Impact Factor ISSN: 1449-2288
Int J Biol Sci 2022; 18(7):3006-3018. doi:10.7150/ijbs.69240 This issue Cite
Research Paper
Stella Regulates the Development of Female Germline Stem Cells by Modulating Chromatin Structure and DNA Methylation
1. Bio-X Institutes, Key Laboratory for the Genetics of Developmental and Neuropsychiatric Disorders, Ministry of Education, Shanghai Jiao Tong University, Shanghai 200240, China.
2. Key Laboratory of Fertility Preservation and Maintenance of Ministry of Education, School of Basic Medical Sciences, Ningxia Medical University, Yinchuan 750004, China.
3. Shanghai Key Laboratory of Reproductive Medicine, Shanghai 200025, China
Abstract
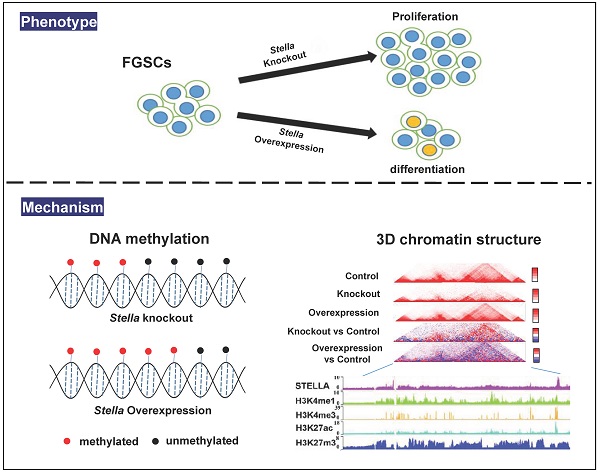
Female germline stem cells (FGSCs) have the ability to self-renew and differentiate into oocytes. Stella, encoded by a maternal effect gene, plays an important role in oogenesis and early embryonic development. However, its function in FGSCs remains unclear. In this study, we showed that CRISPR/Cas9-mediated knockout of Stella promoted FGSC proliferation and reduced the level of genome-wide DNA methylation of FGSCs. Conversely, Stella overexpression led to the opposite results, and enhanced FGSC differentiation. We also performed an integrative analysis of chromatin immunoprecipitation followed by high-throughput sequencing (ChIP-seq), high-throughput genome-wide chromosome conformation capture (Hi-C), and use of our published epigenetic data. Results indicated that the binding sites of STELLA and active histones H3K4me3 and H3K27ac were enriched near the TAD boundaries. Hi-C analysis showed that Stella overexpression attenuated the interaction within TADs, and interestingly enhanced the TAD boundary strength in STELLA-associated regions. Taking these findings together, our study not only reveals the role of Stella in regulating DNA methylation and chromatin structure, but also provides a better understanding of FGSC development.
Keywords: Stella, female germline stem cells, epigenomics, chromatin structure, DNA methylation

 Global reach, higher impact
Global reach, higher impact