Impact Factor ISSN: 1449-2288
Int J Biol Sci 2019; 15(1):114-126. doi:10.7150/ijbs.28679 This issue Cite
Research Paper
COPI-Mediated Nuclear Translocation of EGFRvIII Promotes STAT3 Phosphorylation and PKM2 Nuclear Localization
1. Department of Cell Biology, Stem Cell Research Center, School of Basic Medical Sciences, Peking University Health Science Center, Beijing, People's Republic of China
2. Neuroscience Research Institute, School of Basic Medical Sciences, Peking University Health Science Center, Beijing, People's Republic of China
3. Beijing Cellonis Biotechnologies Co. Ltd, Zhongguancun Bio-Medicine Park, Beijing, People's Republic of China
4. Beijing DongFang YaMei Gene Science and Technology Research Institute, Beijing, People's Republic of China
5. Research Center of Artificial Organ and Tissue Engineering, Second Department of Hepatobiliary Surgery, ZhuJiang Hospital, Southern Medical University, Guangzhou, Guangdong Province, People's Republic of China
6. Medical and Health Analytical Center, Peking University Health Science Center, People's Republic of China
# These authors have contributed equally to this work
Received 2018-7-22; Accepted 2018-9-26; Published 2019-1-1
Abstract
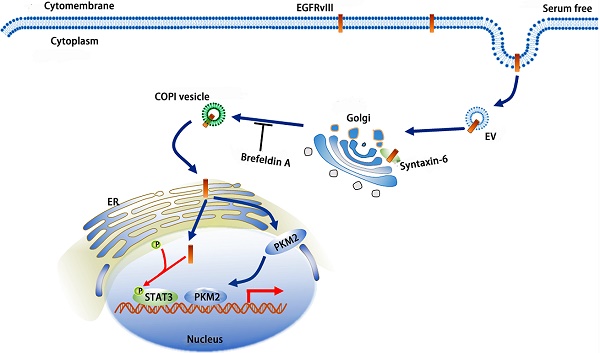
As a non-ligand-dependent activation protein, EGFRvIII is the most common mutant of EGFR, and its existence or especially its nuclear translocation in tumors can exacerbate the malignancy. Compared with the nuclear translocation of EGFR, which has been studied extensively, the specific mechanism by which EGFRvIII undergoes nuclear translocation has not yet been reported. Here, we found that EGFRvIII eventually reached the nucleus with the involvement of the Golgi and endoplasmic reticulum (ER) in glioma cells. In this process, syntaxin-6 was responsible for the identification and transport of EGFRvIII on Golgi. We also demonstrated that COPI mediated the reverse transport of EGFRvIII from the Golgi to ER, which process was also important for EGFRvIII's nuclear accumulation. EGFRvIII's nuclear translocation can significantly promote STAT3 phosphorylation and PKM2 nuclear localization. Finally, we showed that EGFRvIII's nuclear translocation obviously induced the growth of gliomas in an intracranial xenotransplantation experiment. These data suggested that searching methods that inhibit EGFRvIII entry into the nucleus will be effective glioma treatments.
Keywords: EGFRvIII, nuclear translocation, Syntaxin-6, COPI, PKM2 nuclear localization
Introduction
EGFR belongs to the family of receptor tyrosine kinases (RKTs) and can activate downstream signaling cascades in a ligand-dependent manner [1, 2]. Signal pathway disorders caused by the overexpression, amplification and mutation of EGFR are the root cause of many malignant tumors, occurring 50%-70% in glioblastoma multiforme (GBM) cases [3-5].
In EGFR-induced glioma patients, the most common mutation in EGFR is EGFRvIII, which lacks an extracellular domain. Deletion of exons 2-7 in EGFRvIII at the genetic level results in the deletion of extracellular amino acids 6-273 [5-7]. The presence of EGFRvIII is an important marker of more highly malignant gliomas and a worse prognosis in patients, accounting for 25%-33% of all gliomas [8-10].
Based on existing studies, there are two main causes of EGFRvIII-aggravated gliomas: ligand-independent activation of EGFRvIII can continuously activate signaling pathways that are related to proliferation, migration, and invasion of tumor cells[11-13]; and besides, as a cell membrane localization protein, EGFRvIII, which undergoes nuclear localization, exposing its nuclear localization signal (NLS) during endocytosis and then occurring transposition into the nucleus with the aid of some protein molecules [14, 15], where it regulates downstream signals by functioning as a transcription factor. During the nuclear translocation process of EGFRvIII, the most commonly studied cascade is EGFRvIII-STAT3-mediated pro-inflammatory signaling pathway, which is the pathogenesis of many tumors. [3, 15-17].
Thus, the specific process by which EGFRvIII undergoes nuclear translocation must be examined to determine the pathogenesis of glioma and identify targets for its treatment. Unfortunately, there are few reports on the nuclear translocation process of EGFRvIII, but existing studies on the nuclear translocation of EGFR can provide us with a reference [18-20].
Recent studies have shown that many RTK family members exert their unique functions after being transported to different organelles along the secretory pathway [21-23]. After endocytosis, EGFR may be transferred to the Golgi apparatus and ER, known as retrograde trafficking process, and then localizes in the nucleus [18, 19].
As an EGFR mutant, EGFRvIII is likely to undergo a similar process. In addition, retrograde transport between the Golgi and ER is mediated by COPI, which is an important step in nuclear translocation. COPI, consisting of the small GTPase ADP- ribosylation factor (ARF) and seven subunits (α, β, β', γ, δ, ε, and ζ), is coat protein complex I [24]. After GDP-GTP exchange, ARF recruits a coatomer complex that is preassembled by the seven subunits to form COPI that has transport function [25, 26]. Thus, the activation of ARF plays an important role in COPI-mediated protein transport; the inactivation of ARF will cause the disassembly of COPI, suppressing reverse transport. Brefeldin A (BFA), used in this study, is an inhibitor that can inactivate ARF [27]. In this paper, the involvement of the Golgi and ER apparatuses, especially COPI, was studied in the retrograde trafficking of EGFRvIII, and the process by which EGFRvIII undergoes nuclear translocation was determined.
In the alternative splicing of PKM pre-messenger RNA, exon 10 is retained in PKM2, whereas exon 9 is excluded from PKM1, which two share the same pre-messenger RNA [28-30]. PKM2 is an isoenzyme of glycolytic pyruvate kinase, which is highly expressed in many tumors and participates in excess glucose metabolism and lactate production [31-33]. Previous studies have shown that EGFR can promote the nuclear translocation of PKM2 [34, 35], which can further enhance the Warburg effect and possibility of tumorigenesis. Therefore, based on these findings, we investigated the regulation function of EGFRvIII in PKM2 nuclear translocation and prompted us to determine the major role of EGFRvIII nuclear translocation in promoting the nuclear localization of PKM2. This is an overlooked but important discovery, because it has implications for the future treatment of tumors, especially GBM.
Materials and Methods
Tumor samples
Primary tumor samples were obtained in accordance with research ethics board approval from the Department of Pathology, San Bo Brain Hospital, Capital Medical University, Haidian District.
Cell lines, reagents and processing
F98 related cells and C6 rat glioma cell were purchased from the American Type Culture Collection (ATCC) and cultured in Dulbecco's modified Eagle's medium (DMEM) supplemented with 10% fetal bovine serum (HyClone, USA) and 5% CO2 at 37°C. U251 human glioma cell was purchased from Institute of Basic Medical of Science, Chinese Academy of Medical Sciences and cultured in Eagle's Minimal Essential Medium (EMEM) supplemented with 10% fetal bovine serum and 5% CO2 at 37°C. Starvation treatment mentioned in the article refers to serum-free starvation culture for 24h and then cells were used in the subsequent experiment. Brefeldin A (BFA, 20350-15-6, ApexBio) was used in culturing cells for 30min firstly, then replaced with serum-free medium to continue culture for 24h.
