Impact Factor ISSN: 1449-2288
Int J Biol Sci 2022; 18(16):6068-6083. doi:10.7150/ijbs.73431 This issue Cite
Research Paper
GALNT1 Enhances Malignant Phenotype of Gastric Cancer via Modulating CD44 Glycosylation to Activate the Wnt/β-catenin Signaling Pathway
1. Digestive Disease Center, The Seventh Affiliated Hospital of Sun Yat-Sen University, Shenzhen, Guangdong 518107, P.R. China.
2. Department of Gastrointestinopancreatic Surgery, The First Affiliated Hospital of Sun Yat-Sen University, Guangzhou, Guangdong 510080, P.R. China.
*These authors have contributed equally to this work.
Received 2022-3-30; Accepted 2022-9-24; Published 2022-10-17
Abstract
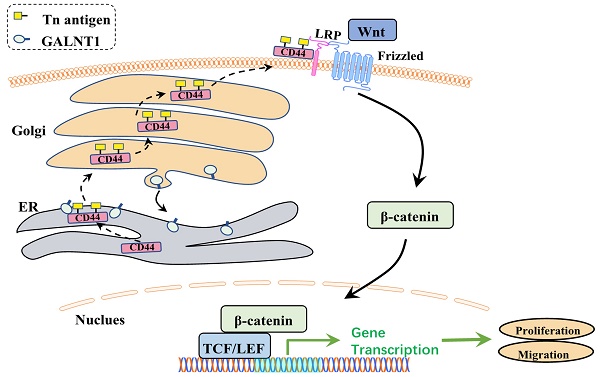
O-glycosylation is a widespread post-translational modification of proteins. Aberrant O-glycosylation is a hallmark of cancer. Here, we show that the polypeptide N-acetylgalactosamine-transferase 1 (GALNT1) is frequently upregulated in gastric cancer and is correlated with poor survival. Overexpression of GALNT1 promoted, whereas knockdown suppressed proliferation, migration, and invasion of gastric cancer cells in vitro and in vivo. Mechanistically, GALNT1 enhances aberrant initiation of O-glycosylation and results in CD44 glycoproteins modified with abundant Tn antigens, thereby activating the Wnt/β-catenin signaling pathway. Collectively, this study demonstrates that GALNT1 overexpression in gastric cancer promotes the Wnt/β-catenin signaling pathway via abnormal O-glycosylation of CD44 to enhance malignancy, providing a novel strategy for the development of therapeutic reagents against gastric cancer.
Keywords: Gastric cancer, GALNT1, CD44, O-Glycosylation, Wnt/β-catenin signaling pathway
Introduction
Gastric cancer (GC) is the fifth most frequent tumor and the fourth leading cause of cancer-related mortality worldwide, despite its decreasing incidence over the past 50 years [1]. Although there have been considerable improvements in diagnosis and therapy for GC, metastasis remains the primary cause of death in patients with GC [2], rendering prognosis challenging. Invasive tumor phenotypes are often associated with metastasis and poor survival. Therefore, elucidating the mechanisms that drive the aggressive phenotype of malignant GC progression is vital.
Glycosylation, one of the most prevalent post-translational modifications in proteins, is frequently altered in tumors and associated with tumor progression [3, 4]. A frequent and noticeable characteristic of malignancy is the abundance of shortened O-glycans, such as the Tn antigen, which can be recognized by vicia villosa (VVA) lectins and is formed by aberrant initiation of N-acetylgalactosamine (GalNAc)-type O-glycosylation [5, 6]. Accumulated studies indicate that the Tn antigen is abnormally increased in human cancers, including breast cancer, lung cancer, and GC tissues [7-9], which is strongly correlated with cancer progression characteristics, such as prognosis, invasion, metastasis, and immune escape [10, 11]. The above studies indicate that O- glycosylation is intimately associated with the emergence and progression of cancers. Hence, understanding the involvement of O-glycosylation in GC may open new avenues for elucidating the pathogenesis of gastric cancer.
Polypeptide N-acetylgalactosamine-transferases (GALNTs) are abnormally expressed in multiple cancers and affect tumor progression by initiating GalNAc-type O-glycosylation of various proteins [12-15]. Moreover, the initiation of O-glycosylation had previously been demonstrated to begin in the Golgi apparatus [16]. In contrast, the initiation of O-glycosylation is associated with GALNTs relocated from the Golgi apparatus to the endoplasmic reticulum (ER) in tumor progression and metastasis (GALNTs activation [GALA] pathway) [8, 17, 18]. GALNT1, as a member of the GALNT family, initiates mucin-type O-glycosylation by transferring α-GalNAc from UDP-GalNAc to serine (Ser) or threonine (Thr) residues in proteins [11, 19]. Importantly, GALNT1 plays a vital role in cancer growth and metastasis by modifying the O-glycosylation of various glycoproteins, such as mucin 1 (MUC1), osteopontin (OPN), matrix metalloproteinase-14 (MMP14), and integrin α3 [17, 18, 20]. These reports suggest that GALNT1 may be an efficient prognostic biomarker and a therapeutic target for treating a variety of malignancies. However, the function and detailed molecular mechanisms of GALNT1 remain largely unknown.
Therefore, the goal of this study was to explore the underlying mechanism by which GALNT1 controls GC progression and assess its therapeutic potential.
Materials and methods
Clinical data and tissue samples
At the First Affiliated Hospital of Sun Yat‐sen University, tissue samples from 150 patients with GC with detailed medical records were collected between January 2010 and December 2013. Twenty-three fresh GC samples and their neighboring normal samples were collected at the Seventh Affiliated Hospital of Sun Yat-sen University. Total RNA and total protein were extracted from 13 and 10 pairs of tissue samples, respectively. Informed consent was obtained from patients involved in the study. This study was approved by the Ethics Committees of the First and Seventh Affiliated Hospitals of Sun Yat-sen University.
Cell culture
We used one gastric epithelial cell line (GES1) and eight GC cell lines (AGS, MGC803, KATO III, BGC803, MKN28, SNU719, HGC27 and SGC7901) obtained from the Chinese Academy of Sciences (Shanghai, China). AGS cells were cultured in DMEM/F12 (Gibco, CAT# C11330500BT, Beijing, China) and 10% fetal bovine serum (Gibco, CAT# 10099141C, Beijing, China). Other cells were cultivated in RPMI‐1640 (Gibco, CAT# C11875500BT, Beijing, China) with 10% fetal bovine serum. All the cells were maintained at 37 °C in a 5% CO2 cell culture incubator.
Cell transfection and lentiviral infection
GeneChem (Shanghai, China) provided lentiviral vectors for GALNT1 knockdown (shGALNT1), whereas those for CD44 knockdown (shCD44) were purchased from GeneCopoeia (Guangzhou, China) (Table S1). The GALNT1 expressing lentiviral vector was constructed by cloning GALNT1 into the GV341 vector (GeneChem, Shanghai, China). Lentiviral transduction was generated and purified according to the manufacturer's instructions. Puromycin (2 μg/ml) was added to screen the transgenic cells.
Quantitative real-time PCR
Total RNA from GC cells and fresh tissues was extracted with AG RNAex Pro RNA reagent (Accurate Biology, CAT#AG21102, Hunan, China) according to the manufacturer's instructions. RNA was reverse transcribed to cDNA using the Evo M‐MLV reverse transcription master mix (Accurate Biology, CAT#AG11706, Hunan, China). Quantitative real-time PCR (qRT-PCR) was performing using a SYBR Green Pro Taq HS premixed qPCR kit (Accurate Biology, CAT# AG11702, Hunan, China). Primer sequences are listed in Supplementary Table S2. The 2-ΔΔCT technique was used to analyze the data.
Western blotting and antibodies
Protein lysis buffer was used to extract total proteins from GC cells and fresh tissues on ice for 30 min in accordance with the procedure. A nuclear and cytoplasmic protein extraction kit (Beyotime Biotechnology, CAT#P0027, Shanghai, China) was used to fractionate the nuclear and cytoplasmic fractions as per the manufacturer's instructions. Protein concentrations were quantified using the BCA protein assay kit (Biosharp, CAT# BL521A). Equal quantities of proteins were fractionated by 10% SDS-PAGE and transferred to PVDF or NC membrane. The transferred protein was blotted with the corresponding antibody overnight at 4 °C after blocking with 5% skim milk for 1 h. The following primary antibodies were used: anti-GALNT1 (1:500, Abcam, CAT#ab253025), anti-GAPDH (1:2000, Proteintech, CAT#10494-1-AP), anti-CD44 (1:2000, Proteintech, CAT# 15675-1-AP-100), anti-β-catenin (1:5000, Proteintech, CAT#51067-2-AP), anti-phospho-β-catenin (1:1000, Cell Signaling Technology, CAT#4176S), anti-c-Myc (1:2000, Proteintech, CAT#10828-1-AP), anti-Lamin B1(1:1000, Cell Signaling Technology, CAT#13435), and VVA (2 μg/ml, Vector Laboratories, CAT#B-1235-2). HRP‐conjugated anti‐rabbit IgG (1:1000, Beyotime, CAT#A0208), anti‐mouse IgG (1:1000, Beyotime, CAT#A0216), and Streptavidin HRP (1:10000, Abcam, CAT#ab7403-500) were used as the secondary antibodies. Proteins were detected using an extremely sensitive ECL assay kit (Meilunbio, CAT#MA0186-2).
