/AKT signaling pathway. | The USP7 inhibitor P5091 suppresses cell proliferation and metastasis and promotes hepatoblastoma (HB) apoptosis. | [84] | ||
| USP7 activates the Wnt/β-catenin signaling pathway. | USP7 inhibitor P22077 induces apoptosis and DNA damage and suppresses cell migration and invasion in melanoma. | [85] | ||
| USP8 | Oncogene | Overexpressed USP8 increases p-AKT, activates AKT signaling, and regulates intrinsic apoptosis pathway. | USP8 suppresses apoptosis and promotes proliferation, invasion, and metastasis in cholangiocarcinoma. | [86] |
| TSG (Tumor suppressor gene) | USP8 is a protective factor and prognosticates better clinical outcomes. | USP8 is downregulated in breast cancer. | [87] | |
| USP9X | Oncogene | USP9X activates EMT. | USP9X overexpression promotes invasion and migration, and inhibits apoptosis in pancreatic ductal adenocarcinoma (PDAC). | [88,89] |
| USP9X bans TIF1γ from ubiquitinating SMAD4 and maintains its nuclear retention to induce the TGF-β signaling. | Plasma-free fatty acids (FFA) promote the SMAD4-USP9X interaction via ERK to elicit TGF-β-induced metastasis for obese breast cancer patients. | |||
| USP10 | Oncogene | USP10 deubiquitinates Smad4 (K48) and activates TGF-β. | USP10-siRNA and Spautin1 inhibitor can downregulate USP10 to suppress Smad4 and metastasis in hepatocellular carcinoma. | [90] |
| USP10 deubiquitinates NLRP7 to induce M2 TAM polarization via CCL2 secretion. | USP10 is highly expressed and stabilizes NLRP7 to promote cell proliferation and metastasis in colorectal cancer. | [91] | ||
| USP11 | Oncogene | USP11 deubiquitinates and stablizes PPP1CA to activate the ERK/MAPK pathway. | USP11 is overexpressed and promotes metastasis in colorectal cancer. | [92-95] |
| USP11 deubiquitinates and stabilizes nuclear factor (NF90). | USP11 can promote proliferation and metastasis in hepatocellular carcinoma. | |||
| USP11 stabilizes TGFβ receptor type 2 (TGFBR2). | USP11 enhances TGFβ-induced EMT to promote breast cancer metastasis. | |||
| USP11 is an independent prognostic predictor. | USP11 is overexpressed and promotes migration and metastasis in hepatocellular carcinoma. | |||
| TSG | USP11 deubiquitinates ARID1A and prevents its degradation to inhibit SDC2 activation. | USP11 antagonizes with TRIM32 to stabilize ARID1A and to suppress proliferation and metastasis in squamous cell carcinomas (SCCs). | [96, 97] | |
| USP11 deubiquitinates and stabilizes PTEN and subsequently suppresses the PI3K/AKT pathway. | PTEN inhibits the PI3K/AKT-mediated phosphorylation of FOXO to increase its nuclear localization and to enhance USP11 transcription. The PTEN-PI3K/AKT-FOXO-USP11 regulatory feedforward loop regulates the tumor- suppressive activity of PTEN. | |||
| USP12 | Oncogene | USP12 deubiquitinates and stabilizes midkine (MDK). | The USP12-MDK axis promotes angiogenesis to faciliate breast cancer metastasis. | [98] |
| USP14 | Oncogene | USP14 deubiquitinates PI3K. | USP14 inhibitor Lidocaine (Lido) suppresses proliferation and migration while aggravating hepatocellular carcinoma cell apoptosis. | [99] |
| USP14 overexpression promotes proliferation and migration and prevents apoptosis. | USP14 is remarkably upregulated in pancreatic ductal adenocarcinoma (PDAC). | [100] | ||
| USP15 | Oncogene | USP15 overexpression promotes proliferation and prevents apoptosis. | High USP15 expression indicates a worse prognosis, and USP15 could be a therapeutic target in hepatocellular carcinoma. | [101] |
| USP15 promotes β-catenin nuclear translocation and activates the Wnt/β-catenin pathway. | USP15 is upregulated and promotes EMT, cell proliferation, and metastasis. | [102] | ||
| USP18 | Oncogene | USP18 deubiquitinates ZEB1. | USP18 is overexpressed and induces ZEB1-mediated EMT to promote metastasis in esophageal squamous cell carcinomas (ESCC). | [103] |
| USP20 | Oncogene | USP20 deubiquitinates β-catenin. | USP20 highly expresses and regulates the Wnt/β-catenin pathway to potentiate tumorigenesis in colon cancer. | [104] |
| USP21 | Oncogene | USP21 deubiquitinates EZH2. | USP21 upregulates and promotes EMT and metastasis in bladder cancer. | [105, 106] |
| USP21 deubiquitinates Fos-related-antigen-1 (Fra-1) and enhances AP-1 target gene expression. | USP21 overexpresses and promotes Fra1-dependent metastasis in colorectal cancer. | |||
| USP22 | Oncogene | USP22 activates AP4 transcription to induce EMT. | USP22 and AP4 overexpress and promote liver metastasis in colorectal cancer. | [107-111] |
| USP22 stabilizes BMI1 protein to maintain cancer stemness. | USP22 and BMI1 overexpress and facilitate proliferation in gastric cancer. | |||
| High USP22 enhances angiogenesis, metastasis, and recurrence. | USP22 knockout suppresses metastasis and sensitizes cisplatin and irradiation in non-small cell lung cancer. | |||
| USP22 increases the relative abundance of myeloid cells vs. cytotoxic T cells via its deubiquitinase activity. | USP22 ablation can suppress metastasis and improve the response to immunotherapy in pancreatic ductal adenocarcinoma (PDA). | |||
| USP22 promotes gastric cancer progression by modulating FOXO1 and the YAP signaling pathways via c-Myc/NAMPT/SIRT1. | USP22 is overexpressed, and its depletion suppresses invasion and metastasis in gastric cancer. | |||
| TSG | USP22 decreases mTOR activity. | USP22 deficiency activates mTOR and tumorigenesis, reversed by mTOR inhibitor treatment in colorectal cancer. | [112] | |
| USP25 | Oncogene | mi-RNA 200c reduces the USP25 gene mRNA and protein levels to inhibit invasion and migration. | USP25 protein and mRNA levels are highly expressed in non-small cell lung cancer. | [113] |
| USP26 | Oncogene | USP26 deubiquitinates and stabilizes Snail. | USP26 is highly expressed in esophageal squamous cell carcinoma (ESCC). | [114] |
| USP28 | Oncogene | USP28 stabilizes lysine specific demethylase1. | USP28 is overexpressed in gastric cancer. | [115, 116] |
| USP28 antagonizes GSK3β-Fbw7-dependent HIF-1α ubiquitination degradation to affect HIF-1α-dependent angiogenesis and carcinogenesis. | Expression of USP28 is elevated in colon and breast carcinomas. | |||
| USP29 | Oncogene | USP29 interacts simultaneously with Snail and SCP1 to stabilize Snail via deubiquitination and dephosphorylation. | TNFα, TGFβ, and Hypoxia can induce USP29 to promote gastric cancer cell migration. | [117] |
| USP33 | Oncogene | USP33 deubiquitinates specificity protein 1 (SP1) to upregulate c-met. | USP33 is overexpressed and is a prognostic biomarker and therapeutic target in hepatocellular carcinoma. | [118, 119] |
| TSG | USP33 can deubiquitinate and stabilize Robo1 to inhibit EMT and cell migration in a Slit-Robo pathway-dependent manner. | USP33 expression is downregulated and it is an independent prognostic marker in colorectal cancer and gastric cancer. | [120, 121] | |
| USP37 | Oncogene | USP37 deubiquitinates Snail. | Upregulated expression of USP37 promotes lung cancer cell migration. | [122-124] |