Plasmids Construction
The full-length open reading frame of EGFRvIII was amplified from pEGFRP-N1-EGFRvIII plasmid (a gift from Dr. Janusz Rak, Cancer and Angiogenesis Laboratory, MUHCRI-Montreal Children's Hospital Research Institute, Canada) and we directly cloned EGFRvIII into pHBLV-CMVIE-IRES-puro lentiviral plasmid vector, which product is abbreviated as pHBLV-EGFRvIII. On the basis of pHBLV-EGFRvIII, we knocked out the nucleic acid sequence of EGFRvIII expressing 645RRRHIVRKR653 peptide to obtain EGFRvIII-NLS-del with the loss of nuclear localization function, which product is named pHBLV-EGFRvIII-NLS-del. The cDNA of PKM2 come from U251 cell line. We first made a full RNA extraction, and then we got the open reading frame of PKM2 through reverse transcription and PCR technique. PKM2 was cloned into the PITA lentivirus plasmid vector and was added a Flag tag on the N-terminal domain, which was named PITA-Flag-PKM2. On the basis of PITA-Flag-PKM2, we mutated the Arg399/400 of PKM2 into alanine, so that PKM2 lost its nuclear transposition capability and which is named PITA-Flag-R399/400A. All of the above recombinant constructs were verified by DNA sequencing. The shsyntaxin-6-1 (5'-AACCATCAGCATAGTTGAAGCAAATCCTA-3') and shsyntaxin-6-2 (5'-GAAATCGACTGGACCACCAACGAGCTGAG-3') were inserted into lentivirus small hairpin RNA vector pSilencer2.0-u6.
Glioma cells were transfected with lentivirus plasmids according to previously described methods [36]. Followed by selection with puromycin for one week or G418 for ten days. The following are the screening concentrations of the plasmids and the cell lines: pHBLV (puromycin: U251, 0.5ug/ml; C6, 5ug/ml), PITA (G418: C6, 600ug/ml).
Immunofluorescence analysis
The glioma cells were sequentially fixed with 4% polyformaldehyde for 15 minutes and permeabilized with 0.3%Triton-X100 for 10 minutes. Then Cells were blocked with 10% bovine serum albumin at 37°C for 1h. Finally, we incubated cells with primary antibodies overnight at 4°C, and Alex Fluor dye-conjugated secondary antibodies (Fluor®-488 or -594 goat anti-rabbit or anti-mouse IgG (ZSGB-BIO, China)) and Hoechst 33342 together for 1h at 37°C. The primary antibodies include EGFRvIII murine monoclonal antibody (#HTA0001, Beijing Cellonis Biotechnologies Co. Ltd, China), EGFR (D38B1) XP Rabbit mAb (#4267, CST), syntaxin-6 (C34B2) Rabbit mAb (#2869, CST), Anti-GM130 antibody (ab3156, abcam), Calregulin antibody (sc-11398, Santa Cruz), PKM2 (D78A4) XP Rabbit mAb (#4053P, CST). Images were captured under a TCSSP8 Confocal Microscope (Leica, Buffalo Grove, USA). The co-localization coefficient of merged images was processed with Image-Pro Plus6.0 software.
Immunoblotting and immunoprecipitation
RIPA lysates (P0013B, Beyotime, China) and IP lysates (P0013, Beyotime, China) are used for the extraction of protein in these two experiments respectively. The extraction procedure is based on the manufacturer's instructions. Some of the antibodies used have been described in the immunofluorescence analysis, and the rest of the antibodies include GAPDH antibody (TA-08, ZSGB-BIO, China), Lamin B1 antibody (PA532474, Pierce), Histone H3 (D1H2) XPR Rabbit mAb (#4499, CST), p-STAT3 antibody (#9145, CST), STAT3 antibody (#12640, CST), p-ERK1/2 antibody (sc-7383, Santa Cruz), ERK1/2 antibody (sc-514302, Santa Cruz), p-PKM2 ser37 (12822-1, signalway antibody). We used image J software to analyze the gray levels of the immunoblotting results.
Subcellular fractionation
Nuclei and cytosol were isolated using a nuclear extract kit (#P0028) from a Chinese biology company Beyotime. The extraction procedure is based on the manufacturer's instruction, whose isolation process has no much difference compared with general separation steps. The extracted protein was used for immunoblotting analysis.
Xenograft tumor models
For the intracranial xenotransplantation tumor model, 1×105(10μl) cells in PBS were inoculated into the brain of female BALB/c nude mice. All animal experiments were performed in accordance with the Guidelines of Peking University Health Science Center Animal Care and Use Committee.
Immunohistochemistry
Immunohistochemistry analysis involves clinical primary glioma samples and intracranial tumorigenic tissue of nude mice. Glioma tissue samples were routinely fixed in 10% formalin and embedded in paraffin. 5μm-thick tissue sections were deparaffinized in xylene and rehydrated over a graded ethanol series. For antigen retrieval, rehydrated sections were treated in 10mM citrate buffer at pH 6.0 for 20 min in a steamer. Incubating with 3% H2O2 for 10 minutes at room temperature and goat serum for 20 minutes at 37°C in turn. Then sections were immune-stained with primary antibodies against EGFRvIII or p-STAT3. The catalogs have been described in the immunofluorescence analysis. Immunohistochemistry was performed on the Dako Autostainer Plus automated slide-processing system using the Ultravision LP Large Volume Detection System HRP Polymer (ZSGB-BIO, China) for detection of antibody binding according to the manufacturer's protocol. Slides were counterstained with hematoxylin (ZSGB-BIO, China) for several seconds.
Bioluminescence imaging with IVIS
Mice were intraperitoneally injected with 100μl of 7.5 mg/ml D-luciferin (Xenogen), and after 10 minutes they were anaesthetized with isoflurane inhalation. And then Bioluminescence imaging with a CCD camera (IVIS, Xenogen) was used for living imaging with automatic exposure time. Flurorescence emission was normalized to photons per second per centimeter squared per steradian (p/s/cm2/sr) and the final imaging results of three weeks were unified to the same scale size.
Statistical analysis
All data are expressed as the mean ± SD from at least three independent experiments. The Student's two-tailed t-test with significance set at p<0.05 was used to determine statistical significance for each assay. One-way ANOVA was performed to compare more than two groups. Error bars indicate standard deviation. The software of GraphPad Prism 5 was used for statistical analysis and related mapping results.
Results
EGFRvIII nuclear localization is evident in primary human glioblastoma and glioma cells
To verify that EGFRvIII undergoes nuclear translocation, we stained for EGFRvIII in a series of human primary glioblastoma tissues. EGFRvIII experienced nuclear translocation in nearly every tissue sample; we present the typical nuclear translocation results in three different glioma patients' samples (Fig. 1A). According to previous results, EGFRvIII nuclear localization will lead to downstream gene transcription, cell proliferation, drug inhibition, and other effects, contributing to a poor prognosis in patients with glioma. However, the mechanism by which EGFRvIII is transported into the nucleus has not been determined, although EGFR's nuclear translocation has been reported in many studies. The aim of this study was to identify the pathway of EGFRvIII nuclear transport and examine how EGFRvIII nuclear translocation affects downstream signaling molecules to find new therapeutic interventions for the treatment of glioma.