Immunohistochemistry
Immunohistochemistry (IHC) was performed according to a standard protocol. Tumor tissue sections were deparaffinized in xylol and rehydrated with a stepwise decrease in ethanol concentration. The sections were then heated in a pressure cooker for antigen retrieval with citric acid buffer and cooled naturally to room temperature. After washing with distilled water, the tissue was treated with 3.0% hydrogen peroxide for 15 min at room temperature, followed by 30 min incubation with 5% goat serum. After washing thrice with phosphate-buffered saline (PBS), rabbit anti-GALNT1 (1:200, Abcam, CAT# ab253025) was used to incubate the sections overnight at 4 °C. Subsequently, the sections were incubated with horseradish peroxidase (HRP) (goat anti-rabbit/mouse, CAT#A0208, Beyotime) for one hour at room temperature after washing in PBS containing Tween-20 (PBST). After 3,3'-diaminobenzidine (DAB) and hematoxylin staining, GALNT1 expression was evaluated using an inverted microscope. VVA expression in GC tissues was also detected using IHC as described above. Non-specific binding was blocked by incubating the sections with Carbo-Free™ Blocking Solution (Vector Laboratories, CAT#SP-5040) for 30 min at room temperature. VVA (1:2000, Vector Laboratories, CAT#B-1235-2) was used to incubate the sections overnight at 4 °C, then streptavidin HRP (1:10000, Abcam, CAT# ab7403) for 1h at room temperature.
Immunofluorescence assay
After fixing in 4% paraformaldehyde for 10 min, cells were treated with 0.2% Triton X-100 for 10 min at room temperature, followed by blocking with goat serum at 37 °C for 60 min. The cells were then immunostained with the appropriate antibody or lectin. The following primary antibodies or lectins were used: anti-GALNT1 (1:200; Abnova, CAT#H00002589-M10), anti-β-catenin (1:200; Proteintech, CAT#51067), anti-GM130 (1:200; Proteintech, CAT#11308-1-AP-150), anti-PDI (1:200; Cell Signaling Technology, CAT# 3501T), and VVA (1:200; Vector Laboratories, CAT# B-1235-2). Alexa Fluor 546-conjugated donkey anti‐rabbit IgG (1:1000, Thermo Fisher, CAT#A11003), DyLight™ 488 streptavidin (1:200, BioLegend, CAT# 405218), and Alexa Fluor 488-conjugated donkey anti-mouse IgG (1:1000, Thermo Fisher, CAT#A11001) were used as secondary antibodies. DAPI nucleic acid stain (Abcam, CAT# ab104139) was used to counterstain the sections. An inverted microscope was used to capture the images.
Immunoprecipitation assays
The cells were lysed with IP Lysis Buffer (Pierce, CAT#87788), followed by immunoprecipitation with Protein A/G Magnetic Beads (Biomake, CAT#B23202-5) conjugated with CD44 immunoprecipitation specific polyclonal antibody. Then, they were incubated overnight at 4 °C and displayed via western blotting with biotin-conjugated VVA, as described above. Similarly, total cell lysates were immunoprecipitated using streptavidin-agarose (Sigma, CAT#S1638-1ML) conjugated with VVA immunoprecipitation-specific lectin. The immunocomplexes were then analyzed by western blotting.
Luciferase reporter assay
Super TOP flash plasmids (Beyotime, CAT#D2506) and Renilla TK-luciferase vector were co-transfected into cells by Lipofectamine 6000 (Beyotime, CAT# C0526). The luciferase assay was performed using the Dual Luciferase Reporter Gene Assay Kit (YEASEN, CAT#11402ES60*) according to the manufacturer's protocol. And the firefly luciferase activity was normalized to Renilla.
Cell viability and colony formation assay
A total of 1000-1500 cells per well were seeded in 96-well plates. Next, cell counting kit-8 (CCK-8) was used to determine the viability of cells incubated at 37 °C for 2 h at 0, 24, 48, 72, and 96 h, in accordance with the manufacturer's protocol. For colony formation assays, six-well plates containing 500 cells per well were used for colony formation experiments, and the cells were cultivated for two weeks. The colonies were then fixed with 4% paraformaldehyde for 15 min and stained with crystal violet (Beyotime Biotechnology, CAT#C0121) for 20 min.
Cell migration and invasion assays
Transwell plates with 8-micrometer holes (Corning) were used to conduct migration and invasion assays. Cells were seeded in the top compartment with 0.2 ml serum-free medium, whereas 0.8 ml culture media with 10% fetal bovine serum was injected into the bottom chamber for the migration assay. When the cells grew for 24 h in an incubator, crystal violet staining was performed for 20 min with 4% paraformaldehyde for 15 min. Cotton swabs were used to remove the remaining unmigrated cells from the top layer. The protocols for the invasion experiments were roughly the same as for the migration assays, except for a 10% Matrigel (BD Biosciences) precoated in the top compartment and a doubled number of seeded cells. A 10× microscope (Olympus CKX53) was used to view and capture migrating or invading cells. ImageJ (X64) was used to count cells in the images.
Animal studies
The TOPBIOTECH Animal Model Center's Experimental Animal Ethics Committee authorized in vivo studies, which were carried out in accordance with the institution's ethical rules for animal experimentation. Four-week-old female BALB/c mices were inoculated subcutaneously in the right anterior axilla with control (n = 6) or GALNT1-knockdown (n = 6) stably transfected SGC7901 cells (5.0 × 106 cells). After 28 days, the mice were sacrificed. In the same time, four-week-old female BALB/c mice were injected with control (n = 7) or GALNT1-knockdown (n = 7) stably transfected SGC7901 cells (1.5 × 106 cells) via the tail vein. After 7 weeks, the mice were euthanized, and the number of metastatic lung nodules was recorded.
Statistical analysis
Unless otherwise stated, the data are presented as mean ± SD. Statistical Package for the Social Sciences (SPSS, version 26.0) and GraphPad Prism (Version 8.0) were used for all statistical analyses. To compare the two groups, the student's t-test was used. Paired t-tests were used to analyze paired GC tissues. The student's t-test was also used to examine the correlation between GALNT1 or VVA expression and the clinicopathological characteristics of patients with GC. OS and DFS were examined by performing Kaplan-Meier curves, log-rank tests, and univariate/multivariate Cox regression analyses. Statistical significance was set at p< 0.05 (***p< 0.001, **p< 0.01, *p< 0.05).
Results
GALNT1 is identified as a candidate target in GC by bioinformatics analysis
To investigate the potential targets of O-glycosylation transferases in GC, we first investigated the expression of GALNT1-20 in GC from The Cancer Genome Atlas (TCGA) database. Our study comprised 211 healthy tissues and 408 samples from patients with GC and their clinical medical records. Interestingly, the analysis revealed different expression patterns of these genes in the GC tissues. GALNT1, GALNT3, GALNT4, GALNT5, and GALNT6 were significantly higher in GC tissues, whereas GALNT10 expression was higher in normal tissues (Fig. S1, p<0.05). In the TCGA pan-cancer database, increased GALNT1 expression was frequently observed in various cancers (Fig. 1A-B). Among these, we mainly focused on GALNT1 because its function in GC remains elusive. Therefore, the emphasis of this study was on the impact and molecular mechanisms of GALNT1 in GC.
GALNT1 as a candidate target in gastric cancer by bioinformatics analysis. A, GALNT1 expression in different types of human cancers. B, Analysis of GALNT1 expression in gastric cancer using the GEPIA database. C-E, Analysis of GALNT1 expression grouped by gastric cancer grade (C), pathological stages (D), and lymph node metastasis status (E). *** p<0.001. F, Overall survival comparing low and high expression of GALNT1 in gastric cancer from TCGA datasets is shown. G, The receiver-operating characteristic (ROC) curve analysis of GALNT1 in patients with gastric cancer from TCGA datasets.
