ISSN: 1449-2288
Int J Biol Sci 2023; 19(14):4476-4492. doi:10.7150/ijbs.87635 This issue Cite
Research Paper
CD36-BATF2\MYB Axis Predicts Anti-PD-1 Immunotherapy Response in Gastric Cancer
1. Department of Gastroenterology and Hepatology, Shanghai Institute of Liver Diseases, Zhongshan Hospital, Fudan University, Shanghai, 200032, China.
2. Liver Cancer Institute, Zhongshan Hospital, Fudan University, Shanghai, 200032, China.
3. Department of Oncology, Minhang Hospital, Fudan University, China.
4. Key Laboratory of Whole-Period Monitoring and Precise Intervention of Digestive Cancer (SMHC), Minhang Hospital & AHS, Fudan University, China.
5. School of Medicine, Anhui University of Science and Technology, Anhui, 232000, China.
6. Department of Medical Oncology, Zhongshan Hospital, Fudan University, Shanghai 200032, China.
7. NHC Key Laboratory of Glycoconjugate Research, Department of Biochemistry and Molecular Biology, School of Basic Medical Sciences, Fudan University, Shanghai, 200032, China.
8. Shanghai Baoshan District Wusong Central Hospital (Zhongshan Hospital Wusong Branch, Fudan University), Shanghai 200940, China.
†These authors contribute equally to this article.
Abstract
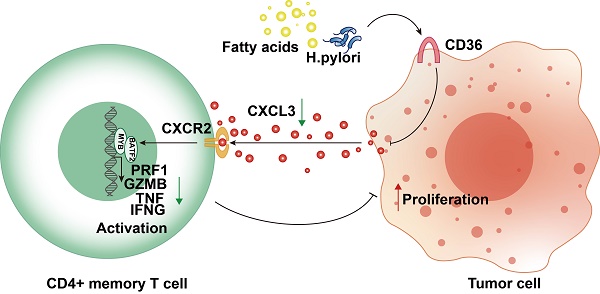
Despite the utilization of anti-PD-1 therapy in gastric cancer (GC), the absence of a reliable predictive biomarker continues to pose a challenge. In this study, we utilized bioinformatic analysis and immunohistochemistry to develop a prediction model for activated CD4+ memory T cells, considering both mRNA and protein levels. An elevation of activated CD4+ memory T cells in GC was noted, which exhibited a strong association with the patients' overall survival. By utilizing WGCNA and DEG analysis, we discovered that BATF2, MYB, and CD36 are genes that exhibit differential expression and are linked to activated CD4+ memory T cells. Afterwards, a forecast model was built utilizing Stepwise regression and immunohistochemistry relying on the three genes. The model's high-risk score showed significant associations with a suppressive immune microenvironment. Moreover, our model exhibited encouraging prognostic value and superior performance in predicting response to immune checkpoint blockade therapy compared with the conventional CD8+PD-L1 model. In terms of mechanism, CD36 could function as a receptor upstream that identifies Helicobacter pylori and fatty acids. This recognition then results in the reduction of the BATF2-MYB protein complex and subsequent alterations in the transcription of genes associated with classical T cell activation. As a result, the activation state of CD4+ memory T cells is ultimately suppressed. The CD36-BATF2/MYB signature serves as a robust predictor of anti-PD-1 immunotherapy response in GC.
Keywords: Gastric cancer, Immune cell related genes, Prognosis, Risk signature, Activated CD4+ memory T cells, Immunotherapy

 Global reach, higher impact
Global reach, higher impact