| USP37 stabilizes the hedgehog (Hh) pathway component Gli-1. | USP37 can regulate the stemness, cell invasion, cisplatin sensitivity, and EMT via the Hh pathway in breast cancer. | |||
| USP37 binds and deubiquitinates Snai1. | Overexpression of USP37 upregulates Snai1 to promote cancer cell migration. | |||
| USP43 | TSG | USP43 physically binds to the chromatin remodeling NuRD complex and catalyzes H2BK120 deubiquitination to repress the EGFR gene. | EGFR/PI3K/Akt-mediated phosphorylated USP43 binds to the 14-3-3β/ε heterodimer and sequestrates in the cytoplasm to drive breast carcinogenesis. | [52] |
| USP44 | Oncogene | USP44 deubiquitinates EZH2, a histone H3 lysine 27 methyltransferase. | USP44 knockdown decreases the EZH2 protein level and inhibits prostate cancer cells' tumorigenesis and cancer stem cell-like behaviors. | [125, 126] |
| USP44 expression in breast cancer stem cells (CSC) contributes to the formation of vasculogenic mimicry (VM) to promote transendothelial migration. | USP44 silencing abates VM and USP44+CSC subclones act as an independent prognostic biomarker in breast cancer. | |||
| USP46 | Oncogene | USP46 deubiquitinates ENO1 and promotes EMT. | USP46 is overexpressed in esophageal squamous cell carcinoma (ESCC). | [127] |
| USP47 | Oncogene | USP47 deubiquitinates and stabilizes Snail to induce EMT. | Inhibiton of USP47 with P5091 can reverse the EMT phenotype. | [128-130] |
| USP47, as a novel target of Sox9, mediates hypoxia-induced EMT via deubiquitinating Snail. | The expression of USP47 is elevated, and silencing USP47 can promote Snail degradation and attenuate EMT in colorectal cancer. | |||
| USP47 abrogates the SMURF2-mediated ubiquitination of special AT-rich sequence-binding protein-1 (SATB1) to promote colon cacer cell proliferation and metastasis. | USP47 depletion sensitizes colon cancer cells to 5-FU treatment-induced apoptosis. | |||
| USP48 | Oncogene | USP48 promotes migration and invasion. | Ablation of USP48 increases the responsiveness to carboplatin treatment in ovarian cancer. | [131] |
| USP51 | Oncogene | CDK4/6-mediated phosphorylated USP51 can deubiquitinate and stabilize ZEB1 to induce EMT. | The overexpressed p-USP51 is correlated to a poor prognosis for breast cancer patients, and the CDK4/6-USP51-ZEB1 axis could be a viable therapeutic target. | [132-134] |
| USP51 increases FAT4 protein level and is imperative for FAT4's function. | USP51 suppression contributes to the inhibition of FAT4 and promotes proliferation and invasion of endometrial cancer (EC). | |||
| USP54 | Oncogene | USP54 is of pro-tumorigenic properties. | USP54 is upregulated in colorectal carcinoma and is a promising therapeutic target. | [135] |
| OTUB1 | Oncogene | OTUB1 stabilizes Snail to promote metastasis. | OTUB1 is highly expressed in esophageal squamous cell carcinoma (ESCC), and higher expression of OTUB1 predicts poor prognosis. | [136, 137] |
| OTUB1 induces EMT to promote metastasis. | OTUB1 is overexpressed and related to poor survival and serves as an independent prognostic factor in colorectal cancer (CRC). | |||
| OTUB2 | Oncogene | OTUB2 can deubquitinate U2AF2 and activate the AKT/mTOR pathway. | OTUB2 and U2AF2 are highly expressed and associated with metastasis and poor survival. OTUB2 may serve as a potential prognostic indicator and therapeutic target in NSCLC. | [138, 139] |
| EGF/KRAS-induced SUMOylation of OTUB2 can deubiquitinate and activate YAP/TAZ. | OTUB2 can promote cancer stemness and metastasis via the Hippo-independent pathway. | |||
| OTUD1 | TSG | OTUD1 deubiquitinates K48-linked and K33-linked SMAD7 to enhance SMURF2 binding to suppress TGFβ. | High-level OTUD1 inhibits TGFβ-induced cancer stemness and metastasis in breast cancer. | [48] |
| OTUD3 | Oncogene | OTUD3 stabilizes GRP78 to promote lung tumorigenesis, reversed by CHIP which can ubiquitinate OTUD3. | CHIP knockdown increases lung cancer cell invasion in an OTUD3 and GRP78- dependent manner. | [140] |
| TSG | OTUD3 deubiquitinates and stabilizes PTEN. | Reduction of OTUD3 causes decreased PTEN abundance and correlates with breast cancer progression. | [141] | |
| OUTD6B | TSG | OTUD6B couples pVHL to form the CBCVHL complex to decrease its ubiquitination degradation, thereby attenuating HIF-1α. | OTUD6B is positively correlated with pVHL, but negatively with HIF-1α and vascular endothelial growth factor in hepatocellular carcinoma. | [142] |
| OUTD7B | TSG | OTUD7B promotes proliferation and metastasis via the Akt/VEGF signal pathway. | OTUD7B is highly expressed in lung squamous carcinoma and adenocarcinoma, and correlates with a worse prognosis, and may be an independent predictive indicator. | [143] |
| BAP1 | Oncogene | BAP1 deubiquitinates transcription factor KLF5. | BAP1 konckdown inhibits breast cancer tumorigenicity and lung metastasis, and BAP1 could be a therapeutic target. | [144] |
| TSG | BAP1 inhibits the ERK1/2 and JNK/c-Jun pathway to repress intrahepatic cholangiocarcinoma (ICC). | mRNA and protein level of BAP1 are downregulated, related to ICC aggressive characteristics. BAP1 may be a prognostic and therapeutic target. | [41, 145-147] | |
| BAP1 mutants induce migration. | BAP1 blocks metastasis in solid pseudopapillary neoplasms (SPN). | |||
| BAP1 somatic mutations, not germline mutations, infrequently occur in uveal melanoma. | ||||
| CYLD | TSG | CYLD regulates genes involved in proliferation, migration, and angiogenesis. | CYLD-deficiency enhances melanoma progression. | [148-150] |
| CYLD can reverse the K63 ubiquitination of c-Jun and c-Fos to repress the JNK/AP1 pathway. | CYLD mutant enhances squamous cell carcinoma growth and migration in an AP1-dependent manner. | |||
| Snail1 inhibits CYLD to promote BCL-3 nuclear translocation, activating cycling D1 and N-cadherin. | Upregulation of CYLD expression can repress proliferation and invasion in melanoma. | |||
| UCHL1 | Oncogene | UCHL1 promotes EMT. | UCHL1 is overexpressed, and knockdown can induce MET in metastatic prostate cancer. | [151-156] |
| UCHL1 activates the AKT/MAPK pathway. | AKT negative mutant and silencing UCHL1 suppress invasion and metastasis in non-small cell lung cancer. | |||
| UCHL1 activates the MAPK/ERK pathway. | ERK inhibitor U0126 can block multidrug resistance and invasion in UCHL1-overexpressed breast cancer cells. | |||
| UCHL1 compromises VHL-mediated ubiquitination of HIF-1α to promote metastasis. | UCHL1 is overexpressed in breast and lung cancer. It may be a prognostic marker and therapeutic target. | |||
| UCHL1 expression is positively associated with renal cell cancer's (RCC) metastatic phenotype. | UCHL1 might serve as a potential diagnostic and prognosis biomarker for RCC patients. | |||
| UCHL1 deubiquitinates TGFβ type I receptor and SMAD2. | The UCHL1 inhibitor 6RK73 suppresses TGFβ-induced metastasis, and UCHL1 could potentially target triple-negative breast cancer (TNBC) treatment. | |||
| UCHL5 | Oncogene | UCHL5 can activate the Wnt/β-catenin pathway and upregulate β-catenin. | UCHL5 is overexpressed, and promotes tumorigenesis and growth in endometrial cancer (EC), which can be abrogated by the Wnt/β-catenin pathway inhibitor XAV939. | [157] |