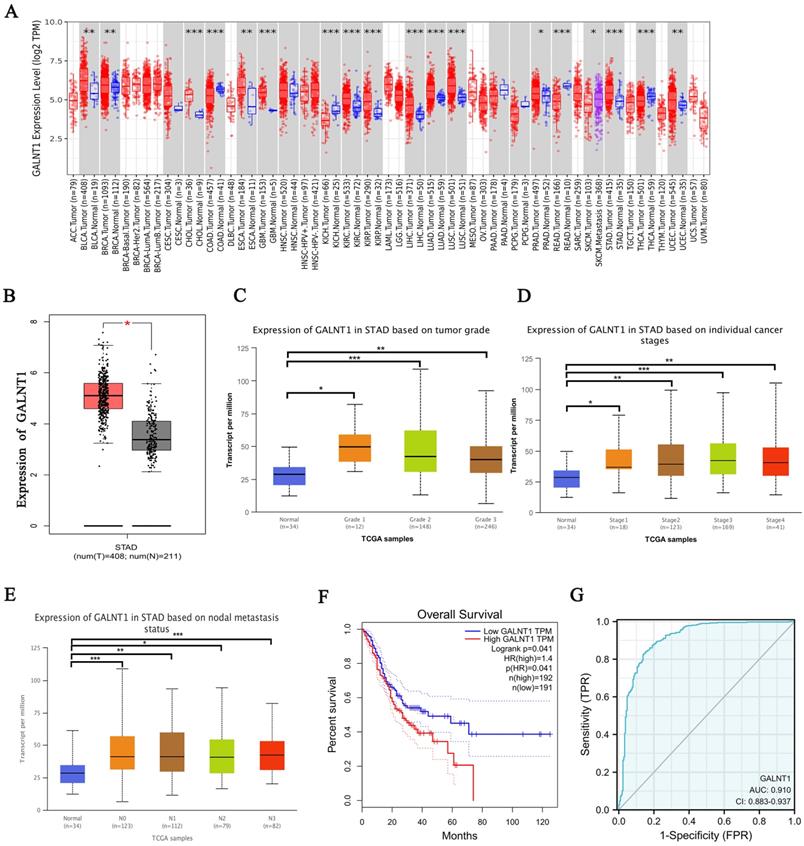
Analysis of TCGA database by applying UALCAN, we examined GALNT1 expression in relation to several clinicopathological parameters in patients with GC, including tumor grade (grade 1, 2, 3), tumor stage (stages I, II, III, and IV), and lymph node stage (N0 1, 2, and 3). The data indicated that GALNT1 expression in any grade and stage was higher than that in the normal tissues (Fig. 1C-D). Moreover, GALNT1 expression was related to lymph node metastasis, with the highest expression observed in N3 GC (Fig. 1E). Furthermore, the study suggested that patients with GC who had higher GALNT1 expression had worse overall survival (OS) by using the Gene Expression Profiling Interactive Analysis (GEPIA) database (Fig. 1F). Interestingly, the receiver-operating characteristic (ROC) curve analysis of GALNT1 in patients with GC from TCGA datasets confirmed that GALNT1 has good diagnostic accuracy with an area under the curve (AUC) of 0.91 (95% CI: 0.883-0.937) (Fig. 1G). The above data indicate that GALNT1 may be a candidate target in GC.
Upregulated GALNT1 expression promotes abnormal O-glycosylation in GC and correlates with aggressive clinicopathological features and poor survival
To confirm the above findings, paired GC tissues from 13 patients were analyzed using qRT-PCR. The results showed that 76.9% of GC tissues showed a positive increase in GALNT1 mRNA expression compared to paired normal tissues (Fig. 2A). Consistently, western blotting analysis found that protein levels of GALNT1 were elevated in GC tissues of matched specimens (Fig. 2B). To further investigate the relationship between GALNT1 levels and clinicopathological characteristics, 150 GC tissue samples were used for immunohistochemical analyses (Fig. 2C). Immunohistochemical analysis of VVA in 150 GC specimens was performed to further investigate glycosylation levels (Fig. 2C). The results revealed that GALNT1 and VVA were mainly expressed in the cytoplasm of GC cells and were considerably greater in primary tumor tissues than in neighboring normal tissues. Interestingly, correlation analyses between GALNT1 and VVA levels based on the immunohistochemical scores showed that increased GALNT1 levels promote abnormal O-glycosylation (Fig. 2D). Similarly, immunofluorescence analyses also showed that Tn levels were upregulated in GC cells (Fig. 2E).
The expression of GALNT1 and VVA in patients with gastric cancer. A, The mRNA levels of GALNT1 in 13 paired gastric cancer tissues and adjacent normal tissues (normalized to GAPDH). B, Western blot analysis depicting the GALNT1 levels in paired primary tumors and adjacent normal tissues. C, Representative immunohistochemical images of GALNT1 and VVA expression in samples and para-carcinoma tissues of patients with gastric cancer. D, Correlation analysis between GALNT1 and VVA immunohistochemistry score. E, Upregulated VVA expression in paired primary tumor tissues compared to adjacent normal tissues as examined by immunofluorescence assay. F-I, Kaplan-Meier analysis of overall survival and disease-free survival for patients with gastric cancer according to the immunohistochemistry of GALNT1 and VVA.
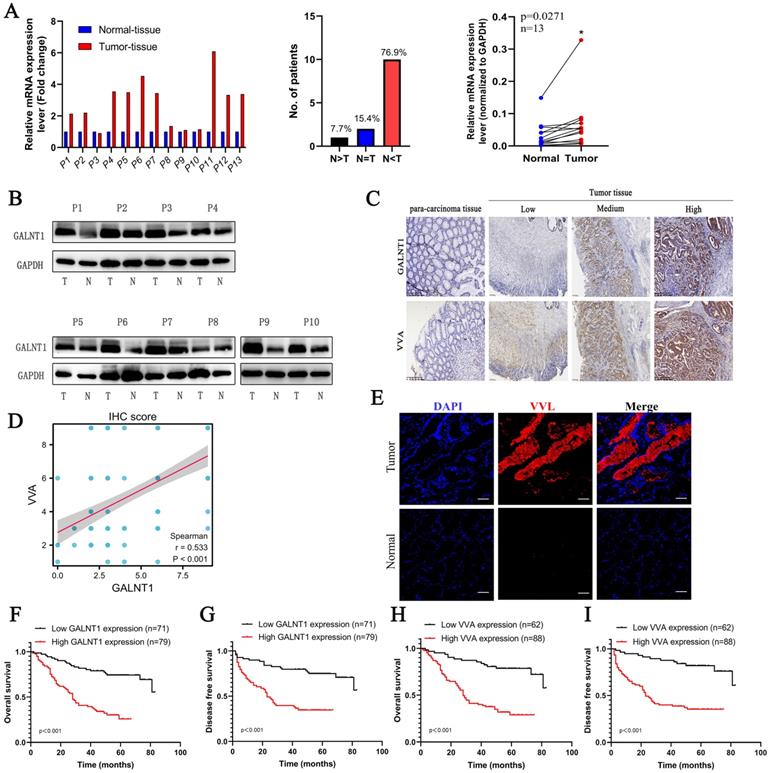
In addition, we analyzed the association between GALNT1 levels and clinicopathological factors (Table 1) and found that patients with larger tumor size (≥5 cm; p=0.002), Borrmann type (III/IV; p=0.005), pathological tumor-node-metastasis (pTNM) stage (III/IV; p<0.001), depth of invasion (T3/4; p=0.001), lymph node metastasis (p<0.001), distant metastasis (p=0.005), and lymphovascular invasion (LVI; p=0.022) tended to have higher levels of GALNT1 in primary tumors (Table 1). Similar results were observed for VVA expression in GC (Table 1). To further understand the clinical importance of GALNT1 and VVA in GC, we assessed the prognoses associated with GALNT1 and VVA expression by Kaplan-Meier analysis. Patients with higher GALNT1 and VVA expression had poorer OS and disease-free survival (DFS) than those with lower GALNT1 and VVA expression levels (log-rank test, p<0.001; Fig. 2F-I). Multivariate Cox regression analysis indicated that tumor size (hazard ratio (HR): 2.098, 95%CI: 1.201-3.664, p=0.009; DFS: HR:2.057, 95%CI: 1.155-3.664, p=0.014), lymph node metastasis (OS: HR:2.965, 95%CI: 1.183-7.436, p=0.02; DFS: HR: 2.817, 95%CI: 1.116-7.114, p=0.028), and high GALNT1 expression (OS: HR:2.458, 95%CI: 1.297-4.660, p=0.006; DFS: HR: 2.542, 95%CI: 1.335-4.841, p=0.005) were independent poor prognostic factors for OS and DFS (Tables 2 and 3). Collectively, these data suggest that GALNT1 is frequently upregulated in GC and promotes abnormal O-glycosylation, which is associated with poor survival. This indicates that GALNT1 is a potential prognostic marker for GC.
Associations of GALNT1 and VVA expression with clinical parameters in GC patients
| Characteristic | GALNT1 | VVA | ||||
|---|---|---|---|---|---|---|
| Low (%) | High (%) | P | Low (%) | High (%) | P | |
| Age (years) | 0.403 | 0.471 | ||||
| <60 | 32(43.8) | 41(56.2) | 28(38.4) | 45(61.6) | ||
| ≥60 | 39(50.6) | 38(49.4) | 34(44.2) | 43(55.8) | ||
| Gender | 0.327 | 0.418 | ||||
| Male | 46(50.5) | 45(49.5) | 40(44.0) | 51(56.0) | ||
| Female | 25(42.4) | 34(57.6) | 22(37.3) | 37(26.7) | ||
| Tumor size | <0.001 | 0.015 | ||||
| ≤5 cm | 52(60.5) | 34(39.5) | 43(50.0) | 43(50.0) | ||
| >5 cm | 19(30.2) | 31(69.8) | 19(30.2) | 44(69.8) | ||
| Borrmann type | 0.005 | 0.023 | ||||
| I-II | 20(76.9) | 6(23.1) | 15(57.7) | 11(42.3) | ||
| III-IV | 40(45.4) | 48(54.5) | 29(33.0) | 59(67.0) | ||
| Differentiation | 0.666 | 0.617 | ||||
| Well+ moderate | 19(55.9) | 15(44.1) | 16(47.1) | 18(52.9) | ||
| poor | 49(51.6) | 46(48.4) | 40(42.1) | 55(57.9) | ||
| pTNM stage | <0.001 | <0.001 | ||||
| I | 20(74.1) | 7(25.9) | 20(74.1) | 7(25.9) | ||
| II | 28(87.5) | 4(12.5) | 21(65.6) | 11(34.4) | ||
| III | 21(27.6) | 55(72.4) | 18(23.7) | 58(76.3) | ||
| IV | 2(13.3) | 13(86.7) | 3(20.0) | 12(80.0) | ||
| Depth of invasion | 0.001 | <0.001 | ||||
| T1/2 | 24(72.7) | 9(27.3) | 24(72.7) | 9(27.3) | ||
| T3/4 | 47(40.2) | 70(59.8) | 38(32.5) | 79(67.5) | ||
| Lymph node metastasis | <0.001 | <0.001 | ||||
| N0 | 41(82.0) | 9(18.0) | 35(70.0) | 15(30.0) | ||
| N+ | 30(30.0) | 70(70.0) | 27(27.0) | 73(73.0) | ||
| Distant metastasis | 0.005 | 0.077 | ||||
| M0 | 69(51.1) | 66(48.9) | 59(43.7) | 76(56.3) | ||
| M1 | 2(13.3) | 13(86.7) | 3(20.0) | 12(80.0) | ||
| CEA level (μg/L) | 0.319 | 0.912 | ||||
| ≤5 | 61(49.2) | 63(50.8) | 51(41.1) | 73(58.1) | ||
| >5 | 10(38.5) | 16(61.5) | 11(42.3) | 15(57.7) | ||
| LVI | 0.011 | 0.009 | ||||
| Yes | 12(32.4) | 25(67.6) | 9(24.3) | 28(75.7) | ||
| No | 57(57.0) | 43(43.0) | 49(24.3) | 51(75.7) | ||
| PNI | 0.949 | 0.917 | ||||
| Yes | 8(50.0) | 8(50.0) | 7(43.8) | 9(56.3) | ||
| No | 60(50.8) | 58(49.2) | 50(42.4) | 68(57.6) | ||
Bold indicates statistically significant value.