EGFRVIII's nuclear translocation was observed in primary human glioblastoma and glioma cells. A: Immuno-histochemical staining for EGFRvIII of primary human GBM. The red arrows indicate the glioma cells that undergo EGFRvIII's nuclear translocation. Scales bar, 30μm and 10μm. B: Schematic of EGFR/EGFRvIII. EGFRvIII-NLS-del is the protein that the red amino acid sequence was truncated based on EGFRvIII. ECD, extracellular domain; TM, transmembrane domain; JM, juxtamembrane domain; KD, kinase domain; CR, C-terminal regulatory region. NLS, nuclear localization signal. C: EGFRvIII and EGFRvIII-NLS-del were overexpressed in U251 and C6 respectively by lentivirus infection, and followed by immunoblot analysis after one week selection with puromycin. The EGFRvIII channel was detected by EGFRvIII antibody, which can recognize EGFRvIII and EGFRvIII-NLS-del, while the EGFR channel was detected by EGFR antibody, which can identify EGFR, EGFRvIII and EGFRvIII-NLS-del. The successfully constructed cell lines include U251EGFRvIII, U251EGFRvIII-NLS-del, C6EGFRvIII and C6EGFRvIII-NLS-del. F98, F98EGFR and F98npEGFRvIII were referenced cell lines. D: Immunofluorescence staining with EGFRvIII antibody was performed to detect EGFRvIII or EGFRvIII-NLS-del in our constructed cells. Green signals also indicated we successfully constructed our targeted cell lines. F98, F98EGFR and F98npEGFRvIII were referenced cell lines. Scale bar, 50μm. E: Comparison of EGFRvIII's nuclear translocation by immunoblot analysis in U251EGFRvIII and U251EGFRvIII-NLS-del. Non-nucleus and Nucleus were separated after 24h serum-free starvation culture (representing as “starvation” in all figures) and then immunoblot test was carried out. The intensity of bands were determined with image J program and the lower histogram is the relative protein levels nuclear EGFRvIII. Here, we declare that in this article all the gray density statistics are done by image J software, and all the histogram and statistical analysis results are completed by GraphPad Prism 5 software, of which *P<0.05, **P<0.01, ***P<0.001. F: Comparison of EGFRvIII's nuclear translocation by immunofluorescence staining in U251EGFRvIII and U251EGFRvIII-NLS-del. Lower histogram is the statistical results of cell numbers occurring EGFRvIII's nuclear translocation in 100 cells. Scale bar are respectively 50μm and 10μm. G: Comparison of EGFRvIII's nuclear translocation by immunoblot analysis in C6EGFRvIII and C6EGFRvIII-NLS-del. Lower histogram represents the relative densities of the bands. H: Comparison of EGFRvIII's nuclear translocation by immunofluorescence staining in C6EGFRvIII and C6EGFRvIII-NLS-del. Lower histogram is the statistical results of cell numbers occurring EGFRvIII's nuclear translocation in 100 cells. Scale bar are respectively 50μm and 10μm.
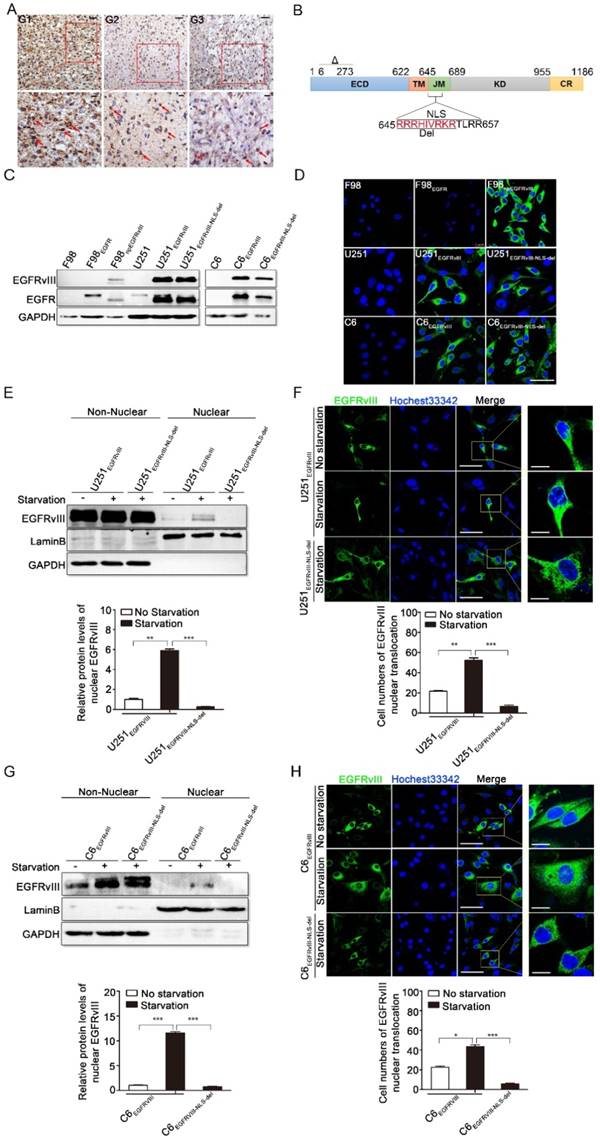
Prior to our research, we constructed a series of cell lines based on the U251 human glioma cell line and C6 rat glioma cell line. Originally, there was no EGFRvIII expression in the U251 and C6 cells. And thus, we overexpressed EGFRvIII and EGFRvIII-NLS-del, respectively, in these lines by lentiviral infection. EGFRvIII-NLS-del has a partial deletion in the nuclear localization signal of EGFRvIII, whose nuclear localization sequence is distributed throughout the juxta-membrane domain (Fig. 1B).
By western blot, using EGFRvIII and EGFR antibodies, we identified successfully constructed cell lines, which we named U251EGFRvIII, U251EGFRvIII-NLS-del, C6EGFRvIII, and C6EGFRvIII-NLS-del; The F98, F98EGFR, and F98npEGFRvIII cell lines were purchased from ATCC as references (Fig. 1C). EGFRvIII antibody can only recognize EGFRvIII and its mutant form, EGFRvIII-NLS-del, whereas EGFR antibody can recognize EGFR, EGFRvIII, and EGFRvIII-NLS-del. The immunofluorescence staining results (Fig. 1D), performed only with EGFRvIII antibody, were consistent with the western blot findings and again suggested we have successfully constructed the target cell lines.
As a constitutively active protein, the nuclear translocation of EGFRvIII does not require ligands stimulation. According to previous studies, EGFRvIII's nuclear translocation is increased by serum-free starvation (referred to as “starvation” in all figures) for 12-24 hours [17, 37]. This model is also based on the fact that glioma cells are sometimes subjected to nutrient deprivation in vivo.
Compared with normal culture conditions, the amount of nuclear EGFRvIII in U251EGFRvIII cells rose after serum-free starvation for 24h in western-blot result, whereas nuclear EGFRvIII-NLS-del was not detected in U251EGFRvIII-NLS-del, indicating that EGFRvIII-NLS-del, with its partially deleted nuclear localization signal, did not undergo nuclear translocation (Fig. 1E). We also performed an immunoblot analysis of nuclear EGFRvIII by using histone H3 as a nuclear marker in U251EGFRvIII, and obtained a consistent EGFRvIII nuclear translocation result, indicating that EGFRvIII can indeed enter the nuclear (Fig. S1-A). Figure 1F shows the immunofluorescence results of U251EGFRvIII and U251EGFRvIII-NLS-del cells, demonstrating that nuclear translocation occurs in U251EGFRvIII but rarely in U251EGFRvIII-NLS-del cells after serum-free starvation for 24h. To be more rigorous, we performed sequential confocal imaging of EGFRvIII through the sections of the nucleus in U251EGFRvIII and U251EGFRvIII-NLS-del (Fig. S1-B), and results showed that EGFRvIII can enter the nucleus instead of EGFRvIII-NLS-del. Figure 1G and 1H, respectively, showed the western blot and immunofluorescence results about EGFRvIII's nuclear transloncation in C6EGFRvIII and C6EGFRvIII-NLS-del cells, also revealing contrasting nuclear translocation results of EGFRvIII and EGFRvIII-NLS-del in corresponding cell lines. In C6EGFRvIII and C6EGFRvIII-NLS-del, we also performed immunoblot analysis of nuclear EGFRvIII by using histone H3 as a nuclear marker (Fig. S2-A) and sequential confocal imaging of EGFRvIII through the sections of the nucleus (Fig. S2-B), which all have consistent experimental verification results about EGFRvIII's nuclear translocation.
These results suggest that EGFRvIII has a nuclear transposition in the clinical tumor tissues and glioma cell lines, and the nuclear transposition was blocked when we knocked out its nuclear location sequence.