| COPS5 | Oncogene | COPS5 deubiquitinates HK2 and attenuates its degradation to regulate glycolysis. | COPS5 relates to HK2 overexpression, and Curcumin can inhibit CSN5 activity to decrease HK2, and to repress glycolysis and metastasis in hepatocellular carcinoma. | [158, 159] |
| COPS5 deubiquitinates and stabilizes ZEB1. | COPS5 expression is elevated and its knockdown can suppress EMT and metastasis. | |||
| COPS6 | Oncogene | COPS6 increases CHIP self-ubiquitination to elevate EGFR stability. | COPS6 is overexpressed and the CSN6-CHIP-EGFR axis could be a therapeutic target in glioblastoma. | [160, 161] |
| COPS6 inhibits the autophagy of CathepsinL (CTSL) via the mTOR pathway. | COPS6 and CTSL are overexpressed and indicate aggressive cervical cancer. | |||
| ATXN3 | Oncogene | ATXN3 deubiquitinates KLF4. | High ATXN3 and KLF4 expression are associated with a poor prognosis in breast cancer patients. | [162] |
DUBs inhibiting metastasis
Ub carboxyl-terminal hydrolase BAP1 (BAP1)
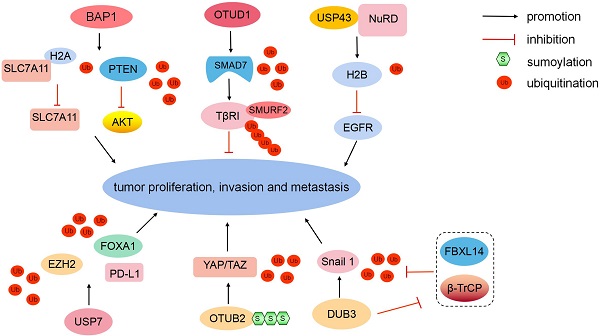
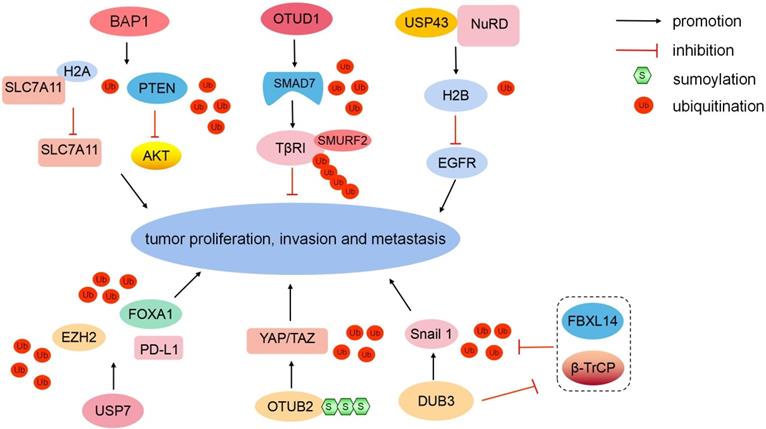
BAP1 belongs to the UCH domain-containing proteins, and it can physically bind to and deubiquitinate PTEN to stabilize PTEN protein. Downregulated BAP1 leads to the decrease of PTEN protein levels and the activation of the Akt signaling pathway, therefore promoting malignant transformation and metastasis in prostate cancer. Clinically, low BAP1 expression is positively correlated with aggressive prostate tumor proliferation and lymphatic metastasis [40]. In ICC, BAP1 inhibits ERK1/2 and JNK/c-Jun pathways [41]. Moreover, BAP1 mediates the metabolic regulation of ferroptosis and tumor suppression. BAP1 reduces histone 2A ubiquitination (H2AUb) on the cystine/glutamate transporter (SLC7A11) promoter and represses SLC7A11 expression in a deubiquitination-dependent manner [42]. The loss of cystine transport mediated by SLC7A11 induces ferroptosis to inhibit tumor development and metastasis [43] (Fig. 3). In summary, as a major tumor suppressor, mutated BAP1 is associated with numerous human malignancies, which is defined as “BAP1 cancer syndrome” [44].
Different DUBs regulate tumorigenesis. BAP1 can deubiquitinate PTEN and reduce H2AUb on the SLC7A11 promoter to inhibit tumor development and metastasis. OTUD1 can eliminate the K48-linked Ub chain from SMAD7. The USP43-NuRD complex catalyzes H2BK120 deubiquitination to suppress downstream EGFR. USP7 can stabilize EZH2 and FOXA1 via deubiquitination to promote tumor growth. USP7 upregulates PD-L1 expression in the tumor microenvironment. SUMOylated-OTUB2 deubiquitinates YAP/TAZ to potentiate tumor cell proliferation and metastasis in a Hippo-independent manner. Dub3 can stabilize Snail1 via deubiquitination and inhibiting β-TrCP1 and FBXL14-mediated ubiquitination degradation.
OTU domain-containing protein 1 (OTUD1)
In the TGF-β signaling pathway, SMAD7 recruits E3 ligase SMURF2 to TGF-β type I receptor (TβRI) [45]. However, E3 Ub-protein ligases RNF12 and Itchy homolog can degrade SMAD7 to antagonize its suppression [46, 47]. In breast cancer, OTUD1 can selectively eliminate the K48-linked Ub chain from SMAD7 to stabilize it (Fig. 3). Moreover, OTUD1 can cleave the K33-linked Ub chain on the lysine 220 site and unveil the PY motif of SMAD7 [48]. The exposed SMAD7 PY motif subsequently binds the WW domain of SMURF2 to degrade TβRI via ubiquitination [45, 49]. In summary, OTUD1 suppresses the TGF-β-induced metastasis by exerting dual effects on SMAD7.
Ub carboxyl-terminal hydrolase 43 (USP43)
The EGFR/PI3K/AKT pathway is aberrantly activated in various cancers [50]. PTEN can negatively regulate AKT kinase activity [51]. In addition, another molecular regulatory mechanism exists. Nuclear USP43 is physically associated with the nucleosome remodeling and deacetylase (NuRD) complex. The USP43-NuRD complex coordinately catalyzes H2BK120 deubiquitination to suppress downstream EGFR [52] (Fig. 3). Simultaneously, activated AKT can phosphorylate cytoplasmic USP43 on Ser29, phosphorylated USP43 is detained in the cytoplasm, reduced nuclear USP43 and accumulated EGFR potentiate the EGFR/PI3K/AKT pathway to promote breast cell proliferation and invasion [52]. USP43 is a hub of the USP43-NuRD complex, the reciprocally inhibitory loop between the USP43-NuRD complex and the EGFR/PI3K/AKT pathway synergistically modulates breast carcinogenesis. The ratio of nuclear/cytoplasmic USP43 is a worthy prognostic indicator [52].
DUBs promoting metastasis
USP7
USP7 can coordinate with histone-lysine N-methyltransferase EZH2 (EZH2)-catalyzed methylation to remove ubiquitination and enhance FOXA1 protein stability, promoting prostate cancer growth [53]. Moreover, USP7 can stabilize EZH2 via deubiquitination [54] (Fig. 3). USP7 is upregulated in M2 macrophages. USP7 inhibition can induce the polarization of tumor-associated macrophages from M2 into M1 by activating the P38 MAPK pathway and upregulating the expression of programmed cell death 1 ligand 1 (PD-L1) in the tumor microenvironment [55]. Therefore, USP7 blockade combined with anti-PD-1 immunotherapy exert an inhibitory effect on tumors in lung cancer.
SUMOylated-Ub thioesterase OTUB2 (OTUB2)
Transcriptional coactivator YAP1 (YAP) and tafazzin (TAZ) are generally downregulated by the canonical Hippo pathway [56]. However, YAP and TAZ are hyperactivated and induce tumor proliferation and metastasis, while the Hippo pathway is still active in multiple malignancies, including breast cancer [57, 58]. DUB OTUB2 mediates the activation of YAP and TAZ in a Hippo-independent manner (Fig. 3). Mechanistically, OTUB2 is poly-SUMOylated at lysine 233 (K233), SUMOylated-OTUB2 can subsequently bind YAP/TAZ through SUMO-interacting motif in YAP and TAZ [58]. OTUB2 deubiquitinates and activates YAP and TAZ, and accumulated YAP and TAZ translocate into the nucleus in which they interact with TEA domain family transcription factors and transcriptionally activate genes to potentiate cell proliferation and metastasis [59, 60]. Meanwhile, activated EGF-RAS signaling strengthens OTUB2 SUMOylation and elevates YAP/TAZ protein levels to promote cancer stemness and metastasis [58, 61]. In summary, the novel SUMOylated-OTUB2-mediated regulatory mechanism expands the complexity of YAP/TAZ beyond the Hippo pathway. OTUB2 may be a potential drug target to suppress cancer progression for patients harboring RAS mutations.
Dub3
Snail1 is a critical EMT-driving transcription factor and confers tumor metastatic and cancer stem cell-like properties [62]. The E3 ligases β-TrCP1 and F-box/LRR-repeat protein 14 (FBXL14) can degrade Snail1 via ubiquitination [63]. In breast cancer, Dub3 accounts for Snail1 stabilization, and inflammatory cytokine IL-6 can increase the expression of Dub3. Meanwhile, Dub3 also inhibits the activity of β-TrCP1 and FBXL14 to block Snail1 ubiquitination [64] (Fig. 3). Overall, Dub3 senses inflammatory stimulation and converts it into Snail1 stabilization.