Univariate and multivariate analyses for OS in GC patients
| Variable | Univariate | Multivariate | ||||
|---|---|---|---|---|---|---|
| HR | 95%CI | P | HR | 95%CI | P | |
| Age (years) | ||||||
| ≥60 vs. <60 | 0.868 | 0.553-1.363 | 0.539 | |||
| Gender | ||||||
| Male vs. Female | 0.731 | 0.464-1.152 | 0.178 | |||
| Tumor size | ||||||
| >5 cm vs. ≤5 cm | 3.201 | 1.995-5.137 | <0.001 | 2.098 | 1.201-3.664 | 0.009 |
| Borrmann type | ||||||
| III-IV vs. I-II | 5.506 | 1.992-15.219 | 0.001 | 2.412 | 0.825-7.050 | 0.108 |
| Differentiation | ||||||
| Poor vs. Well+ moderate | 1.717 | 0.908-3.246 | 0.096 | |||
| Depth of invasion | ||||||
| T3-4 vs. T1-2 | 15.71 | 3.851-64.095 | <0.001 | |||
| Lymph node metastasis | ||||||
| N+ vs. N0 | 7.871 | 3.609-17.169 | <0.001 | 2.965 | 1.183-7.436 | 0.020 |
| Distant metastasis | ||||||
| M1 vs. M0 | 3.011 | 1.644-5.514 | <0.001 | |||
| CEA level (μg/L) | ||||||
| >5 vs. ≤5 | 1.787 | 1.059-3.015 | 0.03 | |||
| LVI | ||||||
| Present vs. none | 2.980 | 1.816-4.888 | <0.001 | |||
| PNI | ||||||
| Present vs. none | 1.388 | 0.684-2.817 | 0.364 | |||
| GALNT1 | ||||||
| High vs. Low | 4.406 | 2.574-7.541 | <0.001 | 2.458 | 1.297-4.660 | 0.006 |
Bold indicates statistically significant value.
Univariate and multivariate analyses for DFS in GC patients
| Variable | Univariate | Multivariate | ||||
|---|---|---|---|---|---|---|
| HR | 95%CI | P | HR | 95%CI | P | |
| Age (years) | ||||||
| ≥60 vs. <60 | 0.924 | 0.571-1.493 | 0.746 | |||
| Gender | ||||||
| Male vs. Female | 0.742 | 0.456-1.207 | 0.229 | |||
| Tumor size | ||||||
| >5 cm vs. ≤5 cm | 3.716 | 2.250-6.137 | <0.001 | 2.057 | 1.155-3.664 | 0.014 |
| Borrmann type | ||||||
| III-IV vs. I-II | 5.645 | 2.043-15.602 | 0.001 | 2.572 | 0.885-7.475 | 0.083 |
| Differentiation | ||||||
| Poor vs. Well+ moderate | 1.731 | 0.916-3.272 | 0.091 | |||
| Depth of invasion | ||||||
| T3-4 vs. T1-2 | 29.55 | 4.096-213.16 | 0.001 | |||
| Lymph node metastasis | ||||||
| N+ vs. N0 | 7.604 | 3.466-16.683 | <0.001 | 2.817 | 1.116-7.114 | 0.028 |
| Distant metastasis | ||||||
| M1 vs. M0 | 3.287 | 1.184-9.124 | 0.022 | |||
| CEA level (μg/L) | ||||||
| >5 vs. ≤5 | 1.872 | 1.064-3.296 | 0.030 | |||
| LVI | ||||||
| Present vs. none | 3.217 | 1.923-5.380 | <0.001 | 1.680 | 0.943-2.994 | 0.079 |
| PNI | ||||||
| Present vs. none | 1.093 | 0.469-2.546 | 0.836 | |||
| GALNT1 | ||||||
| High vs. Low | 4.259 | 2.434-7.454 | <0.001 | 2.542 | 1.335-4.841 | 0.005 |
Bold indicates statistically significant value.
GALNT1 regulates proliferation, migration, and invasion of GC cells in vitro
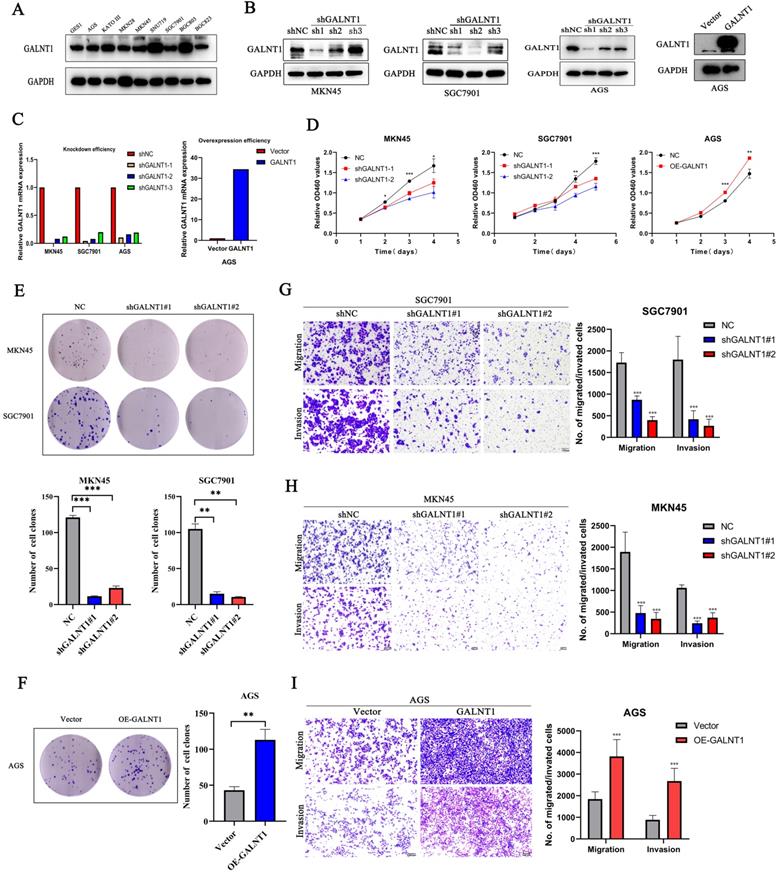
To better understand the effect of GALNT1 in GC, we investigated phenotypic changes in GC cells after GALNT1 knockdown or overexpression. We initially assessed the GALNT1 levels in several cell lines using western blot assays (Fig. 3A). Then, GALNT1 was stably knocked down in MKN45, SGC7901, and AGS cells, and overexpressed in AGS cells. The effectiveness of GALNT1 knockdown and overexpression was examined by western blotting and qRT-PCR assays (Fig. 3B-C). Cell counting kit-8 (CCK8) assays revealed that GALNT1 knockdown significantly impaired the growth of MKN45, SGC7901, and AGS cells, whereas GALNT1 overexpression in AGS cells significantly increased cell proliferation (Fig. 3D, Fig. S2A). Similarly, colony formation assays further indicated that the clonogenic capacity of GC cells was suppressed after GALNT1 deficiency, whereas overexpression of GALNT1 in AGS cells yielded the opposite result (Fig. 3E-F, Fig. S2B). In addition, cell migration and invasion assays demonstrated that GALNT1 knockdown significantly decreased the migration and invasion abilities of GC cells (Fig. 3G-H, Fig. S2C). Furthermore, GALNT1 overexpression facilitated migration and invasion of AGS cells (Fig. 3I). Overall, these results suggest that GALNT1 plays a crucial role in the proliferation, migration, and invasion of GC cells.
