Impact Factor
ISSN: 1449-2288
Int J Biol Sci 2021; 17(15):4176-4191. doi:10.7150/ijbs.67379 This issue Cite
Research Paper
Cullin3 deficiency shapes tumor microenvironment and promotes cholangiocarcinoma in liver-specific Smad4/Pten mutant mice
1. Cancer Center, Faculty of Health Sciences, University of Macau, Macau SAR, China.
2. Centre for Precision Medicine Research and Training, Faculty of Health Sciences, University of Macau, Macau SAR, China.
3. Department of Oncology, The Affiliated Hospital of Southwest Medical University, Luzhou, China.
4. Zhuhai Interventional Medical Center, Zhuhai Precision Medical Center, Zhuhai People's Hospital, Zhuhai Hospital Affiliated with Jinan University.
5. MOE Frontieers Science Center for Precision Oncogene, University of Macau, Macau SAR, China.
Abstract
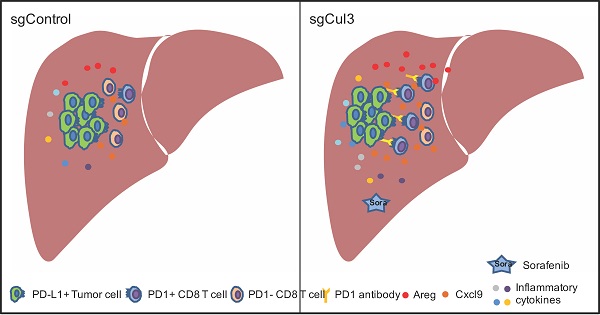
Cholangiocarcinoma (CC), the most lethal type of liver cancer, remains very difficult to treat due to an incomplete understanding of the cancer initiation and progression mechanisms and no effective therapeutic drugs. Thus, identification of genomic drivers and delineation of the underlying mechanisms are urgently needed. Here, we conducted a genome-wide CRISPR-Cas9 screening in liver-specific Smad4/Pten knockout mice (Smad4co/co;Ptenco/co;Alb-Cre, abbreviated as SPC), and identified 15 putative tumor suppressor genes, including Cullin3 (Cul3), whose deficiency increases protein levels of Nrf2 and Cyclin D1 that accelerate cholangiocytes expansion leading to the initiation of CC. Meanwhile, Cul3 deficiency also increases the secretion of Cxcl9 in stromal cells to attract T cells infiltration, and increases the production of Amphiregulin (Areg) mediated by Nrf2, which paracrinely induces inflammation in the liver, and promotes accumulation of exhausted PD1high CD8 T cells at the expenses of their cytotoxic activity, allowing CC progression. We demonstrate that the anti-PD1/PD-L1 blockade inhibits CC growth, and the effect is enhanced by combining with sorafenib selected from organoid mediated drug sensitive test. This model makes it possible to further identify more liver cancer suppressors, study molecular mechanisms, and develop effective therapeutic strategies.
Keywords: Cholangiocarcinoma, tumor microenvironment, inflammatory cytokines, exhausted T cells, anti-PD1/PD-L1 therapy

 Global reach, higher impact
Global reach, higher impact