Conclusion
Although this brief review only scratches the surface of ubiquitination and deubiquitination in cancer, it highlights the significance of E3s and DUBs in a range of processes involved in tumor progression. Ubiquitination components are potential therapeutic targets for cancer treatment [65]. However, several issues remain obstacles for targeted therapy. DUBs share similar structural characteristics among family members, and ubiquitination involves substantial conformational changes. We still endeavor to deal with the challenges ahead, such as defining novel E3s, DUBs, and targeted substrates, investigating whether there exists unknown crosstalk among distinct E3s or DUBs, and decoding the unknown pathways linking ubiquitination with other cellular physiological mechanisms. Several valuable E3s or DUBs are promising clinical prognostic indexes and drug targets. Proteolysis Targeting Chimeras (PROTACs) exploit the intracellular Ub-proteasome system to degrade target proteins [66], selectively. In tumor xenografts, small-molecule PROTACs can significantly attenuate tumor progression [67].
This review provides a glimpse into the importance and extensiveness of ubiquitination component-mediated tumor invasion and metastasis, which represents a worthy research prospect.
Abbreviations
Ub: ubiquitin; DUBs: deubiquitinases; SKP2: S phase kinase-associated protein 2; FBXW2: F-box and WD-repeat domain-containing 2; TAK1: transforming growth factor-β-activated kinase 1; β-TrCP1: β-transducin repeat-containing protein 1; FBW7: F-box and WD repeat domain-containing 7; Brg1: transcription activator Brg1; Mcl-1: induced myeloid leukemia cell differentiation protein Mcl-1; YTHDF2: YTH domain-containing family protein 2; EMT: epithelial-mesenchymal transition; CK1: casein kinase 1; UBR7: putative E3 Ub-protein ligase UBR7; CDH4: cadherin-4; HIF-1α: hypoxia-inducible factor 1α; PHGDH: phosphoglycerate dehydrogenase; PKM: pyruvate kinase PKM; UBE3C: Ub-protein ligase E3C; AHNAK: neuroblast differentiation-associated protein AHNAK; TRIM65: tripartite motif-containing protein 65; ARHGAP35: Rho GTPase-activating protein 35; FBXO22: F-box only protein 22; LKB1: liver kinase B1; VEGFA: vascular endothelial growth factor A; BAP1: Ub carboxyl-terminal hydrolase BAP1; H2AUb: histone 2A ubiquitination; SLC7A11: cystine/glutamate transporter; SMURF2: E3 Ub-protein ligase SMURF2; TβRI: TGF-β type I receptor; OTUD1: OTU domain-containing protein 1; USP43: Ub carboxyl-terminal hydrolase 43; NuRD: nucleosome remodeling and deacetylase; H2BK120: histone H2B at lysine 120; USP: Ub-specific protease; EZH2: histone-lysine N-methyltransferase EZH2; PD-L1: programmed cell death 1 ligand 1; OTUB2: Ub thioesterase OTUB2; YAP: transcriptional coactivator YAP1; TAZ: tafazzin; FBXL14: F-box/LRR-repeat protein 14.
Acknowledgements
We apologize to colleagues whose work was not cited due to space limitations. This study was supported by China's National Natural Science Foundation (No.81972837).
Competing Interests
The authors have declared that no competing interest exists.
References
1. Swatek KN, Komander D. Ubiquitin modifications. Cell Res. 2016;26:399-422
2. Zheng N, Shabek N. Ubiquitin Ligases: Structure, Function, and Regulation. Annual review of biochemistry. 2017;86:129-57
3. Sowa ME, Bennett EJ, Gygi SP, Harper JW. Defining the human deubiquitinating enzyme interaction landscape. Cell. 2009;138:389-403
4. Clague MJ, Urbé S, Komander D. Breaking the chains: deubiquitylating enzyme specificity begets function. Nature reviews Molecular cell biology. 2019;20:338-52
5. Decoding cancer metastasis. Nat Cell Biol. 2018; 20: 859.
6. Massagué J, Obenauf AC. Metastatic colonization by circulating tumour cells. Nature. 2016;529:298-306
7. Pastushenko I, Blanpain C. EMT Transition States during Tumor Progression and Metastasis. Trends in cell biology. 2019;29:212-26
8. Xu J, Zhou W, Yang F, Chen G, Li H, Zhao Y. et al. The β-TrCP-FBXW2-SKP2 axis regulates lung cancer cell growth with FBXW2 acting as a tumour suppressor. Nat Commun. 2017;8:14002
9. Yang F, Xu J, Li H, Tan M, Xiong X, Sun Y. FBXW2 suppresses migration and invasion of lung cancer cells via promoting β-catenin ubiquitylation and degradation. Nat Commun. 2019;10:1382
10. Xia S, Ji L, Tao L, Pan Y, Lin Z, Wan Z. et al. TAK1 Is a Novel Target in Hepatocellular Carcinoma and Contributes to Sorafenib Resistance. Cellular and molecular gastroenterology and hepatology. 2021;12:1121-43
11. Davis RJ, Welcker M, Clurman BE. Tumor suppression by the Fbw7 ubiquitin ligase: mechanisms and opportunities. Cancer cell. 2014;26:455-64
12. Huang LY, Zhao J, Chen H, Wan L, Inuzuka H, Guo J. et al. SCF(FBW7)-mediated degradation of Brg1 suppresses gastric cancer metastasis. Nat Commun. 2018;9:3569
13. Kim MJ, Chen G, Sica GL, Deng X. Epigenetic modulation of FBW7/Mcl-1 pathway for lung cancer therapy. Cancer biology & therapy. 2021;22:55-65
14. Xu F, Li J, Ni M, Cheng J, Zhao H, Wang S. et al. FBW7 suppresses ovarian cancer development by targeting the N(6)-methyladenosine binding protein YTHDF2. Molecular cancer. 2021;20:45
15. Adhikary S, Chakravarti D, Terranova C, Sengupta I, Maitituoheti M, Dasgupta A. et al. Atypical plant homeodomain of UBR7 functions as an H2BK120Ub ligase and breast tumor suppressor. Nat Commun. 2019;10:1398
16. Jain K, Fraser CS, Marunde MR, Parker MM, Sagum C, Burg JM. et al. Characterization of the plant homeodomain (PHD) reader family for their histone tail interactions. Epigenetics & chromatin. 2020;13:3
17. Pohl SG, Brook N, Agostino M, Arfuso F, Kumar AP, Dharmarajan A. Wnt signaling in triple-negative breast cancer. Oncogenesis. 2017;6:e310
18. Li J, Zhou BP. Activation of β-catenin and Akt pathways by Twist are critical for the maintenance of EMT associated cancer stem cell-like characters. BMC Cancer. 2011;11:49
19. Pickrell AM, Youle RJ. The roles of PINK1, parkin, and mitochondrial fidelity in Parkinson's disease. Neuron. 2015;85:257-73
20. Xu L, Lin DC, Yin D, Koeffler HP. An emerging role of PARK2 in cancer. Journal of molecular medicine (Berlin, Germany). 2014;92:31-42
21. Liu J, Zhang C, Zhao Y, Yue X, Wu H, Huang S. et al. Parkin targets HIF-1α for ubiquitination and degradation to inhibit breast tumor progression. Nat Commun. 2017;8:1823
22. LaGory EL, Giaccia AJ. The ever-expanding role of HIF in tumour and stromal biology. Nat Cell Biol. 2016;18:356-65
23. Liu J, Zhang C, Wu H, Sun XX, Li Y, Huang S. et al. Parkin ubiquitinates phosphoglycerate dehydrogenase to suppress serine synthesis and tumor progression. The Journal of clinical investigation. 2020;130:3253-69