Golgi, especially syntaxin-6, participates in EGFRvIII's reverse transposition to the nucleus
To examine whether the Golgi is involved in EGFRvIII's retrograde trafficking, we first asked whether serum-free starvation can stimulate EGFRvIII's localization on the Golgi. We performed double staining for EGFRvIII and syntaxin-6 (STX6), which is a marker of trans Golgi and is involved in protein transport, and then performed imaging with confocal microscopy, by which we observed that EGFRvIII localized on the Golgi apparatus and after serum-free stimulation for 24h, their co-localization signals were enhanced (Fig. 2A). Rigorously, we carried out a statistical analysis of syntaxin-6's positive area with and without serum-free starvation and results showed that serum-free starvation didn't expand the positive region of syntaxin-6 in U251EGFRvIII or C6EGFRvIII (Fig. S3). Besides, we performed reciprocal IP-western blot and observed that serum-free starvation improved the co-localization of EGFRvIII and syntaxin-6 in U251EGFRvIII (Fig. 2B). We did not detect any increase in the amount of their co-localization in U251EGFRvIII-NLS-del, further suggesting that syntaxin-6 is likely to be involved in the nuclear translocation of EGFRvIII. The same co-localization results were validated between EGFRvIII and GM130, a Golgi endosomal marker, by confocal fluorescence microscopy (Fig. 2C).
Golgi marker syntaxin-6 mediates nuclear translocation of EGFRvIII. A: Co-localization results of EGFRvIII and syntxain-6 with or without 24h serum-free starvation culture in U251EGFRvIII and C6EGFRvIII respectively by immunofluorescence staining. Yellow signal indicates the co-localization of the two proteins. Lower histogram is a comparison of co-localization coefficients. Here, we declare that all the statistics of co-localization coefficients in this article are made by Image-Pro Plus 6.0 software. Scale bar are respectively 50μm and 25μm. B: Western blot results after reciprocal immunoprecipitation of EGFRvIII and Syntaxin6. Column below represents the relative densities of bands. C: Co-localization results of EGFRvIII and GM130 with or without 24h serum-free starvation culture in U251EGFRvIII and C6EGFRvIII respectively by immunofluorescence staining. Yellow signal indicates the co-localization of the two proteins. Lower histogram is a comparison of co-localization coefficients. Scale bar are respectively 25μm and 10μm. D: U251EGFRvIII cells were transfected with control shRNA and two shRNA targeting syntaxin-6 (STX6) respectively by lentivirus infection, and after 48h immunoblotting for STX6 were analyzed. The lower histogram was the results of relative densities. E: U251EGFRvIII cells were treated with or without 24h serum-free starvation culture for 24h after STX6 shRNA treatment for 48h, and immunoblotting results of EGFRvIII's nuclear translocation were analyzed in control group and shRNA knockdown group. The lower histogram was the relative densities of nuclear EGFRvIII. F: U251EGFRvIII cells were treated with or without 24h serum-free starvation culture for 24h after STX6 shRNA treatment for 48h, and imunofluorescence results of EGFRvIII's nuclear translocation were analyzed in control group and shRNA knockdown group. Scale bar are respectively 50μm and 25μm. Lower histogram was the cell numbers of EGFRvIII's nuclear translocation in 100 cells.
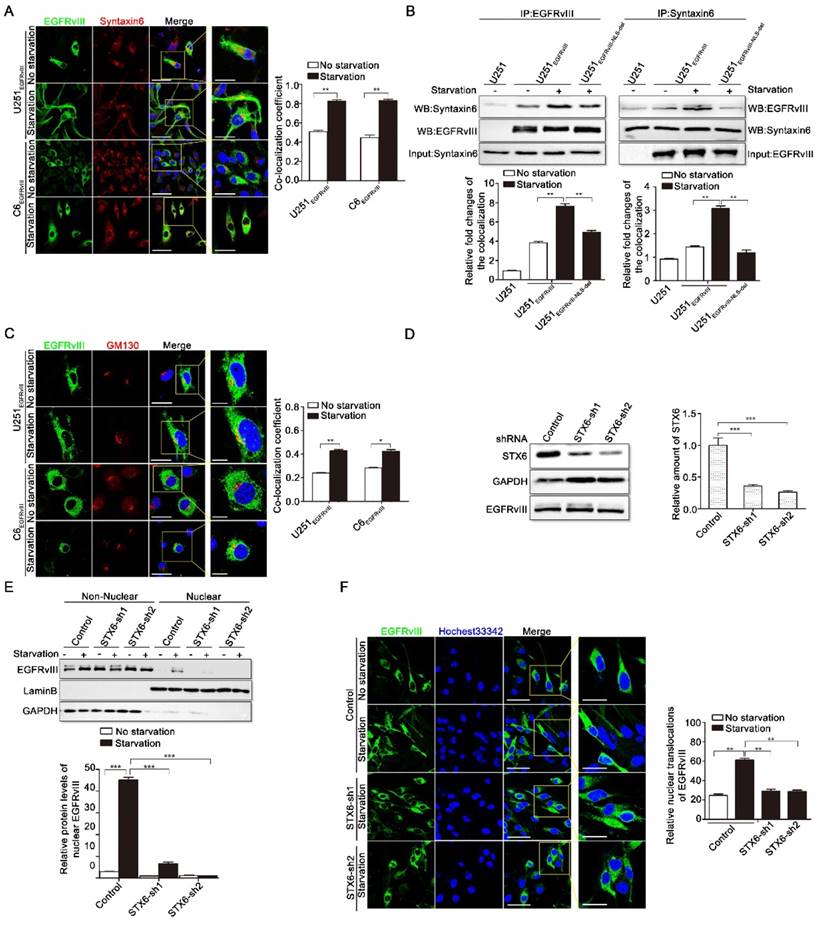
We then studied the function of syntaxin-6 in EGFRvIII's nuclear translocation. To generate positive or negative contrast results, we produced two shRNA constructs targeting syntaxin-6. Figure 2D shows that both compounds have good knockdown efficacy against syntaxin-6.
As indicated in Figure 2E, compared with the control group, the amount of nuclear EGFRvIII decreased when syntaxin-6 was down-regulated by shRNA in U251EGFRvIII cells, showing that syntaxin-6 positively regulates the reverse transport of EGFRvIII to the nucleus. Similarly, the immunofluorescence results in Figure 2F also demonstrated that syntaxin-6 is closely related to the nuclear transport of EGFRvIII, which differed significantly between the control and shRNA groups in the subsequent statistical analysis.
These results suggest that Golgi, in particular syntaxin-6, has a positive effect on EGFRvIII's reverse transposition to the nucleus.
COPI-mediated retrograde trafficking process from Golgi to ER is associated with the nuclear translocation of EGFRvIII
COPI is a pivotal retrograde transporting complex that transfers proteins from cis Golgi to the endoplasmic reticulum. We then examined whether COPI can transfer EGFRvIII from the Golgi apparatus to ER organelles. By double immunofluorescence staining of EGFRvIII with the ER marker calregulin, the localization of EGFRvIII to the ER declined after the addition of BFA compared with serum-free starvation and the DMSO control group (Fig. 3A). BFA is an assembly inhibitor that blocks the transport function of COPI. Moreover, we compared syntaxin-6 control groups with its shRNA group by immunofluorescence imaging and observed that serum-free starvation provoked the co-localization of EGFRvIII and calregulin, while their co-location signal decreased when the expression of syntaxin-6 was down-regulated by shRNA (Fig. 3B). These data indicate that COPI participates in the transport of EGFRvIII from the Golgi apparatus to ER.