GALNT1 promotes the growth and metastatic ability of gastric cancer cells in vitro. A, GALNT1 expression in gastric cancer cell lines as examined by western blot analysis. B-C, Western blot (B) and qPCR (C) analysis depicting the silencing efficiency of GALNT1 in MKN45, AGS, and SGC7901 cells as well as the overexpression of GALNT1 in AGS cells. D-F, Cells growth ability after GALNT1 knockdown and overexpression were determined by CCK8 assay (D) and colony formation assay (E, F). G-I, Knockdown of GALNT1 decreased the abilities of migration and invasion of MKN45 and SGC7901 cells (G, H). Overexpression of GALNT1 enhanced the abilities of migration and invasion of AGS cells (I). Data are shown as means ± S.D. *p<0.05, **p<0.01, ***p<0.001.
GALNT1 knockdown inhibits growth and metastasis of GC cells in vivo
Subcutaneous tumor in vivo experiment was performed to investigate the function of GALNT1 in driving tumorigenesis. Compared to the NC group, the shGALNT1 group had a much smaller mean tumor volume and tumor weight (Fig. 4A-C), manifesting that targeted knockdown of GALNT1 expression effectively inhibited tumor proliferation in vivo.
GALNT1 regulates subcutaneous tumor formation and metastasis of gastric cancer in mice. A, GALNT1 knockdown inhibited subcutaneous tumor formation in a nude mouse model. B-C, Tumor weight (B) and volume (C) analysis of subcutaneous tumors in nude mouse models. D, SGC7901 cells with control or GALNT1 knockdown were injected into the tail veins of nude mice (n = 7 for each group). Representative images of lung taken seven weeks after injection. E, Quantification of metastatic lung tumors by hematoxylin & eosin. F, Representative images of metastatic lung tumors performed by hematoxylin & eosin staining and immunohistochemistry in mice. Data are shown as means ± S.D. *p<0.05, **p<0.01, ***p<0.001.
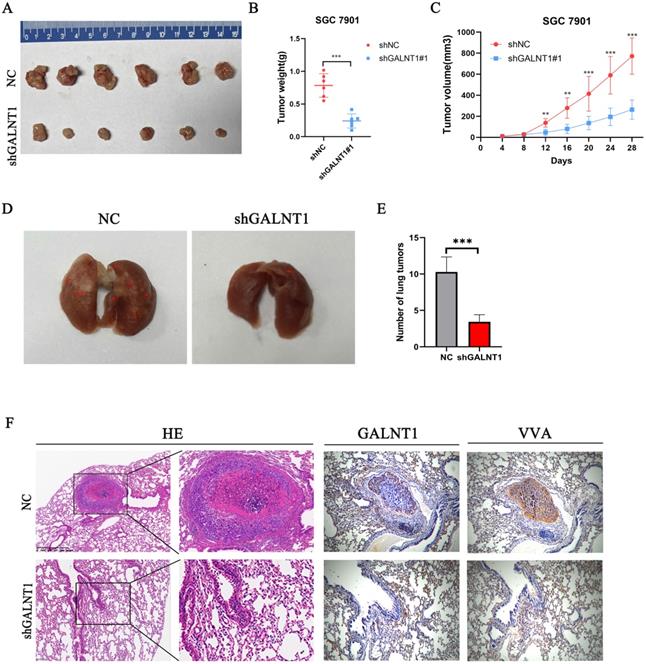
Metastasis is an important risk factor for a poor prognosis. As mentioned above, upregulated GALNT1 is associated with lymph node metastasis and distant metastasis, and GALNT1 can enhance the migration and invasion of GC cells in vitro. To further evaluate the metastatic ability of GC cells in vivo, we directly injected GALNT1-deficient SGC7901 cells and control cells into the tail veins of mice to examine the lung metastasis capacity of GC cells. Representative images of the lungs taken seven weeks after injection are shown in Fig. 4D. Mice implanted with GALNT1-deficient SGC7901 cells had fewer lung metastatic nodules than those in the control group, as shown by hematoxylin and eosin (H&E) staining (Fig. 4E). Furthermore, representative images showed that the expression levels of GALNT1 and VVA were upregulated in the metastatic lung nodules following immunohistochemistry (Fig. 4F). These results indicate the vital role of GALNT1 in enhancing the invasion and metastasis of GC.
GALNT1 activates the Wnt/β-catenin signaling in GC cells
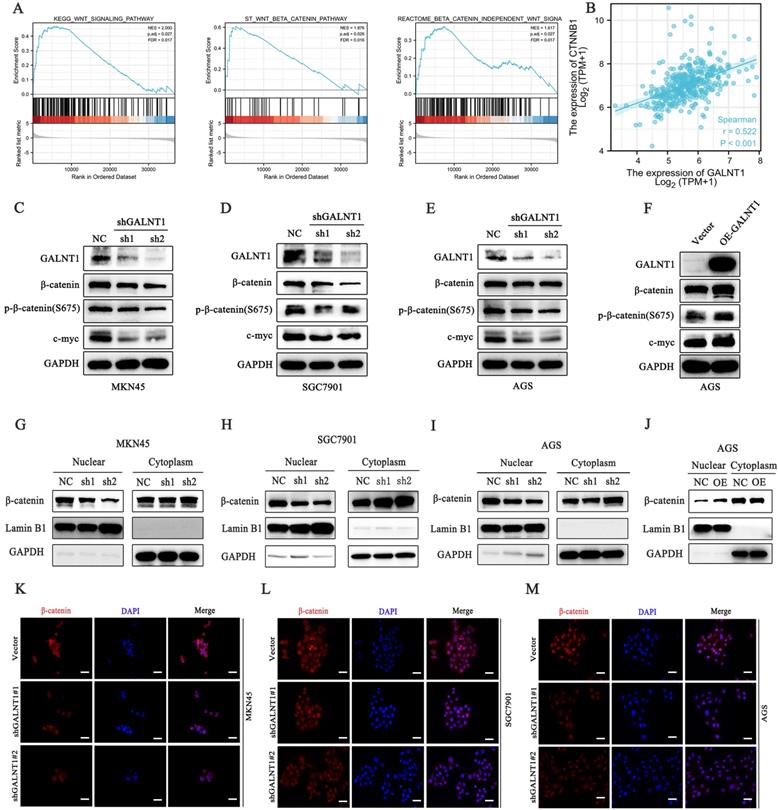
To investigate the underlying mechanisms of GALNT1 in GC development, we first performed gene set enrichment analysis (GSEA) analysis of TCGA data, which indicated that GALNT1 was associated with the Wnt/β-catenin signaling pathway in GC (Fig. 5A). The aberrant Wnt/β-catenin signaling pathway is tightly linked to cell proliferation, invasion, and metastasis in multiple malignancies [21]. To confirm the linkage between GALNT1 and Wnt/β-catenin signaling, correlation analysis was conducted on TCGA data and revealed that GALNT1 expression was positively associated with β-catenin expression in GC (r=0.522, p<0.001; Fig. 5B). Therefore, to further confirm whether Wnt/β-catenin is engaged in GALNT1-mediated pathways, the levels of key molecules in the pathway were examined by western blotting. We first silenced GALNT1 expression using shRNA in multiple GC cell lines, including MKN45, SGC7901, and AGS, and discovered that the Wnt/β-catenin signaling-related molecules β-catenin and phospho-β-catenin (S675) had considerably lower levels, as did c-Myc, a downstream target (Fig. 5C-E). In contrast, GALNT1 overexpression significantly increased the levels of these proteins (Fig. 5F). To investigate this further, nuclear/cytoplasmic separation was performed to detect β-catenin levels in the cytoplasm and nucleus. The expression levels of β-catenin in the nucleus were downregulated in GALNT1-knockdown cell lines of GC (Fig. 5G-I), whereas overexpression of GALNT1 increased β-catenin nuclear expression (Fig. 5J). Moreover, immunofluorescence assays indicated that GALNT1 deficiency reduced, whereas GALNT1 overexpression upregulated the expression and nuclear signals of β-catenin (Fig. 5K-M). The luciferase reporter assay indicated that GALNT1 overexpression significantly increased the activity of β-catenin-driven transcription and vice versa (Fig. S3A-B). Collectively, these findings suggest that the Wnt/β-catenin signaling pathway is regulated by GALNT1 overexpression.
GALNT1 positively regulates the β-catenin signaling pathway in gastric cancer cells. A, GSEA plots showing the Wnt/β-catenin pathways of differentially expressed genes altered by GALNT1 in patients with gastric cancer from the TCGA database. B, Correlation of GALNT1 and β-catenin in gastric cancer from TCGA database were analyzed. C-F, Western blotting analysis-based detection of protein expression of β-catenin, phospho-β-catenin (S675) and its downstream target c-Myc in MKN45 (C), SGC7901 (D) and AGS (E) cells after knockdown of GALNT1, as well as in the AGS cells after overexpression of GALNT1 (F). G-J, Protein expression of β-catenin in the cytoplasm and nucleus of MKN45 (G), SGC7901 (H), and AGS (I) cells after knockdown of GALNT1 as determined by western blot analysis, as well as AGS cells after overexpression of GALNT1 (J). K-M, Nuclear translocation of β-catenin examined by immunofluorescence assays in GALNT1-silenced MKN45 (K), SGC7901 (L), and AGS (M) cells. Scale bar: 50μm.