24. Chen D, Wu H, Feng X, Chen Y, Lv Z, Kota VG. et al. DNA Methylation of Cannabinoid Receptor Interacting Protein 1 Promotes Pathogenesis of Intrahepatic Cholangiocarcinoma Through Suppressing Parkin-Dependent Pyruvate Kinase M2 Ubiquitination. Hepatology (Baltimore, Md). 2021;73:1816-35
25. Xiong J, Wei B, Ye Q, Liu W. MiR-30a-5p/UBE3C axis regulates breast cancer cell proliferation and migration. Biochem Biophys Res Commun. 2019;516:1013-8
26. Jiang JH, Liu YF, Ke AW, Gu FM, Yu Y, Dai Z. et al. Clinical significance of the ubiquitin ligase UBE3C in hepatocellular carcinoma revealed by exome sequencing. Hepatology (Baltimore, Md). 2014;59:2216-27
27. Wen JL, Wen XF, Li RB, Jin YC, Wang XL, Zhou L. et al. UBE3C promotes growth and metastasis of renal cell carcinoma via activating Wnt/β-catenin pathway. PloS one. 2015;10:e0115622
28. Gu J, Mao W, Ren W, Xu F, Zhu Q, Lu C. et al. Ubiquitin-protein ligase E3C maintains non-small-cell lung cancer stemness by targeting AHNAK-p53 complex. Cancer Lett. 2019;443:125-34
29. Aloni-Grinstein R, Shetzer Y, Kaufman T, Rotter V. p53: the barrier to cancer stem cell formation. FEBS letters. 2014;588:2580-9
30. Zhang Y, Xu J, Fu H, Wei Z, Yang D, Yan R. UBE3C promotes proliferation and inhibits apoptosis by activating the β-catenin signaling via degradation of AXIN1 in gastric cancer. Carcinogenesis. 2021;42:285-93
31. Jaffe AB, Hall A. Rho GTPases: biochemistry and biology. Annual review of cell and developmental biology. 2005;21:247-69
32. Etienne-Manneville S, Hall A. Rho GTPases in cell biology. Nature. 2002;420:629-35
33. Chen D, Li Y, Zhang X, Wu H, Wang Q, Cai J. et al. Ubiquitin ligase TRIM65 promotes colorectal cancer metastasis by targeting ARHGAP35 for protein degradation. Oncogene. 2019;38:6429-44
34. Sun R, Xie HY, Qian JX, Huang YN, Yang F, Zhang FL. et al. FBXO22 Possesses Both Protumorigenic and Antimetastatic Roles in Breast Cancer Progression. Cancer Res. 2018;78:5274-86
35. Ge MK, Zhang N, Xia L, Zhang C, Dong SS, Li ZM. et al. FBXO22 degrades nuclear PTEN to promote tumorigenesis. Nat Commun. 2020;11:1720
36. Zhang L, Chen J, Ning D, Liu Q, Wang C, Zhang Z. et al. FBXO22 promotes the development of hepatocellular carcinoma by regulating the ubiquitination and degradation of p21. J Exp Clin Cancer Res. 2019;38:101
37. Zhu XN, He P, Zhang L, Yang S, Zhang HL, Zhu D. et al. FBXO22 mediates polyubiquitination and inactivation of LKB1 to promote lung cancer cell growth. Cell Death Dis. 2019;10:486
38. Zheng Y, Chen H, Zhao Y, Zhang X, Liu J, Pan Y. et al. Knockdown of FBXO22 inhibits melanoma cell migration, invasion and angiogenesis via the HIF-1α/VEGF pathway. Investigational new drugs. 2020;38:20-8
39. Sun J, Shi X, Mamun MAA, Gao Y. The role of deubiquitinating enzymes in gastric cancer. Oncology letters. 2020;19:30-44
40. Deng R, Guo Y, Li L, He J, Qiang Z, Zhang H. et al. BAP1 suppresses prostate cancer progression by deubiquitinating and stabilizing PTEN. Mol Oncol. 2021;15:279-98
41. Chen XX, Yin Y, Cheng JW, Huang A, Hu B, Zhang X. et al. BAP1 acts as a tumor suppressor in intrahepatic cholangiocarcinoma by modulating the ERK1/2 and JNK/c-Jun pathways. Cell Death Dis. 2018;9:1036
42. Zhang Y, Shi J, Liu X, Feng L, Gong Z, Koppula P. et al. BAP1 links metabolic regulation of ferroptosis to tumour suppression. Nat Cell Biol. 2018;20:1181-92
43. Dixon SJ, Lemberg KM, Lamprecht MR, Skouta R, Zaitsev EM, Gleason CE. et al. Ferroptosis: an iron-dependent form of nonapoptotic cell death. Cell. 2012;149:1060-72
44. Carbone M, Harbour JW, Brugarolas J, Bononi A, Pagano I, Dey A. et al. Biological Mechanisms and Clinical Significance of BAP1 Mutations in Human Cancer. Cancer discovery. 2020;10:1103-20
45. Kavsak P, Rasmussen RK, Causing CG, Bonni S, Zhu H, Thomsen GH. et al. Smad7 binds to Smurf2 to form an E3 ubiquitin ligase that targets the TGF beta receptor for degradation. Mol Cell. 2000;6:1365-75
46. Park SH, Jung EH, Kim GY, Kim BC, Lim JH, Woo CH. Itch E3 ubiquitin ligase positively regulates TGF-β signaling to EMT via Smad7 ubiquitination. Molecules and cells. 2015;38:20-5
47. Zhang L, Huang H, Zhou F, Schimmel J, Pardo CG, Zhang T. et al. RNF12 controls embryonic stem cell fate and morphogenesis in zebrafish embryos by targeting Smad7 for degradation. Mol Cell. 2012;46:650-61
48. Zhang Z, Fan Y, Xie F, Zhou H, Jin K, Shao L. et al. Breast cancer metastasis suppressor OTUD1 deubiquitinates SMAD7. Nat Commun. 2017;8:2116
49. Chong PA, Lin H, Wrana JL, Forman-Kay JD. Coupling of tandem Smad ubiquitination regulatory factor (Smurf) WW domains modulates target specificity. Proceedings of the National Academy of Sciences of the United States of America. 2010;107:18404-9
50. Horn D, Hess J, Freier K, Hoffmann J, Freudlsperger C. Targeting EGFR-PI3K-AKT-mTOR signaling enhances radiosensitivity in head and neck squamous cell carcinoma. Expert opinion on therapeutic targets. 2015;19:795-805
51. Song MS, Salmena L, Pandolfi PP. The functions and regulation of the PTEN tumour suppressor. Nature reviews Molecular cell biology. 2012;13:283-96
52. He L, Liu X, Yang J, Li W, Liu S, Liu X. et al. Imbalance of the reciprocally inhibitory loop between the ubiquitin-specific protease USP43 and EGFR/PI3K/AKT drives breast carcinogenesis. Cell Res. 2018;28:934-51
53. Park SH, Fong KW, Kim J, Wang F, Lu X, Lee Y. et al. Posttranslational regulation of FOXA1 by Polycomb and BUB3/USP7 deubiquitin complex in prostate cancer. Science advances. 2021 7
54. Zheng N, Chu M, Lin M, He Y, Wang Z. USP7 stabilizes EZH2 and enhances cancer malignant progression. American journal of cancer research. 2020;10:299-313
55. Dai X, Lu L, Deng S, Meng J, Wan C, Huang J. et al. USP7 targeting modulates anti-tumor immune response by reprogramming Tumor-associated Macrophages in Lung Cancer. Theranostics. 2020;10:9332-47
56. Yu FX, Zhao B, Guan KL. Hippo Pathway in Organ Size Control, Tissue Homeostasis, and Cancer. Cell. 2015;163:811-28
57. Moya IM, Halder G. Hippo-YAP/TAZ signalling in organ regeneration and regenerative medicine. Nature reviews Molecular cell biology. 2019;20:211-26
58. Zhang Z, Du J, Wang S, Shao L, Jin K, Li F. et al. OTUB2 Promotes Cancer Metastasis via Hippo-Independent Activation of YAP and TAZ. Mol Cell. 2019;73:7-21.e7
59. Crawford JJ, Bronner SM, Zbieg JR. Hippo pathway inhibition by blocking the YAP/TAZ-TEAD interface: a patent review. Expert opinion on therapeutic patents. 2018;28:867-73
60. Koo JH, Guan KL. Interplay between YAP/TAZ and Metabolism. Cell metabolism. 2018;28:196-206
61. Cordenonsi M, Zanconato F, Azzolin L, Forcato M, Rosato A, Frasson C. et al. The Hippo transducer TAZ confers cancer stem cell-related traits on breast cancer cells. Cell. 2011;147:759-72