COPI mediates reverse transport of EGFRvIII. A: U251EGFRvIII cells were treated with or without 24h serum-free starvation culture, or 24h serum-free starvation culture after adding brefeldin A (BFA) or DMSO for 30min. DMSO is the control of BFA. Immunofluorescence staining were performed to analyze the co-localization of EGFRvIII and calregulin. Right histogram is a comparison of the co-localization coefficients between them. Scale bar, 10μm. B: Immunofluorescence results of the co-localization between EGFRvIII and calregulin in control group and shRNA knockdown group targeting syntaxin6 after treating with or without 24h serum-free starvation culture. Right histogram is a comparison of the co-localization coefficients between them. Scale bar, 25μm. C: U251EGFRvIII cells were treated with or without 24h serum-free starvation culture, or 24h serum-free starvation culture after adding brefeldin A (BFA) or DMSO for 30min, and then immunoblotting analysis were performed to detect the protein levels of nuclear EGFRvIII. DMSO is the control of BFA. Lower histogram was the relative densities of nuclear EGFRvIII. D: U251EGFRvIII cells were treated with or without 24h serum-free starvation culture, or 24h serum-free starvation culture after adding brefeldin A (BFA) or DMSO for 30min, and then immunofluorescence staining were performed to detect the protein levels of nuclear EGFRvIII. DMSO is the control of BFA. Lower histogram was the cell numbers of EGFRvIII's nuclear translocation in 100 cells. Scale bar, 25μm.
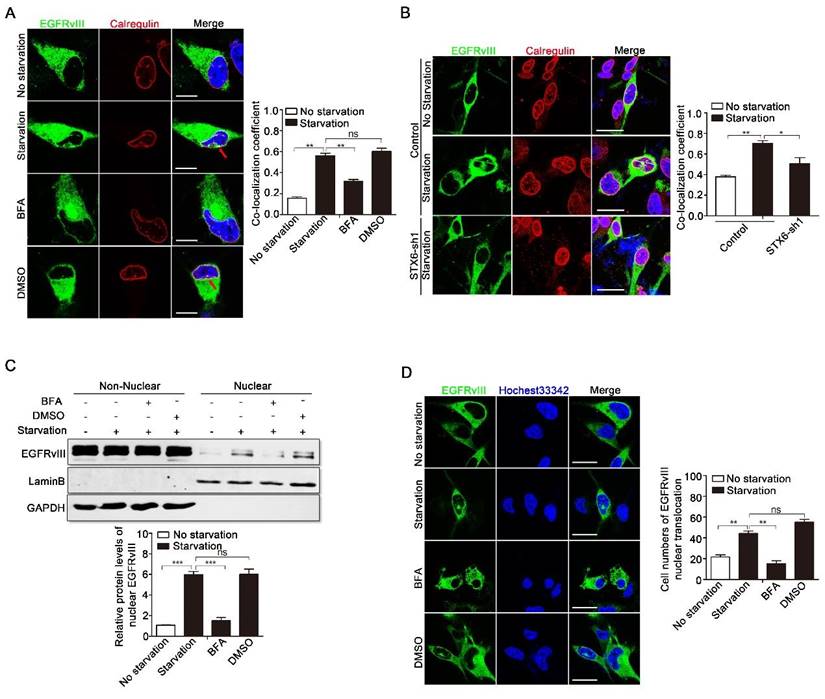
To validate whether COPI is involved in the nuclear translocation of EGFRvIII, we performed immunoblotting analysis, in which BFA was administered to inhibit the activation of ARF1-guanine exchange factors, resulting in the disassembly of the COPI coat on the Golgi; DMSO was used as a control (Fig. 3C). Results showed that compared with the serum-free starvation or DMSO control group, BFA reduced the amount of EGFRvIII in the nucleus. The immunofluorescence results in Figure 3D provided further support that pretreatment with BFA in U251EGFRvIII decreased the cell number of EGFRvIII's nuclear translocation, which suggested that intact COPI is required for EGFRvIII's nuclear translocation.
These results strongly indicate that COPI mediates the retrograde trafficking of EGFRvIII from the Golgi to ER, and meanwhile COPI participates in the reverse nuclear transport of EGFRvIII.
EGFRvIII's nuclear positioning can increase the phosphorylation level of STAT3
As seen in the immunoblotting results in Figure 4A, EGFRvIII, especially under serum-free starvation culture condition, increased the levels of p-STAT3, which is the active form of STAT3. This result suggests that EGFRvIII's nuclear translocation may be closely related to the activation of STAT3. Subsequently, we confirmed this association in the following experiment. In U251EGFRvIII cells, serum-free starvation up-regulated p-STAT3, and meanwhile, compared with the DMSO control group, BFA inhibited the phosphorylation level of STAT3 (Fig. 4B). Existing researches showed that phosphorylated STAT3 often translocate to the nucleus to play its transcriptional regulation function [38-40]. Thus, we performed nuclear and non-nuclear fractionation experiments and detected the level of nuclear p-STAT3 with western blot experiment. EGFRvIII promoted nuclear distribution of p-STAT3 in C6EGFRvIII cells versus in C6 cells, especially under serum-free starvation culture, whereas this increase was not observed in C6EGFRvIII-NLS-del cells (Fig. 4C).
EGFRvIII's nuclear translocation promoted phosphorylation of STAT3. A: Immunoblotting results of p-STAT3 in U251 and U251EGFRvIII with or without serum-free starvation culture for 24h. Lower histogram was the relative densities of p-STAT3. B: U251EGFRvIII cells were treated with or without 24h serum-free starvation culture, or 24h serum-free starvation culture after adding brefeldin A (BFA) or DMSO for 30min, and then immunoblotting analysis were performed to detect the protein levels of p-STAT3. DMSO is the control of BFA. Lower histogram was the relative densities of p-STAT3. C: Immunoblotting analysis of non-nuclear and nuclear p-STAT3 in C6, C6EGFRvIII and C6EGFRvIII-NLS-del after treating with or without 24h serum-free starvation culture. Lower histogram was the relative densities of nuclear p-STAT3. D: Immunoblotting analysis of p-STAT3 in U251, U251EGFRvIII and U251EGFRvIII-NLS-del after treating with or without 24h serum-free starvation culture. Lower histogram was the relative densities of p-STAT3. E: Immunoblotting analysis of p-STAT3 in C6, C6EGFRvIII and C6EGFRvIII-NLS-del after treating with or without 24h serum-free starvation culture. Lower histogram was the relative densities of p-STAT3.
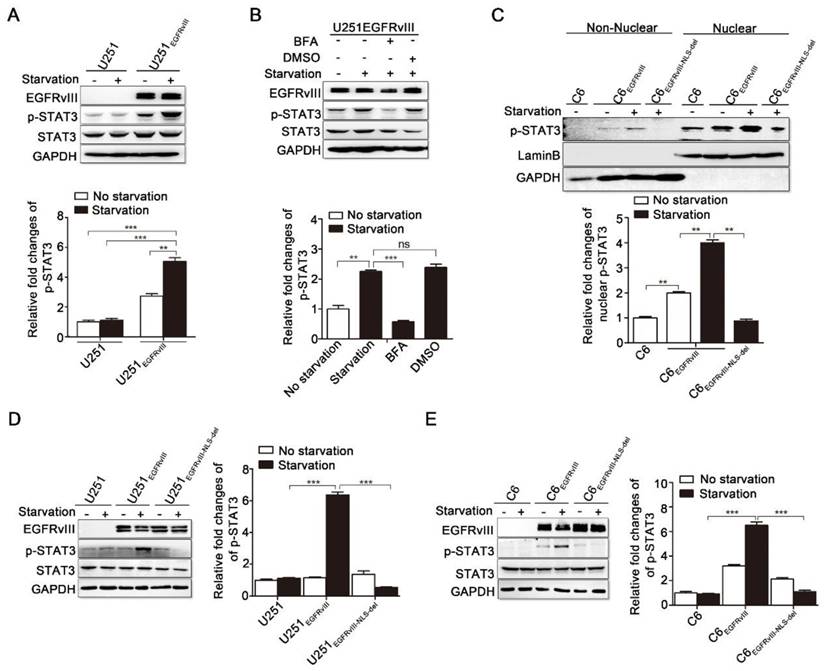
Next, we separately pretreated U251, U251EGFRvIII, and U251EGFRvIII-NLS-del cells in normal culture or under serum-free starvation culture and found that the latter culture condition significantly up-regulated p-STAT3 in U251EGFRvIII but not in U251 or U251EGFRvIII-NLS-del cells (Fig. 4D). We obtained similar outcomes in C6, C6EGFRvIII, and C6EGFRvIII-NLS-del cells (Fig. 4E).