GALNT1 mediates β-catenin signaling through O-glycosylation of CD44
As GALNT1 is an O-glycosyltransferase, O-glycosylation of proteins may be modified to mediate the Wnt/β-catenin signaling pathway in GC cells. A previous study has revealed that CD44 is a specific substrate for GALNT1 [22]. Therefore, we used NetOGlyc-4.0 to predict multiple O-glycosylation sites in CD44. Furthermore, we investigated the primary role of GALNT1 in CD44 modified O-glycans in GC cells using VVA lectins, which recognize tumor-associated Tn antigens. To confirm whether GALNT1 can modify O-glycosylation on CD44, cell lysates were immunoprecipitated (IP) for CD44 in GC cells with GALNT1 silencing or overexpression, and western blotting with biotin-conjugated VVA was used to differentiate the levels of Tn antigen. These results show that GALNT1 silencing reduced VVA binding to CD44 in MKN45, SGC7901, and AGS cells (Fig. 6A-B; Fig. S4A). Conversely, GALNT1 overexpression increased the binding of VVA to CD44 in AGS cells (Fig. 6C). To further verify these results, lectin pull-down of glycoproteins using biotinylated VVA and streptavidin agarose were immunoblotted (IB) with an anti-CD44 antibody. Consistently, western blot analysis indicated that GALNT1 silencing downregulated (Fig. 6D-E; Fig. S4B), whereas overexpression of GALNT1 enhanced the expression of CD44 in glycoproteins pulled down by VVA lectin (Fig. 6F). This finding suggests that GALNT1 plays a crucial role in modifying O-glycans on CD44 in GC cells.
GALNT1 positively regulates Wnt/β-catenin signaling pathway through O-glycosylation of CD44. A-C, GALNT1 modification of O-glycosylation of CD44 measured by co-immunoprecipitation in gastric cancer cells. Cell lysates were immunoprecipitated (IP) for CD44, and then western blotted with biotin-conjugated VVA to distinguish Tn antigen expression. D-F, Lectin pull down glycoproteins using biotinylated VVA, and streptavidin agarose immunoblotted with anti-CD44 antibody. G, Western blot and qPCR analysis for the silencing efficiency of CD44 in AGS cells. H, The growth abilities of AGS cells after GALNT1 overexpression and CD44 knockdown were determined by colony formation assays. I, The migration and invasion abilities of AGS cells transfected with OE-GALNT1 and shCD44 measured by transwell assays. Representative images of the migrated or invaded cells and statistical graphs are shown. J-K, Immunofluorescence assay (J), and western blot analysis (K) depicting the nuclear translocation of β-catenin in AGS cells transfected with OE-GALNT1 and shCD44. Scale bar: 50 µm. Data are shown as means ± S.D. *P<0.05, **P<0.01, ***P<0.001.
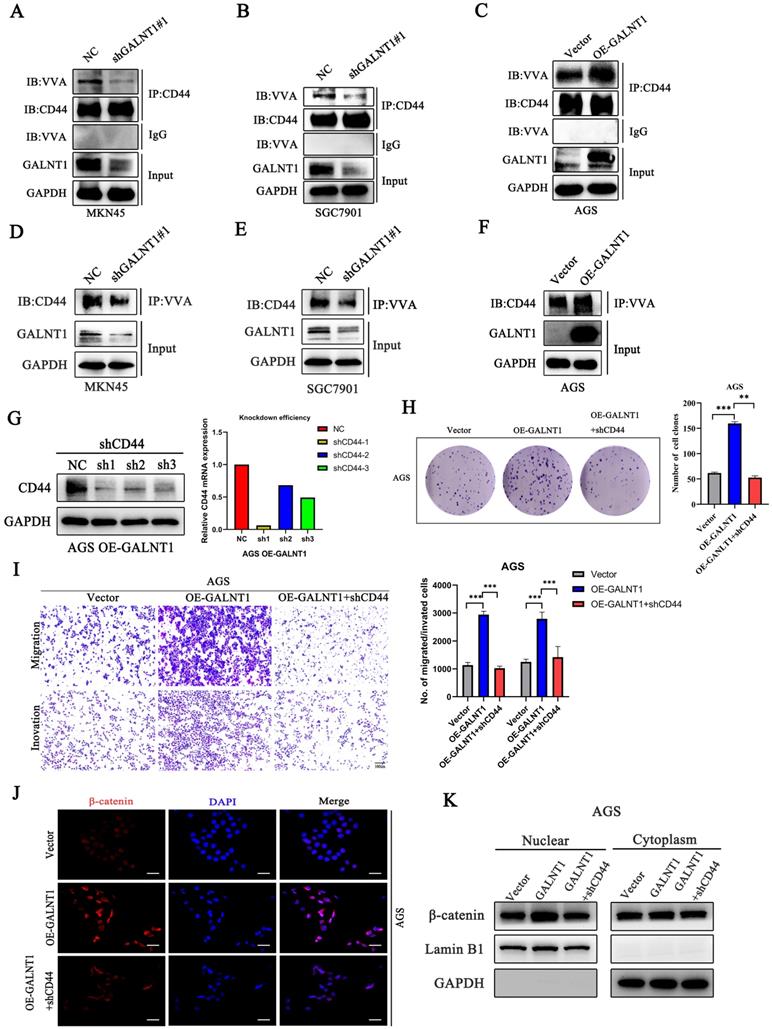
To investigate the mechanism by which GALNT1 regulates the Wnt/β-catenin signaling pathway through O-glycosylation of CD44, we conducted a rescue assay in AGS cells overexpressing GALNT1 and the corresponding control cells. We first stably knocked down CD44 in AGS cells overexpressing GALNT1, and then western blot and qPCR analyses were utilized to investigate the efficiency of knockdown (Fig. 6G). As shown in Fig. 6H, clonogenic capacity was reversed by CD44 knockdown in AGS cells overexpressing GALNT1. Consistently, CD44 silencing in AGS cells with GALNT1 overexpression reversed the favorable effects of GALNT1 on migration and invasion (Fig. 6I). Moreover, immunofluorescence assays revealed that CD44 knockdown abolished the favorable effects of GALNT1 on the nuclear signals of β-catenin in AGS cells (Fig. 6J). Meanwhile, nuclear/cytoplasmic separation analysis also revealed that nuclear β-catenin signals were reversed by CD44 knockdown in AGS cells overexpressing GALNT1 (Fig. 6K). Collectively, these findings suggest that GALNT1 mediates the β-catenin signaling pathway through the O-glycosylation of CD44.
GALNT1/Tn antigen is relocated into the ER
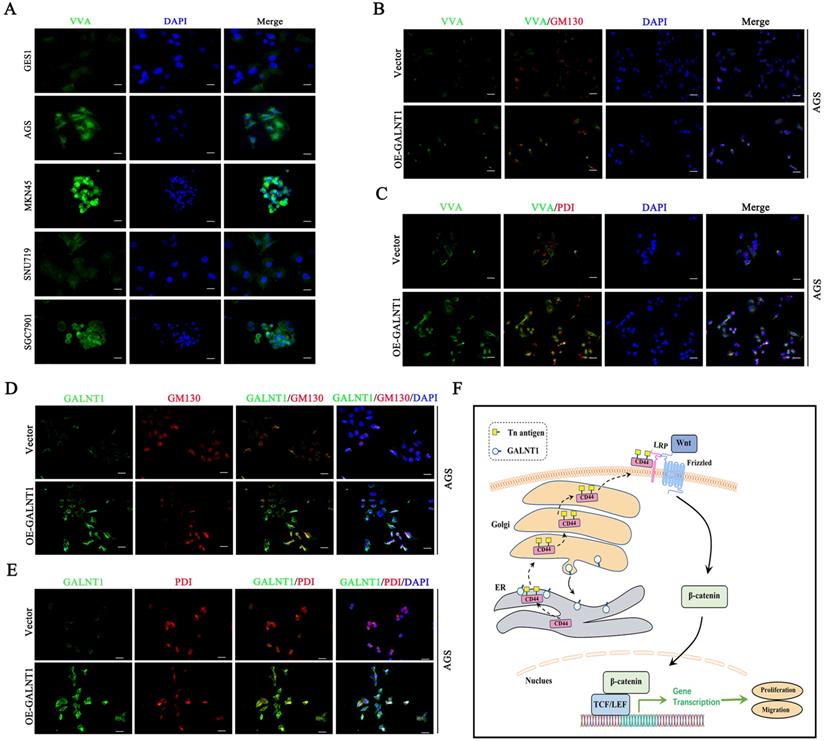
Interestingly, immunofluorescence analysis of multiple GC cells showed that Tn antigen expression was upregulated in the aggressive GC cells MKN45 and SGC7901, whereas it was decreased in less-invasive AGS and SNU719 cells (Fig. 7A). This was similar to our western blot findings of GALNT1 expression levels in GC cells. Tn antigen expression is associated with the initiation of O-glycosylation, and the Golgi apparatus is the primary site of O-glycosylation [16]. Recent studies have found that O-glycosylation initiated by GALNT1 is relocated from the Golgi apparatus to the endoplasmic reticulum (ER) during tumor progression and metastasis (GALA pathway) [8, 17, 18]. Therefore, to verify the mechanism, we first investigated the interaction between the Tn antigen (VVA, green) and Golgi-marker GM130 (red) or ER-marker PDI in AGS cells overexpressing GALNT1. Strikingly, we found that abundant Tn antigen was located in the ER (Fig. 7B-C). In addition, immunofluorescence analysis of GALNT1 with GM130 or PDI also indicated that GALNT1 was located in the Golgi as well as obviously ER in AGS cells overexpressing GALNT1 (Fig. 7D-E). Collectively, the initiation of O-glycosylation is associated with the relocation of GALNT1 to the ER during tumor progression and metastasis.