62. Mazzolini R, Gonzàlez N, Garcia-Garijo A, Millanes-Romero A, Peiró S, Smith S. et al. Snail1 transcription factor controls telomere transcription and integrity. Nucleic acids research. 2018;46:146-58
63. Viñas-Castells R, Beltran M, Valls G, Gómez I, García JM, Montserrat-Sentís B. et al. The hypoxia-controlled FBXL14 ubiquitin ligase targets SNAIL1 for proteasome degradation. The Journal of biological chemistry. 2010;285:3794-805
64. Wu Y, Wang Y, Lin Y, Liu Y, Wang Y, Jia J. et al. Dub3 inhibition suppresses breast cancer invasion and metastasis by promoting Snail1 degradation. Nat Commun. 2017;8:14228
65. Deng L, Meng T, Chen L, Wei W, Wang P. The role of ubiquitination in tumorigenesis and targeted drug discovery. Signal Transduct Target Ther. 2020;5:11
66. Liu J, Ma J, Liu Y, Xia J, Li Y, Wang ZP. et al. PROTACs: A novel strategy for cancer therapy. Semin Cancer Biol. 2020;67:171-9
67. An S, Fu L. Small-molecule PROTACs: An emerging and promising approach for the development of targeted therapy drugs. EBioMedicine. 2018;36:553-62
68. Sonego M, Pellarin I, Costa A, Vinciguerra GLR, Coan M, Kraut A. et al. USP1 links platinum resistance to cancer cell dissemination by regulating Snail stability. Science advances. 2019;5:eaav3235
69. Ma A, Tang M, Zhang L, Wang B, Yang Z, Liu Y. et al. USP1 inhibition destabilizes KPNA2 and suppresses breast cancer metastasis. Oncogene. 2019;38:2405-19
70. Mussell A, Shen H, Chen Y, Mastri M, Eng KH, Bshara W. et al. USP1 Regulates TAZ Protein Stability Through Ubiquitin Modifications in Breast Cancer. Cancers (Basel). 2020 12
71. Liao Y, Shao Z, Liu Y, Xia X, Deng Y, Yu C. et al. USP1-dependent RPS16 protein stability drives growth and metastasis of human hepatocellular carcinoma cells. J Exp Clin Cancer Res. 2021;40:201
72. Zhao Y, Wang X, Wang Q, Deng Y, Li K, Zhang M. et al. USP2a Supports Metastasis by Tuning TGF-β Signaling. Cell Rep. 2018;22:2442-54
73. Xiong B, Huang J, Liu Y, Zou M, Zhao Z, Gong J. et al. Ubiquitin-specific protease 2a promotes hepatocellular carcinoma progression via deubiquitination and stabilization of RAB1A. Cellular oncology (Dordrecht). 2021;44:329-43
74. Wu X, Liu M, Zhu H, Wang J, Dai W, Li J. et al. Ubiquitin-specific protease 3 promotes cell migration and invasion by interacting with and deubiquitinating SUZ12 in gastric cancer. J Exp Clin Cancer Res. 2019;38:277
75. Li F, Hu Q, He T, Xu J, Yi Y, Xie S. et al. The Deubiquitinase USP4 Stabilizes Twist1 Protein to Promote Lung Cancer Cell Stemness. Cancers (Basel). 2020 12
76. Geng N, Li Y, Zhang W, Wang F, Wang X, Jin Z. et al. A PAK5-DNPEP-USP4 axis dictates breast cancer growth and metastasis. Int J Cancer. 2020;146:1139-51
77. Meng J, Ai X, Lei Y, Zhong W, Qian B, Qiao K. et al. USP5 promotes epithelial-mesenchymal transition by stabilizing SLUG in hepatocellular carcinoma. Theranostics. 2019;9:573-87
78. Xue S, Wu W, Wang Z, Lu G, Sun J, Jin X. et al. USP5 Promotes Metastasis in Non-Small Cell Lung Cancer by Inducing Epithelial-Mesenchymal Transition via Wnt/β-Catenin Pathway. Frontiers in pharmacology. 2020;11:668
79. Zeng H, Yuan F, Mi Y, Xian G, Qin C, Zhang D. As an independent prognostic factor, USP6 promotes the invasion and metastasis of colon cancer. Biochem Biophys Res Commun. 2018;505:816-22
80. Vishnoi M, Boral D, Liu H, Sprouse ML, Yin W, Goswami-Sewell D. et al. Targeting USP7 Identifies a Metastasis-Competent State within Bone Marrow-Resident Melanoma CTCs. Cancer Res. 2018;78:5349-62
81. Wu HT, Kuo YC, Hung JJ, Huang CH, Chen WY, Chou TY. et al. K63-polyubiquitinated HAUSP deubiquitinates HIF-1α and dictates H3K56 acetylation promoting hypoxia-induced tumour progression. Nat Commun. 2016;7:13644
82. Ma M, Yu N. Ubiquitin-specific protease 7 expression is a prognostic factor in epithelial ovarian cancer and correlates with lymph node metastasis. Onco Targets Ther. 2016;9:1559-69
83. Yang X, Jin J, Yang J, Zhou L, Mi S, Qi G. Expression of Ubiquitin-specific protease 7 in oral squamous cell carcinoma promotes tumor cell proliferation and invasion. Genetics and molecular biology. 2021;44:e20210058
84. Ye M, He J, Zhang J, Liu B, Liu X, Xie L. et al. USP7 promotes hepatoblastoma progression through activation of PI3K/AKT signaling pathway. Cancer biomarkers: section A of Disease markers. 2021;31:107-17
85. Xiang M, Liang L, Kuang X, Xie Z, Liu J, Zhao S. et al. Pharmacological inhibition of USP7 suppresses growth and metastasis of melanoma cells in vitro and in vivo. J Cell Mol Med. 2021;25:9228-40
86. Jing X, Chen Y, Chen Y, Shi G, Lv S, Cheng N. et al. Down-regulation of USP8 Inhibits Cholangiocarcinoma Cell Proliferation and Invasion. Cancer management and research. 2020;12:2185-94
87. Qiu H, Kong J, Cheng Y, Li G. The expression of ubiquitin-specific peptidase 8 and its prognostic role in patients with breast cancer. J Cell Biochem. 2018;119:10051-8
88. Liu L, Yao D, Zhang P, Ding W, Zhang X, Zhang C. et al. Deubiquitinase USP9X promotes cell migration, invasion and inhibits apoptosis of human pancreatic cancer. Oncology reports. 2017;38:3531-7
89. Wu Y, Yu X, Yi X, Wu K, Dwabe S, Atefi M. et al. Aberrant Phosphorylation of SMAD4 Thr277-Mediated USP9x-SMAD4 Interaction by Free Fatty Acids Promotes Breast Cancer Metastasis. Cancer Res. 2017;77:1383-94
90. Yuan T, Chen Z, Yan F, Qian M, Luo H, Ye S. et al. Deubiquitinating enzyme USP10 promotes hepatocellular carcinoma metastasis through deubiquitinating and stabilizing Smad4 protein. Mol Oncol. 2020;14:197-210
91. Li B, Qi ZP, He DL, Chen ZH, Liu JY, Wong MW. et al. NLRP7 deubiquitination by USP10 promotes tumor progression and tumor-associated macrophage polarization in colorectal cancer. J Exp Clin Cancer Res. 2021;40:126
92. Sun H, Ou B, Zhao S, Liu X, Song L, Liu X. et al. USP11 promotes growth and metastasis of colorectal cancer via PPP1CA-mediated activation of ERK/MAPK signaling pathway. EBioMedicine. 2019;48:236-47
93. Zhang C, Xie C, Wang X, Huang Y, Gao S, Lu J. et al. Aberrant USP11 expression regulates NF90 to promote proliferation and metastasis in hepatocellular carcinoma. American journal of cancer research. 2020;10:1416-28
94. Garcia DA, Baek C, Estrada MV, Tysl T, Bennett EJ, Yang J. et al. USP11 Enhances TGFβ-Induced Epithelial-Mesenchymal Plasticity and Human Breast Cancer Metastasis. Mol Cancer Res. 2018;16:1172-84
95. Zhang S, Xie C, Li H, Zhang K, Li J, Wang X. et al. Ubiquitin-specific protease 11 serves as a marker of poor prognosis and promotes metastasis in hepatocellular carcinoma. Laboratory investigation; a journal of technical methods and pathology. 2018;98:883-94