In summary, EGFRvIII's nuclear translocation can enhance the phosphorylation level and nuclear distribution of STAT3, which has a great help for STAT3 to play its transcriptional regulatory function in the nucleus.
Nuclear translocation of EGFRvIII enhances nuclear localization of PKM2
In U251EGFRvIII cells, serum-free starvation improved the cell number of PKM2's nuclear localization compared with normal culture, whereas no such increase occurred in U251EGFRvIII-NLS-del cells (Fig. 5A). We then compared nuclear PKM2 levels by immunoblotting analysis in U251, U251EGFRvIII, and U251EGFRvIII-NLS-del cells. The amount of nuclear PKM2 in U251EGFRvIII cells was significantly higher than in U251 and U251EGFRvIII-NLS-del cells in the same serum-free starvation environment, which environment can increase the nuclear translocation of EGFRvIII (Fig. 5B). Figure 5C and 5D show additional immunofluorescence and western blot results in C6-associated cell lines, confirming the same changes in PKM2's nuclear translocation. Thus, we can say that the EGFRvIII with the complete nuclear transposition function rather than the EGFRvIII-NLS-del can further promote the nuclear translocation of PKM2.
EGFRvIII's nuclear translocation promoted the nuclear localization of PKM2. A: Immunofluorescence results of EGFRvIII in U251EGFRvIII and U251EGFRvIII-NLS-del cells after treating with or without 24h serum-free starvation culture. Scale bar, 10μm. Right histogram is the cell numbers of PKM2's nuclear localization in 100 cells. B: Immunoblotting results of non-nuclear and nuclear EGFRvIII in U251, U251EGFRvIII and U251EGFRvIII-NLS-del cells after treating with or without 24h serum-free starvation culture. Right histogram was the relative density analysis of nuclear PKM2. C: Immunofluorescence results of EGFRvIII in C6EGFRvIII and C6EGFRvIII-NLS-del cells after treating with or without 24h serum-free starvation culture. Scale bar, 10μm. Right histogram is the cell numbers of PKM2's nuclear localization in 100 cells. D: Immunoblotting results of non-nuclear and nuclear EGFRvIII in C6, C6EGFRvIII and C6EGFRvIII-NLS-del cells after treating with or without 24h serum-free starvation culture. Right histogram was the relative density analysis of nuclear PKM2. E: U251EGFRvIII cells were treated with or without 24h serum-free starvation culture, or 24h serum-free starvation culture after adding brefeldin A (BFA) or DMSO for 30min, and then immunoblotting analysis were performed to detect the protein levels of non-nuclear and nuclear PKM2. DMSO is the control of BFA. Right histogram was the relative density analysis of nuclear PKM2. F: Flag-R399/400A was overexpressed in C6EGFRvIII or C6EGFRvIII-NLS-del respectively by lentivirus infection, and followed by ten days selection with neomycin. Immunoblotting analysis were performed to detect the protein levels of non-nuclear and nuclear Flag-R399/400A after treating cells with or without 24h serum-free starvation serum-free starvation culture. Right histogram was the relative density analysis of nuclear PKM2.
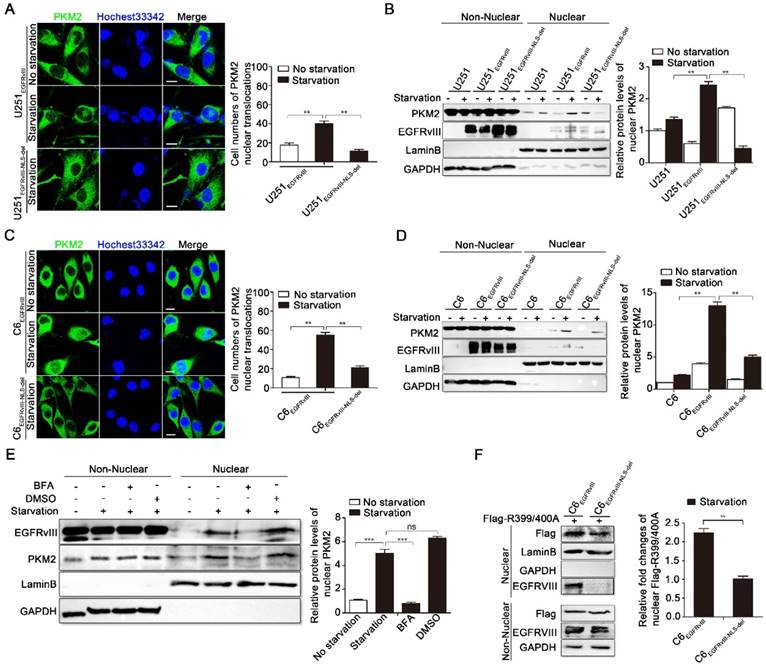
In U251EGFRvIII, we performed further validation with EGFRvIII's nuclear translocation inhibitor BFA. Results showed that, when EGFRVIII's nuclear translocation was inhibited by BFA, the amount of nuclear PKM2 also decreased compared with the serum-free starvation group and DMSO control group (Fig.5E).
To further confirm the role of EGFRvIII's nuclear translocation in regulating PKM2 entry into the nucleus, we constructed two cell lines for subsequent experiments. We respectively overexpressed Flag-R399/400A in C6EGFRvIII and C6EGFRvIII-NLS-del cells via lentiviral infection; R399/400A is a mutant of PKM2 in which Arg399/400 is changed to alanine, causing a failure in PKM2's nuclear translocation. Figure 5F indicated that, in the same serum-free starvation environment, EGFRvIII enhanced the nuclear translocation of Flag-R399/400A more effectively than EGFRvIII-NLS-del.
In conclusion, EGFRVIII's nuclear translocation can increase the localization of PKM2 in the nucleus, while nuclear PKM2 is an important prerequisite for its many regulatory functions.
EGFRvIII's nuclear translocation can promote the progression of glioblastoma in vivo
Next, we determined the effects of EGFRvIII's nuclear translocation on the development of GBM.
C6, C6EGFRvIII, and C6EGFRvIII-NLS-del cells that stably overexpress luciferase were injected separately into the brains of nude mice, and these three kinds of GBM animals were imaged through a luciferase reporter assay device on days 7, 14, and 21 after heterologous transplantation. The luminescence signals in the C6EGFRvIII group were higher than the C6 and C6EGFRvIII-NLS-del groups in the three imaging results, and meanwhile the signal intensity in C6EGFRvIII-NLS-del cells exceeded that in C6 cells (Fig. 6A); the statistical results describe the trend in signal intensity in the three groups (Fig. 6B). In addition, we sampled the above three kinds of tumorigenic brain tissues and analyzed them by immune-histochemical staining. Figure 6C showed the p-STAT3 level in three tissues, and the highest expression of p-STAT3 occurred in EGFRvIII- overexpressing tissues.
EGFRvIII's nuclear translocation promoted the progress of GBM in vivo. A: C6、C6EGFRvIII and C6EGFRvIII-NLS-del cells with stably expressing luciferase were injected into the brain of BALB/c nude mice with intracranial injection technique. Luciferase signal in different mouse models was detected every 7 days in vivo by bioluminescence imaging (IVIS 100 Imaging System) to monitor the development of GBM until 3 times. n=5. B: Plotting diagram shows the changes of relative luminescence in the indicated groups of mice during 3 weeks. Data are means ± S.D. n=5. C: Immuno-histochemical staining for EGFRvIII and p-STAT3 in xenotransplantation glioma tissues originating from C6, C6EGFRvIII, C6EGFRvIII-NLS-del cell lines. Scare bar, 30μm.