GALNT1/Tn translocates to the ER. A, Immunofluorescence of Tn antigen in multiple gastric cancer cells. B, Co-staining Tn antigen (VVA, green) and GM130 (red) in the indicated cells. C, Co-staining Tn antigen (VVA, green) and PDI (red) in the indicated cells. D, Colocalization of GALNT1 (green) and GM130 (red) in AGS cells after overexpression of GALNT1. E, Colocalization of GALNT1 (green) and PDI (red) in AGS cells after overexpression of GALNT1. Scale bar: 50μm. F, Schematic illustration of the working hypothesis.
Discussion
Previous studies have indicated that GALNT1 is upregulated in multiple malignancies, including hepatocellular carcinoma, breast cancer, and bladder cancer, and is strongly associated with tumor progression [17, 18, 23]. However, the function and detailed molecular mechanism of GALNT1 in GC remain largely unknown. In the present study, we confirmed that GALNT1 expression is commonly upregulated in GC and that it is closely associated with the pTNM stage, depth of invasion, metastasis, and poor survival. Moreover, we revealed its involvement in maintaining malignancy and facilitation of metastasis. Mechanistic investigations suggest that overexpression of GALNT1 in GC cells modifies the O-glycosylation of CD44, stimulating the Wnt/β-catenin signaling pathway to promote the progression and metastasis of GC. The main conclusions of this study are shown in Fig. 7F. This is the first study to determine that GALNT1 plays a crucial role in GC progression.
Although significant advances have been made in detecting and treating GC, metastasis remains the primary reason for the high mortality of patients with GC [2]. Hence, it is essential to understand the detailed molecular mechanisms of metastasis and develop more effective and targeted reagents for the treatment of GC. Bioinformatics analysis showed that GALNT1 is overexpressed in GC as well as other malignant tumors, including breast cancer, bladder cancer, and hepatocellular carcinoma. Our findings were consistent with the previously reported data [17, 18, 23]. These results suggest that GALNT1 may be a crucial oncogene during tumor evolution. Moreover, we further confirmed that GALNT1 is frequently upregulated in GC and promotes abnormal O-glycosylation, which is associated with poor survival, by analyzing the clinicopathological features of GC patients. Hence, to further confirm the effects of GALNT1 on gastric carcinogenesis and progression, the effects of upregulating and downregulating GALNT1 on cell function were examined. We further demonstrated that GALNT1 expression affects the malignant and metastatic phenotypes of GC cells in vivo and in vitro. These findings suggest that GALNT1 may be a valuable biomarker and candidate target for new GC treatments.
In numerous cancers, the Wnt/β-catenin signaling pathway is often inappropriately activated [24, 25]. This pathway plays a vital role in embryonic development, and regulates the development, regulation, and survival of cancer cells [26, 27]. In GC, the Wnt/β-catenin signaling pathway contributes to cancer development and is implicated in recurrence and poor survival [28-30]. Consistently, in the present study, we found that GALNT1 knockdown reduced the protein expression of β-catenin, p-β-catenin (S675), and its downstream target c-Myc in GC cells, whereas GALNT1 overexpression increased the expression of these proteins. Moreover, we revealed that silencing GALNT1 inhibited, whereas GALNT1 overexpression promoted β-catenin translocation into the nucleus in GC cells. Collectively, these results indicate that GALNT1 positively regulates the Wnt/β-catenin signaling pathway in GC cells. Therefore, understanding the detailed regulatory mechanisms involved is essential.
GALNT1 plays a crucial role in the biosynthesis of O-glycans such as the Tn antigen, which is identified by VVA lectin. In breast and liver cancer, previous studies have indicated that higher Tn antigen levels were observed in cancerous tissues than in healthy tissues and were positively linked to progressive tumors and poor survival [17, 18]. These results are consistent with our finding that Tn antigen levels are increased in GC tissues and confer a poorer prognosis. Cancer vaccines based on short O-glycans have been developed as novel cancer therapies [31]. Glycoproteins modified with short O-glycans frequently affect the function of cancer cells and their antigenic properties, thereby promoting the development of invasion and metastasis capabilities [32]. Moreover, new discoveries have revealed that GALNTs relocate from the Golgi apparatus to the ER to initiate O-glycosylation [8, 17, 18, 33]. We also observed this translocation of GALNT1 to the ER. However, the role of short O-glycans in GC progression remains unclear. Nonetheless, our study revealed that GALNT1 is the main GalNAc transferase in GC and that it promotes abnormal O-glycosylation.
In the present study, we found that GALNT1 could modify CD44 O-glycosylation in GC cells. Consistently, a previous study revealed that CD44 is a specific substrate for GALNT1 [22]. Moreover, CD44 has multiple O-glycosylation sites, as predicted by NetOGlyc-4.0. CD44 transmembrane glycoproteins influence cell growth, survival, differentiation, and motility, and are correlated with malignant tumors [34, 35]. Many studies have reported that CD44 is aberrantly expressed in several human tumors, and it plays a crucial role in cancer metastasis and tumor growth [35-40]. In GC, CD44 has also been identified as an oncogene with high malignant potential, which can promote metastasis [41, 42]. In addition, recent research has shown that CD44 acts as a positive regulator of the Wnt/β-catenin signaling pathway [43]. Therefore, we conducted a rescue assay in AGS cells overexpressing GALNT1 and the corresponding control cells to investigate the mechanisms by which the GALNT1-mediated O-glycosylation of CD44 regulates the Wnt/β-catenin signaling pathway. We found that CD44 silencing in AGS cells with GALNT1 overexpression reversed the positive effects of GALNT1 on the migration, invasion, and nuclear signals of β-catenin. To our knowledge, this is the first study to report that GALNT1 regulates the Wnt/β-catenin signaling pathway through the O-glycosylation of CD44 in GC cells. Further validation of the specific O-glycosylation sites in CD44 would help understand the regulatory mechanism of the Wnt/β-catenin signaling pathway.
Conclusion
GALNT1 is upregulated in GC and promotes abnormal O-glycosylation of CD44, thereby activating the Wnt/β-catenin signaling pathway and regulating the malignant behavior of GC cells. This study provides insights into the pathophysiological role of GALNT1 and the significance of abnormal O-glycosylation in GC tumor progression. These findings may pave the way for developing novel therapeutic agents for GC.
Abbreviations
GALNT1: N-acetylgalactosamine-transferase 1; GC: gastric cancer; VVA: vicia villosa; GalNAc: N-acetylgalactosamine; Ser: serine; Thr: threonine; ER: endoplasmic reticulum; GALA: GALNTs activation; MUC1: mucin 1; OPN: osteopontin; MMP14: matrix metalloproteinase-14; TCGA: The Cancer Genome Atlas; ROC: receiver-operating characteristic; AUC: area under the curve; pTNM: pathological tumor-node-metastasis; LVI: lymphovascular invasion; OS: overall survival; GEPIA: Gene Expression Profiling Interactive Analysis; DFS: disease-free survival; PBS: phosphate-buffered saline; PVDF: polyvinylidene fluoride; NC: nitrocellulose; DAB: 3,3'-diaminobenzidine; H&E: hematoxylin and eosin; GSEA: gene set enrichment analysis; IB: immunoblot; shGALNT1: GALNT1 knockdown; shCD44: CD44 knockdown; qRT-PCR: quantitative real-time PCR; IHC: immunohistochemistry; HRP: horseradish peroxidase; CCK-8: cell counting kit-8.
Supplementary Material
Supplementary figures and tables.
Acknowledgements
Funding
This work was supported by grants from National Natural Science Foundation of China (81772579), Guangdong Province Science and Technology Plan Projects (2014A020212693), “3&3” Project of The First Affiliated Hospital of Sun Yat-Sen University (Yulong He) and Sanming Project of Medical in Shenzhen (SZSM201911010). All these study sponsors have no roles in the study design, in the collection, analysis and interpretation of data.
Ethics approval
This study was approved by Ethics Committee of the First and Seventh Affiliated Hospital of Sun Yat-sen University. All patients signed the informed consent form.
Author Contributions
Y.H., C.Z. and M.H. designed and supervised the study. J.Z. and H.W. for implementation of the experiment and collection of specimens, analysis of data and drafting the paper. J.W., C.Y., S.C. and S.L. for carrying out the statistical analysis and provided critical comments. Y.H., C.Z. and M.H. checked and improved the manuscript. All authors read and approved the final manuscript.