96. Luo Q, Wu X, Nan Y, Chang W, Zhao P, Zhang Y. et al. TRIM32/USP11 Balances ARID1A Stability and the Oncogenic/Tumor-Suppressive Status of Squamous Cell Carcinoma. Cell Rep. 2020;30:98-111.e5
97. Park MK, Yao Y, Xia W, Setijono SR, Kim JH, Vila IK. et al. PTEN self-regulates through USP11 via the PI3K-FOXO pathway to stabilize tumor suppression. Nat Commun. 2019;10:636
98. Sheng B, Wei Z, Wu X, Li Y, Liu Z. USP12 promotes breast cancer angiogenesis by maintaining midkine stability. Cell Death Dis. 2021;12:1074
99. Zhang Y, Jia J, Jin W, Cao J, Fu T, Ma D. et al. Lidocaine inhibits the proliferation and invasion of hepatocellular carcinoma by downregulating USP14 induced PI3K/Akt pathway. Pathology, research and practice. 2020;216:152963
100. Hang C, Gong C, Fang Y, Chen L, Zhu J. Ubiquitin-specific protease 14 (USP14) promotes proliferation and metastasis in pancreatic ductal adenocarcinoma. Journal of molecular histology. 2021;52:187-96
101. Yao XQ, Li L, Piao LZ, Zhang GJ, Huang XZ, Wang Y. et al. Overexpression of Ubiquitin-Specific Protease15 (USP15) Promotes Tumor Growth and Inhibits Apoptosis and Correlated With Poor Disease-Free Survival in Hepatocellular Carcinoma. Technology in cancer research & treatment. 2020;19:1533033820967455
102. Zhong M, Zhou L, Fang Z, Yao YY, Zou JP, Xiong JP. et al. Ubiquitin-specific protease 15 contributes to gastric cancer progression by regulating the Wnt/β-catenin signaling pathway. World journal of gastroenterology. 2021;27:4221-35
103. Song C, Peng J, Wei Y, Shao J, Chen X, Zhang X. et al. USP18 promotes tumor metastasis in esophageal squamous cell carcinomas via deubiquitinating ZEB1. Exp Cell Res. 2021;409:112884
104. Wu C, Luo K, Zhao F, Yin P, Song Y, Deng M. et al. USP20 positively regulates tumorigenesis and chemoresistance through β-catenin stabilization. Cell Death Differ. 2018;25:1855-69
105. Chen Y, Zhou B, Chen D. USP21 promotes cell proliferation and metastasis through suppressing EZH2 ubiquitination in bladder carcinoma. Onco Targets Ther. 2017;10:681-9
106. Yun SI, Hong HK, Yeo SY, Kim SH, Cho YB, Kim KK. Ubiquitin-Specific Protease 21 Promotes Colorectal Cancer Metastasis by Acting as a Fra-1 Deubiquitinase. Cancers (Basel). 2020 12
107. Ma Y, Fu HL, Wang Z, Huang H, Ni J, Song J. et al. USP22 maintains gastric cancer stem cell stemness and promotes gastric cancer progression by stabilizing BMI1 protein. Oncotarget. 2017;8:33329-42
108. Li Y, Yang Y, Li J, Liu H, Chen F, Li B. et al. USP22 drives colorectal cancer invasion and metastasis via epithelial-mesenchymal transition by activating AP4. Oncotarget. 2017;8:32683-95
109. Zhang K, Yang L, Wang J, Sun T, Guo Y, Nelson R. et al. Ubiquitin-specific protease 22 is critical to in vivo angiogenesis, growth and metastasis of non-small cell lung cancer. Cell communication and signaling: CCS. 2019;17:167
110. Liu H, Liu N, Zhao Y, Zhu X, Wang C, Liu Q. et al. Oncogenic USP22 supports gastric cancer growth and metastasis by activating c-Myc/NAMPT/SIRT1-dependent FOXO1 and YAP signaling. Aging. 2019;11:9643-60
111. Li J, Yuan S, Norgard RJ, Yan F, Yamazoe T, Blanco A. et al. Tumor Cell-Intrinsic USP22 Suppresses Antitumor Immunity in Pancreatic Cancer. Cancer Immunol Res. 2020;8:282-91
112. Kosinsky RL, Zerche M, Saul D, Wang X, Wohn L, Wegwitz F. et al. USP22 exerts tumor-suppressive functions in colorectal cancer by decreasing mTOR activity. Cell Death Differ. 2020;27:1328-40
113. Li J, Tan Q, Yan M, Liu L, Lin H, Zhao F. et al. miRNA-200c inhibits invasion and metastasis of human non-small cell lung cancer by directly targeting ubiquitin specific peptidase 25. Molecular cancer. 2014;13:166
114. Li L, Zhou H, Zhu R, Liu Z. USP26 promotes esophageal squamous cell carcinoma metastasis through stabilizing Snail. Cancer Lett. 2019;448:52-60
115. Zhao LJ, Zhang T, Feng XJ, Chang J, Suo FZ, Ma JL. et al. USP28 contributes to the proliferation and metastasis of gastric cancer. J Cell Biochem. 2018
116. Flügel D, Görlach A, Kietzmann T. GSK-3β regulates cell growth, migration, and angiogenesis via Fbw7 and USP28-dependent degradation of HIF-1α. Blood. 2012;119:1292-301
117. Qian W, Li Q, Wu X, Li W, Li Q, Zhang J. et al. Deubiquitinase USP29 promotes gastric cancer cell migration by cooperating with phosphatase SCP1 to stabilize Snail protein. Oncogene. 2020;39:6802-15
118. Gan Q, Shao J, Cao Y, Lei J, Xie P, Ge J. et al. USP33 regulates c-Met expression by deubiquitinating SP1 to facilitate metastasis in hepatocellular carcinoma. Life Sci. 2020;261:118316
119. Buus R, Faronato M, Hammond DE, Urbe S, Clague MJ. Deubiquitinase activities required for hepatocyte growth factor-induced scattering of epithelial cells. Curr Biol. 2009;19:1463-6
120. Huang Z, Wen P, Kong R, Cheng H, Zhang B, Quan C. et al. USP33 mediates Slit-Robo signaling in inhibiting colorectal cancer cell migration. Int J Cancer. 2015;136:1792-802
121. Xia Y, Wang L, Xu Z, Kong R, Wang F, Yin K. et al. Reduced USP33 expression in gastric cancer decreases inhibitory effects of Slit2-Robo1 signalling on cell migration and EMT. Cell Prolif. 2019;52:e12606
122. Cai J, Li M, Wang X, Li L, Li Q, Hou Z. et al. USP37 Promotes Lung Cancer Cell Migration by Stabilizing Snail Protein via Deubiquitination. Front Genet. 2019;10:1324
123. Qin T, Li B, Feng X, Fan S, Liu L, Liu D. et al. Abnormally elevated USP37 expression in breast cancer stem cells regulates stemness, epithelial-mesenchymal transition and cisplatin sensitivity. Journal of Experimental & Clinical Cancer Research. 2018 37
124. Xiao Z, Chang L, Kim J, Zhang P, Hang Q, Yap S. et al. USP37 is a SNAI1 deubiquitinase. American journal of cancer research. 2019;9:2749-59
125. Liu T, Sun B, Zhao X, Li Y, Zhao X, Liu Y. et al. USP44+ Cancer Stem Cell Subclones Contribute to Breast Cancer Aggressiveness by Promoting Vasculogenic Mimicry. Mol Cancer Ther. 2015;14:2121-31
126. Park JM, Lee JE, Park CM, Kim JH. USP44 Promotes the Tumorigenesis of Prostate Cancer Cells through EZH2 Protein Stabilization. Molecules and cells. 2019;42:17-27
127. Tian M, Zhu R, Ding F, Liu Z. Ubiquitin-specific peptidase 46 promotes tumor metastasis through stabilizing ENO1 in human esophageal squamous cell carcinoma. Exp Cell Res. 2020;395:112188
128. Silvestrini VC, Thome CH, Albuquerque D, de Souza Palma C, Ferreira GA, Lanfredi GP. et al. Proteomics analysis reveals the role of ubiquitin specific protease (USP47) in Epithelial to Mesenchymal Transition (EMT) induced by TGFbeta2 in breast cells. J Proteomics. 2020;219:103734
129. Choi BJ, Park SA, Lee SY, Cha YN, Surh YJ. Hypoxia induces epithelial-mesenchymal transition in colorectal cancer cells through ubiquitin-specific protease 47-mediated stabilization of Snail: A potential role of Sox9. Sci Rep. 2017;7:15918