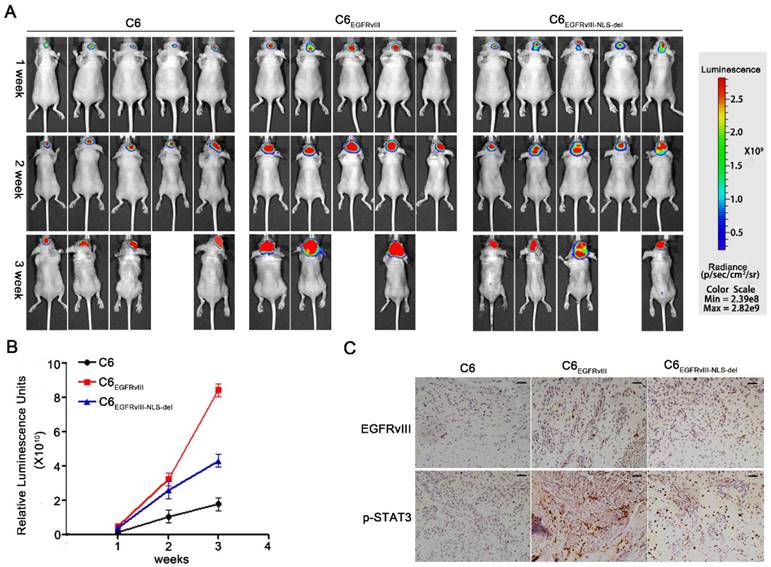
Compared with C6 and C6EGFRvIII-NLS-del groups, EGFRvIII, having intact nuclear transposition ability, in C6EGFRvIII accelerated disease progression significantly. These results are consistent with previous studies that EGFRvIII, as a constitutively active protein, particularly with regard to its nuclear translocation, provokes the occurrence and progression of GBM [3, 41, 42].
Discussion
EGFRvIII, the most common mutant of EGFR, coexists with EGFR in glioma cells and does not exist alone. In the absence of ligands, EGFRvIII also can activate downstream cascade signals by forming a heterogeneous or homologous polymer, promoting the proliferation, migration, and infiltration of glioma cells. As nuclear localization proteins, EGFR/ EGFRvIII can activate STAT3 and STAT5 through its tyrosine kinase function during nuclear translocation and can act as transcription factors in the nucleus, interacting with RHA (RNA helicase A), E2F, and STAT3/5. These pathways are often associated with inflammatory reactions, tumorigenesis or cancer progression. Although the nuclear translocation pathway of EGFR has been reported extensively, there is no detailed study on the mechanism of EGFRvIII's nuclear translocation.
Our research revealed complicated reverse nuclear trafficking of EGFRvIII, which involves the Golgi and ER and the participation of syntaxin-6 and COPI. Based on our results and previous studies, we generated a model of EGFRvIII's reverse transposition from endocytosis to nuclear localization (Fig.7).
A schematic model of EGFRvIII's reverse nuclear trafficking: after endocytosis, EGFRvIII is recognized, submitted and transported to trans Golgi by syntaxin6; COPI mediates the shuttle of EGFRvIII on Golgi apparatus and the retrograde trafficking from Golgi to ER; Finally, EGFRvIII is transported to the nucleus; EGFRvIII's nuclear translocation can increase the phosphorylation of STAT3 and promote the nuclear localization of PKM2.
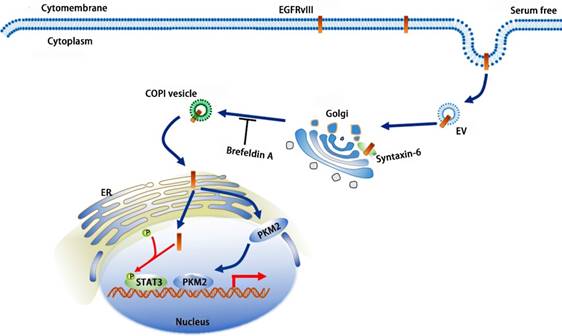
This process includes several key nodes: (1) as a SNARE protein and located primarily on the trans Golgi, syntaxin-6 is responsible for the identification, presentation, and transportation of EGFRvIII toward the cytoplasm; (2) COPI can identify EGFRvIII, shuttle it back to the cis Golgi, and transfer it from the cis Golgi to ER; (3) inhibition of sytaxin6 or COPI disassembly weakens the localization of EGFRvIII in the nucleus, which demonstrates their importance in EGFRvIII's reverse nuclear trafficking; and (4) the nuclear translocation of EGFRvIII facilitates the phosphorylation of STAT3, which often has a role in cancer progression, and when the nuclear translocation of EGFRvIII is inhibited, the levels of p-STAT3 decrease. In conclusion, our study reveals a mechanism by which EGFRvIII undergoes reverse nuclear translocation and syntaxin-6 and COPI implicate in this process.
Overexpression of PKM2 and its nuclear translocation are important markers of Warburg effect and tumorigenesis. Although the promotion of PKM2's nuclear translocation by EGFR has been elucidated [34, 35, 43], we found the relationship between EGFRvIII's nuclear transposition and PKM2's nuclear localization, that is, the former having a significant promoting effect on the latter.
Our nude mouse experiments demonstrated that EGFRvIII's nuclear translocation has a prominent effect on malignant progression of glioma. Numerous literatures have shown that the level of p-STAT3 or PKM2's nuclear transposition has a positive effect on tumors' growth. This paper shows that EGFRvIII's nuclear transposition is the key upstream mechanism for the change of the above two. Therefore, EGFRvIII and its nuclear transposition may be an important target for the treatment of glioma.
In conclusion, EGFRvIII's nuclear transposition is a key point in the progression of glioma, and the revelation of its nuclear translocation process may help us to find new therapeutic strategies.
Supplementary Material
Supplementary figures.
Acknowledgements
This work was co-funded by the National Natural Science Foundation of China (grant number: 81272790), and the National Major Scientific Instruments and Equipment Development Projects (grant number: 2011YQ13001806).
Competing Interests
The authors have declared that no competing interest exists.
References
1. Yarden Y, Sliwkowski MX. Untangling the ErbB signalling network. Nature reviews Molecular cell biology. 2001;2:127-137
2. Marmor MD, Skaria KB, Yarden Y. Signal transduction and oncogenesis by ErbB/HER receptors. International Journal of Radiation Oncology* Biology* Physics. 2004;58:903-913
3. Fan Q-W, Cheng CK, Gustafson WC, Charron E, Zipper P, Wong RA, Chen J, Lau J, Knobbe-Thomsen C, Weller M. EGFR phosphorylates tumor-derived EGFRvIII driving STAT3/5 and progression in glioblastoma. Cancer cell. 2013;24:438-449
4. Brennan CW, Verhaak RG, McKenna A, Campos B, Noushmehr H, Salama SR, Zheng S, Chakravarty D, Sanborn JZ, Berman SH. The somatic genomic landscape of glioblastoma. Cell. 2013;155:462-477
5. Furnari FB, Cloughesy TF, Cavenee WK, Mischel PS. Heterogeneity of epidermal growth factor receptor signalling networks in glioblastoma. Nature Reviews Cancer. 2015;15:302-310
6. Pedersen M, Meltorn M, Damstrup L, Poulsen H. The type III epidermal growth factor receptor mutation: Biological significance and potential target for anti-cancer therapy. Annals of Oncology. 2001;12:745-760
7. Gensburger C, Labourdette G, Sensenbrenner M. Brain basic fibroblast growth factor stimulates the proliferation of rat neuronal precursor cells in vitro. FEBS letters. 1987;217:1-5
8. Cloughesy TF, Cavenee WK, Mischel PS. Glioblastoma: from molecular pathology to targeted treatment. Annual Review of Pathology: Mechanisms of Disease. 2014;9:1-25
9. Yang J, Yan J, Liu B. Targeting EGFRvIII for glioblastoma multiforme. Cancer letters. 2017;403:224-230
10. Furnari FB, Fenton T, Bachoo RM, Mukasa A, Stommel JM, Stegh A, Hahn WC, Ligon KL, Louis DN, Brennan C. Malignant astrocytic glioma: genetics, biology, and paths to treatment. Genes & development. 2007;21:2683-2710
11. Liu F, Hon GC, Villa GR, Turner KM, Ikegami S, Yang H, Ye Z, Li B, Kuan S, Lee AY. EGFR mutation promotes glioblastoma through epigenome and transcription factor network remodeling. Molecular cell. 2015;60:307-318