Data Availability
The data in this study are available with the approval of corresponding authors in reasonable requirement.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A. et al. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J Clin. 2021;71:209-49
2. Luebeck EG, Curtius K, Jeon J, Hazelton WD. Impact of tumor progression on cancer incidence curves. Cancer Res. 2013;73:1086-96
3. Pinho SS, Reis CA. Glycosylation in cancer: mechanisms and clinical implications. Nat Rev Cancer. 2015;15:540-55
4. Fuster MM, Esko JD. The sweet and sour of cancer: glycans as novel therapeutic targets. Nat Rev Cancer. 2005;5:526-42
5. Bard F, Chia J. Cracking the Glycome Encoder: Signaling, Trafficking, and Glycosylation. Trends Cell Biol. 2016;26:379-88
6. Ju T, Otto VI, Cummings RD. The Tn antigen-structural simplicity and biological complexity. Angew Chem Int Ed Engl. 2011;50:1770-91
7. Duarte HO, Freitas D, Gomes C, Gomes J, Magalhães A, Reis CA. Mucin-Type O-Glycosylation in Gastric Carcinogenesis. Biomolecules. 2016;6:33
8. Gill DJ, Tham KM, Chia J, Wang SC, Steentoft C, Clausen H. et al. Initiation of GalNAc-type O-glycosylation in the endoplasmic reticulum promotes cancer cell invasiveness. Proc Natl Acad Sci U S A. 2013;110:E3152-61
9. Springer GF. T and Tn, general carcinoma autoantigens. Science. 1984;224:1198-206
10. Chia J, Tham KM, Gill DJ, Bard-Chapeau EA, Bard FA. ERK8 is a negative regulator of O-GalNAc glycosylation and cell migration. Elife. 2014;3:e01828
11. Bennett EP, Mandel U, Clausen H, Gerken TA, Fritz TA, Tabak LA. Control of mucin-type O-glycosylation: a classification of the polypeptide GalNAc-transferase gene family. Glycobiology. 2012;22:736-56
12. Liu Y, Liu H, Yang L, Wu Q, Liu W, Fu Q. et al. Loss of N-Acetylgalactosaminyltransferase-4 Orchestrates Oncogenic MicroRNA-9 in Hepatocellular Carcinoma. J Biol Chem. 2017;292:3186-200
13. Taniuchi K, Cerny RL, Tanouchi A, Kohno K, Kotani N, Honke K. et al. Overexpression of GalNAc-transferase GalNAc-T3 promotes pancreatic cancer cell growth. Oncogene. 2011;30:4843-54
14. Li Z, Yamada S, Inenaga S, Imamura T, Wu Y, Wang KY. et al. Polypeptide N-acetylgalactosaminyltransferase 6 expression in pancreatic cancer is an independent prognostic factor indicating better overall survival. Br J Cancer. 2011;104:1882-9
15. Wu C, Guo X, Wang W, Wang Y, Shan Y, Zhang B. et al. N-Acetylgalactosaminyltransferase-14 as a potential biomarker for breast cancer by immunohistochemistry. BMC Cancer. 2010;10:123
16. Röttger S, White J, Wandall HH, Olivo JC, Stark A, Bennett EP. et al. Localization of three human polypeptide GalNAc-transferases in HeLa cells suggests initiation of O-linked glycosylation throughout the Golgi apparatus. J Cell Sci. 1998;111( Pt 1):45-60
17. Fang R, Xu F, Shi H, Wu Y, Cao C, Li H. et al. LAMTOR5 raises abnormal initiation of O-glycosylation in breast cancer metastasis via modulating GALNT1 activity. Oncogene. 2020;39:2290-304
18. Nguyen AT, Chia J, Ros M, Hui KM, Saltel F, Bard F. Organelle Specific O-Glycosylation Drives MMP14 Activation, Tumor Growth, and Metastasis. Cancer Cell. 2017;32:639-53.e6
19. Gerken TA, Raman J, Fritz TA, Jamison O. Identification of common and unique peptide substrate preferences for the UDP-GalNAc:polypeptide alpha-N-acetylgalactosaminyltransferases T1 and T2 derived from oriented random peptide substrates. J Biol Chem. 2006;281:32403-16
20. Li C, Yang Z, Du Y, Tang H, Chen J, Hu D. et al. BCMab1, a monoclonal antibody against aberrantly glycosylated integrin α3β1, has potent antitumor activity of bladder cancer in vivo. Clin Cancer Res. 2014;20:4001-13
21. Lim SK, Lu SY, Kang SA, Tan HJ, Li Z, Adrian Wee ZN. et al. Wnt Signaling Promotes Breast Cancer by Blocking ITCH-Mediated Degradation of YAP/TAZ Transcriptional Coactivator WBP2. Cancer Res. 2016;76:6278-89
22. Narimatsu Y, Joshi HJ, Schjoldager KT, Hintze J, Halim A, Steentoft C. et al. Exploring Regulation of Protein O-Glycosylation in Isogenic Human HEK293 Cells by Differential O-Glycoproteomics. Mol Cell Proteomics. 2019;18:1396-409
23. Li C, Du Y, Yang Z, He L, Wang Y, Hao L. et al. GALNT1-Mediated Glycosylation and Activation of Sonic Hedgehog Signaling Maintains the Self-Renewal and Tumor-Initiating Capacity of Bladder Cancer Stem Cells. Cancer Res. 2016;76:1273-83
24. Pan J, Fang S, Tian H, Zhou C, Zhao X, Tian H. et al. lncRNA JPX/miR-33a-5p/Twist1 axis regulates tumorigenesis and metastasis of lung cancer by activating Wnt/β-catenin signaling. Mol Cancer. 2020;19:9
25. Li Y, Chen F, Shen W, Li B, Xiang R, Qu L. et al. WDR74 induces nuclear β-catenin accumulation and activates Wnt-responsive genes to promote lung cancer growth and metastasis. Cancer Lett. 2020;471:103-15
26. Nusse R, Clevers H. Wnt/β-Catenin Signaling, Disease, and Emerging Therapeutic Modalities. Cell. 2017;169:985-99
27. Clevers H, Nusse R. Wnt/β-catenin signaling and disease. Cell. 2012;149:1192-205
28. Ji L, Qian W, Gui L, Ji Z, Yin P, Lin GN. et al. Blockade of β-Catenin-Induced CCL28 Suppresses Gastric Cancer Progression via Inhibition of Treg Cell Infiltration. Cancer Res. 2020;80:2004-16
29. Wang JB, Wang ZW, Li Y, Huang CQ, Zheng CH, Li P. et al. CDK5RAP3 acts as a tumor suppressor in gastric cancer through inhibition of β-catenin signaling. Cancer Lett. 2017;385:188-97
30. Ganesan K, Ivanova T, Wu Y, Rajasegaran V, Wu J, Lee MH. et al. Inhibition of gastric cancer invasion and metastasis by PLA2G2A, a novel beta-catenin/TCF target gene. Cancer Res. 2008;68:4277-86
31. Astronomo RD, Burton DR. Carbohydrate vaccines: developing sweet solutions to sticky situations? Nat Rev Drug Discov. 2010;9:308-24
32. Brockhausen I. Mucin-type O-glycans in human colon and breast cancer: glycodynamics and functions. EMBO Rep. 2006;7:599-604
33. Gill DJ, Clausen H, Bard F. Location, location, location: new insights into O-GalNAc protein glycosylation. Trends Cell Biol. 2011;21:149-58
34. Zöller M. CD44: can a cancer-initiating cell profit from an abundantly expressed molecule? Nat Rev Cancer. 2011;11:254-67
35. Ponta H, Sherman L, Herrlich PA. CD44: from adhesion molecules to signalling regulators. Nat Rev Mol Cell Biol. 2003;4:33-45
36. Jordan AR, Racine RR, Hennig MJ, Lokeshwar VB. The Role of CD44 in Disease Pathophysiology and Targeted Treatment. Front Immunol. 2015;6:182
37. Kim H, Yang XL, Rosada C, Hamilton SR, August JT. CD44 expression in colorectal adenomas is an early event occurring prior to K-ras and p53 gene mutation. Arch Biochem Biophys. 1994;310:504-7
38. Wielenga VJ, Heider KH, Offerhaus GJ, Adolf GR, van den Berg FM, Ponta H. et al. Expression of CD44 variant proteins in human colorectal cancer is related to tumor progression. Cancer Res. 1993;53:4754-6
39. Szatanek R, Baj-Krzyworzeka M. CD44 and Tumor-Derived Extracellular Vesicles (TEVs). Possible Gateway to Cancer Metastasis. Int J Mol Sci. 2021;22:1463
40. Ouhtit A, Rizeq B, Saleh HA, Rahman MM, Zayed H. Novel CD44-downstream signaling pathways mediating breast tumor invasion. Int J Biol Sci. 2018;14:1782-90
41. Nguyen PH, Giraud J, Chambonnier L, Dubus P, Wittkop L, Belleannée G. et al. Characterization of Biomarkers of Tumorigenic and Chemoresistant Cancer Stem Cells in Human Gastric Carcinoma. Clin Cancer Res. 2017;23:1586-97
42. Chen T, Yang K, Yu J, Meng W, Yuan D, Bi F. et al. Identification and expansion of cancer stem cells in tumor tissues and peripheral blood derived from gastric adenocarcinoma patients. Cell Res. 2012;22:248-58
43. Schmitt M, Metzger M, Gradl D, Davidson G, Orian-Rousseau V. CD44 functions in Wnt signaling by regulating LRP6 localization and activation. Cell Death Differ. 2015;22:677-89
Author contact
Corresponding authors: Yulong He, E-mail: hyl1821com; Changhua Zhang, E-mail: zch2108com; Mingyu Huo, E-mail: huomingyu0102com.

 Global reach, higher impact
Global reach, higher impact