130. Yu L, Dong L, Wang Y, Liu L, Long H, Li H. et al. Reversible regulation of SATB1 ubiquitination by USP47 and SMURF2 mediates colon cancer cell proliferation and tumor progression. Cancer Lett. 2019;448:40-51
131. Lei X, Li X, Chen H, Liu Z. USP48 Sustains Chemoresistance and Metastasis in Ovarian Cancer. Current cancer drug targets. 2020;20:689-99
132. Zhang Z, Li J, Ou Y, Yang G, Deng K, Wang Q. et al. CDK4/6 inhibition blocks cancer metastasis through a USP51-ZEB1-dependent deubiquitination mechanism. Signal Transduct Target Ther. 2020;5:25
133. Zhou Z, Zhang P, Hu X, Kim J, Yao F, Xiao Z. et al. USP51 promotes deubiquitination and stabilization of ZEB1. American journal of cancer research. 2017;7:2020-31
134. Che X, Jian F, Jia N, Zheng Y, Jiang Y, Feng W. FAT4-USP51 complex regulates the proliferation and invasion of endometrial cancer via Hippo pathway. American journal of translational research. 2019;11:2784-800
135. Fraile JM, Campos-Iglesias D, Rodríguez F, Español Y, Freije JM. The deubiquitinase USP54 is overexpressed in colorectal cancer stem cells and promotes intestinal tumorigenesis. Oncotarget. 2016;7:74427-34
136. Zhou H, Liu Y, Zhu R, Ding F, Cao X, Lin D. et al. OTUB1 promotes esophageal squamous cell carcinoma metastasis through modulating Snail stability. Oncogene. 2018;37:3356-68
137. Zhou Y, Wu J, Fu X, Du W, Zhou L, Meng X. et al. OTUB1 promotes metastasis and serves as a marker of poor prognosis in colorectal cancer. Molecular Cancer. 2014;13:258
138. Li J, Cheng D, Zhu M, Yu H, Pan Z, Liu L. et al. OTUB2 stabilizes U2AF2 to promote the Warburg effect and tumorigenesis via the AKT/mTOR signaling pathway in non-small cell lung cancer. Theranostics. 2019;9:179-95
139. Zhang Z, Du J, Wang S, Shao L, Jin K, Li F. et al. OTUB2 Promotes Cancer Metastasis via Hippo-Independent Activation of YAP and TAZ. Mol Cell. 2019;73:7-21 e7
140. Zhang P, Li C, Li H, Yuan L, Dai H, Peng Z. et al. Ubiquitin ligase CHIP regulates OTUD3 stability and suppresses tumour metastasis in lung cancer. Cell Death Differ. 2020;27:3177-95
141. Yuan L, Lv Y, Li H, Gao H, Song S, Zhang Y. et al. Deubiquitylase OTUD3 regulates PTEN stability and suppresses tumorigenesis. Nat Cell Biol. 2015;17:1169-81
142. Liu X, Zhang X, Peng Z, Li C, Wang Z, Wang C. et al. Deubiquitylase OTUD6B Governs pVHL Stability in an Enzyme-Independent Manner and Suppresses Hepatocellular Carcinoma Metastasis. Adv Sci (Weinh). 2020;7:1902040
143. Lin DD, Shen Y, Qiao S, Liu WW, Zheng L, Wang YN. et al. Upregulation of OTUD7B (Cezanne) Promotes Tumor Progression via AKT/VEGF Pathway in Lung Squamous Carcinoma and Adenocarcinoma. Front Oncol. 2019;9:862
144. Qin J, Zhou Z, Chen W, Wang C, Zhang H, Ge G. et al. BAP1 promotes breast cancer cell proliferation and metastasis by deubiquitinating KLF5. Nat Commun. 2015;6:8471
145. Amato E, Mafficini A, Hirabayashi K, Lawlor RT, Fassan M, Vicentini C. et al. Molecular alterations associated with metastases of solid pseudopapillary neoplasms of the pancreas. J Pathol. 2019;247:123-34
146. Gupta MP, Lane AM, DeAngelis MM, Mayne K, Crabtree M, Gragoudas ES. et al. Clinical Characteristics of Uveal Melanoma in Patients With Germline BAP1 Mutations. JAMA Ophthalmol. 2015;133:881-7
147. Harbour JW, Onken MD, Roberson ED, Duan S, Cao L, Worley LA. et al. Frequent mutation of BAP1 in metastasizing uveal melanomas. Science (New York, NY). 2010;330:1410-3
148. de Jel MM, Schott M, Lamm S, Neuhuber W, Kuphal S, Bosserhoff AK. Loss of CYLD accelerates melanoma development and progression in the Tg(Grm1) melanoma mouse model. Oncogenesis. 2019;8:56
149. Miliani de Marval P, Lutfeali S, Jin JY, Leshin B, Selim MA, Zhang JY. CYLD inhibits tumorigenesis and metastasis by blocking JNK/AP1 signaling at multiple levels. Cancer prevention research (Philadelphia, Pa). 2011;4:851-9
150. Massoumi R, Kuphal S, Hellerbrand C, Haas B, Wild P, Spruss T. et al. Down-regulation of CYLD expression by Snail promotes tumor progression in malignant melanoma. J Exp Med. 2009;206:221-32
151. Goto Y, Zeng L, Yeom CJ, Zhu Y, Morinibu A, Shinomiya K. et al. UCHL1 provides diagnostic and antimetastatic strategies due to its deubiquitinating effect on HIF-1α. Nat Commun. 2015;6:6153
152. Jang MJ, Baek SH, Kim JH. UCH-L1 promotes cancer metastasis in prostate cancer cells through EMT induction. Cancer Lett. 2011;302:128-35
153. Kim HJ, Kim YM, Lim S, Nam YK, Jeong J, Kim HJ. et al. Ubiquitin C-terminal hydrolase-L1 is a key regulator of tumor cell invasion and metastasis. Oncogene. 2009;28:117-27
154. Wang W, Zou L, Zhou D, Zhou Z, Tang F, Xu Z. et al. Overexpression of ubiquitin carboxyl terminal hydrolase-L1 enhances multidrug resistance and invasion/metastasis in breast cancer by activating the MAPK/Erk signaling pathway. Mol Carcinog. 2016;55:1329-42
155. Seliger B, Fedorushchenko A, Brenner W, Ackermann A, Atkins D, Hanash S. et al. Ubiquitin COOH-terminal hydrolase 1: a biomarker of renal cell carcinoma associated with enhanced tumor cell proliferation and migration. Clin Cancer Res. 2007;13:27-37
156. Liu S, González-Prieto R, Zhang M, Geurink PP, Kooij R, Iyengar PV. et al. Deubiquitinase Activity Profiling Identifies UCHL1 as a Candidate Oncoprotein That Promotes TGFβ-Induced Breast Cancer Metastasis. Clin Cancer Res. 2020;26:1460-73
157. Liu D, Song Z, Wang X, Ouyang L. Ubiquitin C-Terminal Hydrolase L5 (UCHL5) Accelerates the Growth of Endometrial Cancer via Activating the Wnt/β-Catenin Signaling Pathway. Front Oncol. 2020;10:865
158. Huang M, Xiong H, Luo D, Xu B, Liu H. CSN5 upregulates glycolysis to promote hepatocellular carcinoma metastasis via stabilizing the HK2 protein. Exp Cell Res. 2020;388:111876
159. Zhang S, Hong Z, Chai Y, Liu Z, Du Y, Li Q. et al. CSN5 promotes renal cell carcinoma metastasis and EMT by inhibiting ZEB1 degradation. Biochem Biophys Res Commun. 2017;488:101-8
160. Hou J, Deng Q, Zhou J, Zou J, Zhang Y, Tan P. et al. CSN6 controls the proliferation and metastasis of glioblastoma by CHIP-mediated degradation of EGFR. Oncogene. 2017;36:1134-44
161. Mao Z, Sang MM, Chen C, Zhu WT, Gong YS, Pei DS. CSN6 Promotes the Migration and Invasion of Cervical Cancer Cells by Inhibiting Autophagic Degradation of Cathepsin L. Int J Biol Sci. 2019;15:1310-24
162. Zou H, Chen H, Zhou Z, Wan Y, Liu Z. ATXN3 promotes breast cancer metastasis by deubiquitinating KLF4. Cancer Lett. 2019;467:19-28
Author contact
Corresponding authors: Feng Gao, E-mail: gf0731edu.cn. Pengfei Rong, E-mail: rongpengfei66com. Wei Wang, E-mail: cjr.wangwei163.com. Wei Li, E-mail: weililxedu.cn. Tel: +86-0731-88618643

 Global reach, higher impact
Global reach, higher impact