12. Huang H-JS, Nagane M, Klingbeil CK, Lin H, Nishikawa R, Ji X-D, Huang C-M, Gill GN, Wiley HS, Cavenee WK. The enhanced tumorigenic activity of a mutant epidermal growth factor receptor common in human cancers is mediated by threshold levels of constitutive tyrosine phosphorylation and unattenuated signaling. Journal of Biological Chemistry. 1997;272:2927-2935
13. Sugawa N, Ekstrand AJ, James CD, Collins VP. Identical splicing of aberrant epidermal growth factor receptor transcripts from amplified rearranged genes in human glioblastomas. Proceedings of the National Academy of Sciences. 1990;87:8602-8606
14. Edwards J, Traynor P, Munro AF, Pirret CF, Dunne B, Bartlett JM. The role of HER1-HER4 and EGFRvIII in hormone-refractory prostate cancer. Clinical Cancer Research. 2006;12:123-130
15. de la Iglesia N, Konopka G, Puram SV, Chan JA, Bachoo RM, You MJ, Levy DE, DePinho RA, Bonni A. Identification of a PTEN-regulated STAT3 brain tumor suppressor pathway. Genes & development. 2008;22:449-462
16. Lo H-W, Cao X, Zhu H, Ali-Osman F. Cyclooxygenase-2 is a novel transcriptional target of the nuclear EGFR-STAT3 and EGFRvIII-STAT3 signaling axes. Molecular Cancer Research. 2010;8:232-245
17. Latha K, Li M, Chumbalkar V, Gururaj A, Hwang Y, Dakeng S, Sawaya R, Aldape K, Cavenee WK, Bogler O. Nuclear EGFRvIII-STAT5b complex contributes to glioblastoma cell survival by direct activation of the Bcl-XL promoter. International journal of cancer. 2013;132:509-520
18. Du Y, Shen J, Hsu J, Han Z, Hsu M, Yang C, Kuo H, Wang Y, Yamaguchi H, Miller S. Syntaxin 6-mediated Golgi translocation plays an important role in nuclear functions of EGFR through microtubule-dependent trafficking. Oncogene. 2014;33:756-770
19. Wang Y-N, Wang H, Yamaguchi H, Lee H-J, Lee H-H, Hung M-C. COPI-mediated retrograde trafficking from the Golgi to the ER regulates EGFR nuclear transport. Biochemical and biophysical research communications. 2010;399:498-504
20. Dittmann K, Mayer C, Rodemann HP. Nuclear EGFR as novel therapeutic target. Strahlentherapie und Onkologie. 2010;186:1-6
21. Mittar S, Ulyatt C, Howell GJ, Bruns AF, Zachary I, Walker JH, Ponnambalam S. VEGFR1 receptor tyrosine kinase localization to the Golgi apparatus is calcium-dependent. Experimental cell research. 2009;315:877-889
22. Robertson BJ, Park RD, Snider MD. Role of vesicular traffic in the transport of surface transferrin receptor to the Golgi complex in cultured human cells. Archives of biochemistry and biophysics. 1992;292:190-198
23. Akgoz M, Kalyanaraman V, Gautam N. Receptor-mediated reversible translocation of the G protein βγ complex from the plasma membrane to the Golgi complex. Journal of Biological Chemistry. 2004;279:51541-51544
24. Springer S, Spang A, Schekman R. A primer on vesicle budding. Cell. 1999;97:145-148
25. Taylor TC, Kahn RA, Melançon P. Two distinct members of the ADP-ribosylation factor family of GTP-binding proteins regulate cell-free intra-Golgi transport. Cell. 1992;70:69-79
26. Lee MC, Miller EA, Goldberg J, Orci L, Schekman R. Bi-directional protein transport between the ER and Golgi. Annu Rev Cell Dev Biol. 2004;20:87-123
27. Peyroche A, Antonny B, Robineau S, Acker J, Cherfils J, Jackson CL. Brefeldin A acts to stabilize an abortive ARF-GDP-Sec7 domain protein complex: involvement of specific residues of the Sec7 domain. Molecular cell. 1999;3:275-285
28. Christofk HR, Vander Heiden MG, Harris MH, Ramanathan A, Gerszten RE, Wei R, Fleming MD, Schreiber SL, Cantley LC. The M2 splice isoform of pyruvate kinase is important for cancer metabolism and tumour growth. Nature. 2008;452:230-233
29. Christofk HR, Vander Heiden MG, Wu N, Asara JM, Cantley LC. Pyruvate kinase M2 is a phosphotyrosine-binding protein. Nature. 2008;452:181-186
30. Altenberg Ba, Greulich K. Genes of glycolysis are ubiquitously overexpressed in 24 cancer classes. Genomics. 2004;84:1014-1020
31. Vander Heiden MG, Cantley LC, Thompson CB. Understanding the Warburg effect: the metabolic requirements of cell proliferation. science. 2009;324:1029-1033
32. Koppenol WH, Bounds PL, Dang CV. Otto Warburg's contributions to current concepts of cancer metabolism. Nature Reviews Cancer. 2011;11:325-337
33. Cairns RA, Harris IS, Mak TW. Regulation of cancer cell metabolism. Nature Reviews Cancer. 2011;11:85-95
34. Yang W, Zheng Y, Xia Y, Ji H, Chen X, Guo F, Lyssiotis CA, Aldape K, Cantley LC, Lu Z. ERK1/2-dependent phosphorylation and nuclear translocation of PKM2 promotes the Warburg effect. Nature cell biology. 2012;14:1295
35. Yang W, Xia Y, Ji H, Zheng Y, Liang J, Huang W, Gao X, Aldape K, Lu Z. Nuclear PKM2 regulates [bgr]-catenin transactivation upon EGFR activation. Nature. 2011;480:118-122
36. Sun H, Zhang M, Cheng K, Li P, Han S, Li R, Su M, Zeng W, Liu J, Guo J. Resistance of glioma cells to nutrient-deprived microenvironment can be enhanced by CD133-mediated autophagy. Oncotarget. 2016;7:76238
37. Zheng Q, Han L, Dong Y, Tian J, Huang W, Liu Z, Jia X, Jiang T, Zhang J, Li X. JAK2/STAT3 targeted therapy suppresses tumor invasion via disruption of the EGFRvIII/JAK2/STAT3 axis and associated focal adhesion in EGFRvIII-expressing glioblastoma. Neuro-oncology. 2014;16:1229-1243
38. Reich NC, Liu L. Tracking STAT nuclear traffic. Nature Reviews Immunology. 2006;6:602-612
39. Ernst M, Putoczki TL. Stat3: linking inflammation to (gastrointestinal) tumourigenesis. Clinical and Experimental Pharmacology and Physiology. 2012;39:711-718
40. Nicolas CS, Amici M, Bortolotto ZA, Doherty A, Csaba Z, Fafouri A, Dournaud P, Gressens P, Collingridge GL, Peineau S. The role of JAK-STAT signaling within the CNS. Jak-Stat. 2013;2:22162-23996
41. Gururaj AE, Gibson L, Panchabhai S, Bai M, Manyam G, Lu Y, Latha K, Rojas ML, Hwang Y, Liang S. Access to the nucleus and functional association with c-Myc is required for the full oncogenic potential of ΔEGFR/EGFRvIII. Journal of Biological Chemistry. 2013;288:3428-3438
42. Bonavia R, Inda M, Vandenberg S, Cheng S, Nagane M, Hadwiger P, Tan P, Sah D, Cavenee W, Furnari F. EGFRvIII promotes glioma angiogenesis and growth through the NF-κB, interleukin-8 pathway. Oncogene. 2012;31:4054-4066
43. Li N, Feng L, Liu H, Wang J, Kasembeli M, Tran MK, Tweardy DJ, Lin SH, Chen J. PARP inhibition suppresses growth of EGFR-mutant cancers by targeting nuclear PKM2. Cell reports. 2016;15:843-856
Author contact
Corresponding authors: shenliedu.cn (Li Shen), hqhedu.cn (Qihua He).

 Global reach, higher impact
Global reach, higher impact